Escherichia coli is a rod-shaped (bacillus) Gram-negative bacterium that is frequently used as a model organism. Factors such as its ability to grow fast using cheap media and availability of molecular tools to perform genetic manipulations are favorable for using E. coli as a model organism in molecular genetics.
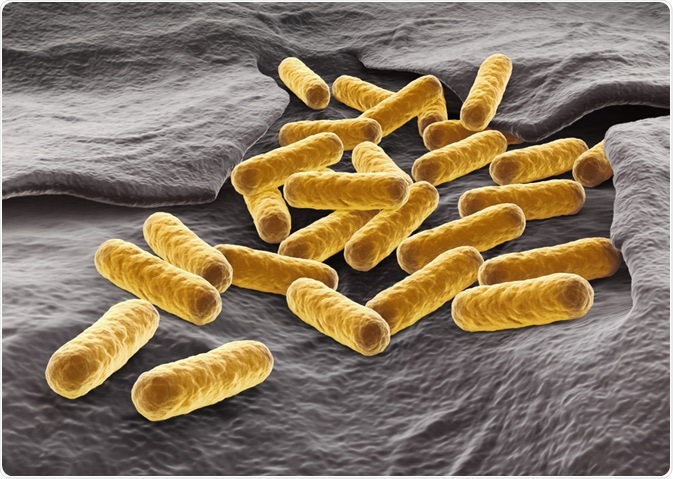 Image Credits: martynowi.cz / Shutterstock.com
Image Credits: martynowi.cz / Shutterstock.com
These tools include plasmids, which are fragments of extrachromosomal DNA that have the ability to confer various phenotypes to E. coli. Its use as a model organism is utilized in metabolic engineering, where the metabolism of E. coli is manipulated to produce specific products.
What is E. coli?
E. coli is a Gram-negative rod-shaped bacterium which is typically around 1µm long and 0.35µm wide. It is able to survive with or without oxygen and thus is a “facultative aerobe”. While it is able to grow fast without forming clumps in various inexpensive chemically defined media, it is not typically tolerant to very high or low temperatures or extreme acidity/alkalinity.
Its potential for fast growth, the number of molecular techniques for genetic manipulations available, and a good amount of knowledge about its genetics means that it has the versatility to be utilized in many ways. Also, E. coli mainly replicates asexually, meaning that modifications made to the genome are maintained and thus effects seen in these mutants are reproducible. These factors make E. coli a good model organism for molecular genetics.
What discoveries were made using E. coli as a model organism?
Several key discoveries in the field of molecular biology, including molecular genetics, were achieved using E. coli as a model organism. This includes an understanding of the genetic code, the mechanisms of DNA replication, the discovery of the genetic operon systems, and the creation of a genetically modified organism.
The E. coli genome
E. coli strains display variation in their pathogenic ability, from strains that do not cause diseases to those that cause severe infections, such as the enterohemorrhagic strain O157:H7. This is evident in the variations seen in E. coli genomes; studies have found that similarity between the genomes of E. coli strains K-12 (a laboratory strain), O157:H7 and CFT073 (a uropathogenic strain) is only 39%.
There are 16,000 genes found in all E. coli strains, which is its pan-genome. However, the core genome of E. coli (i.e. genes present in all strains) is less than 20% of the 16,000 genes in the pan-genome.
The rest of the genome thus is the “flexible genome”; this includes prophages, transposable elements and accessory genes, and they can be acquired horizontally. These genes can confer various phenotypes to E. coli, such as having a more flexible metabolism. This ability of E. coli to incorporate elements in the flexible genome makes it amenable to use as a model organism.
E. coli and its use as a model organism in metabolic engineering
There is a growing demand for a sustainable source of fuels and chemicals. This can potentially be achieved by using renewable sources such as biomass and wastewater as a starting source.
Because there are tools available to manipulate the genome of E. coli, it is a good candidate as a model organism for metabolic engineering; this is where E. coli is genetically manipulated so that it becomes able to produce desired chemicals from various sources during growth.
Modern techniques can be applied to optimize the production of engineered chemicals; this includes the integration of systems biology with metabolic engineering. For example, analysis of the proteins produced by an E. coli strain, or its proteome, can be used as a guide.
In a study, a proteomics approach was used to assess which E. coli membrane proteins were linked to phenylpropanoid tolerance and transport, and thus enabled the identification of potential target proteins which can be utilized in metabolic engineering.
Plasmids; increasing the potential of E. coli as a model organism
Plasmids are fragments of DNA separate from the chromosome, which can replicate itself. The main components of plasmids include the origin of replication and a selection marker such as a gene conferring antibiotic resistance. These plasmids can be manipulated, so a new phenotype can be conferred to E. coli strains by the incorporation of these modified plasmids.
How can plasmids be used to engineer an E. coli strain?
Plasmids can play a part in improving an E. coli strain in the eyes of metabolic engineering. A study by Rodriguez and co. used a modified plasmid to study shikimate production in E. coli.
Shikimate is a product which is formed as a part of the aromatic amino acid biosynthesis pathway. Shikimate is also the starting material for the production of an influenza treatment, therefore there is interest in finding efficient ways of producing it.
Rodrigues and co. modified an E. coli strain to be used as a model organism that can produce shikimate to study how this affects the overall metabolism of E. coli. They placed a plasmid into an E. coli strain which gave it the ability to use glucose to produce high amounts of shikimate.
They found that the modified E. coli strain utilized more glucose, but its growth was not affected. Further analysis showed that the increase in shikimate production reduced oxidative and fermentation pathways, indicating that there is a shift in overall metabolism caused by the production of shikimate.
Sources
Blount, Z. D. (2015) The Natural History of Model Organisms: The unexhausted potential of E. coli. eLife DOI: 10.7554/eLife.05826
Idalia, V.-M. N., and Bernardo, F. (2017) Escherichia coli as a Model Organism and Its Application in Biotechnology. Escherichia coli - Recent Advances on Physiology, Pathogenesis and Biotechnological Applications DOI: 10.5772/67306
Adamczyk, P. A., and Reed, J. L. (2017) Escherichia coli as a model organism for systems metabolic engineering. Current Opinion in Systems Biology. https://doi.org/10.1016/j.coisb.2017.11.001
Rodriguez, A. et al. (2017) Plasmid-encoded biosynthetic genes alleviate metabolic disadvantages while increasing glucose conversion to shikimate in an engineered Escherichia coli strain. Biotechnology and Bioengineering
Further Reading
Last Updated: May 28, 2020