To combat the global spread of coronavirus disease (COVID-19), improved methods for detecting SARS-CoV-2 and viral load quantification are urgently needed. Albeit the reverse transcription-polymerase chain reaction (RT-qPCR) is a current gold standard, quantification necessitates external standards or references with often variable results.
This is where digital PCR comes into play, with very sensitive detection and viral load analysis of SARS-CoV-2; nonetheless, the reaction time is protracted and takes approximately four hours compared to qPCR, which requires one hour.
Likewise, CRISPR-based methods can also detect SARS-CoV-2 within one hour but do not enable the absolute quantification of viral particles, which would reduce inter-laboratory variability and open the scientific field even more.
This research endeavor, led by Dr. Xiaolin Wu from the Singapore-MIT Alliance for Research and Technology, reveals an optimized RApid DIgital Crispr Approach (RADICA) that enables absolute quantification of viral nucleic acids at a constant temperature in one hour time.
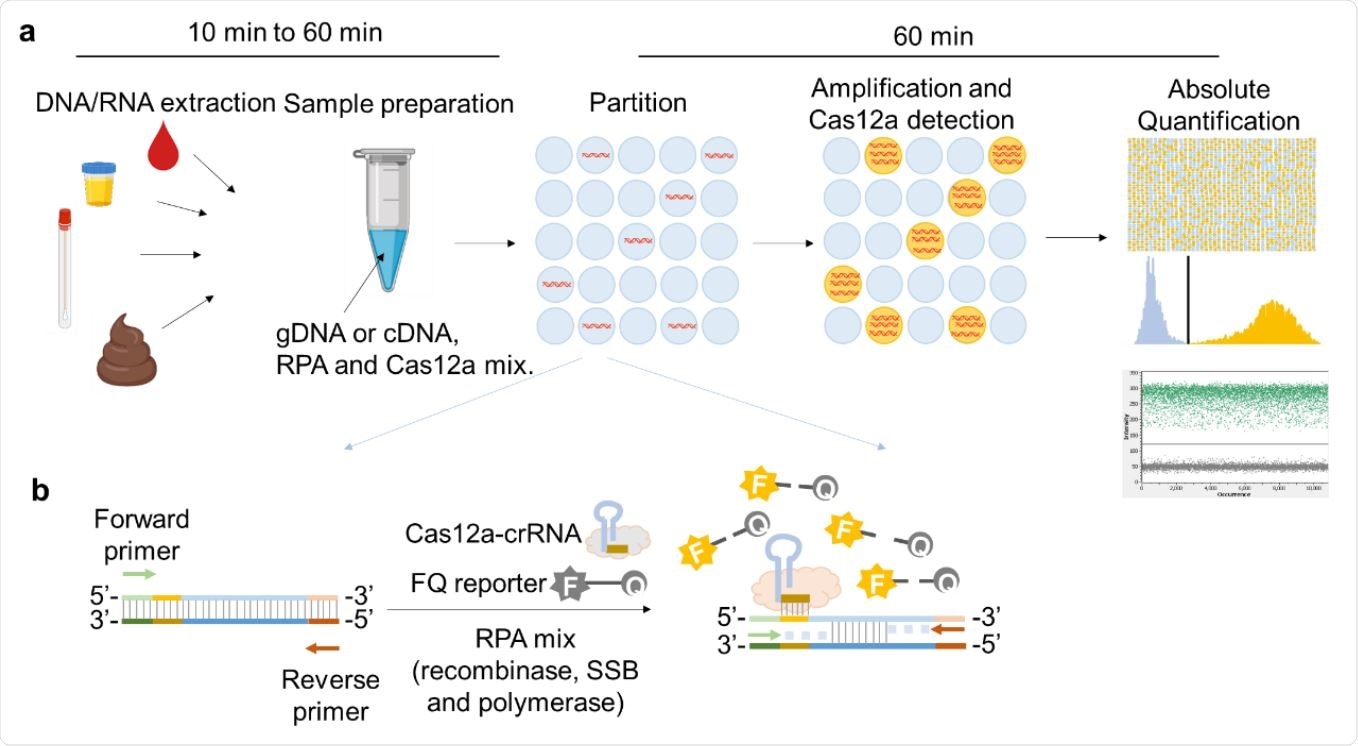
Schematic illustration of RADICA. a, The workflow of RADICA sample partitioning on a chip for absolute quantification of nucleic acid targets. Generally, after the DNA/RNA extraction step, different kind of clinical samples can be used for detection and quantification of various targets. The sample mixture containing DNA/cDNA, RPA reagents, and Cas12a-crRNA715 FQ probes is distributed randomly into thousands of partitions. In each partition, the DNA is amplified by RPA and detected by Cas12a-crRNA, resulting in a fluorescent signal in the partition. Based on the proportion of positive partitions and on Poisson distribution, the absolute copy number of the nucleic acid target is quantified. b, Illustration of RPA-Cas12a reaction in each positive partition. In each partition containing the target nucleic acid, the primers bind to the target nucleic acid and initiate amplification with the aid of recombinase and DNA polymerase. Because of the strand displacement of DNA polymerase, the exposed crRNA-targeted ssDNA sites are bound by Cas12a-crRNA complexes. Cas12a is then activated and cleaves the nearby FQ reporters to produce a fluorescence readout.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Streamlined one-pot reaction on a commercial high-density chip
The method presented in this paper combines all the advantages of quantitative digital PCR, rapid isothermal amplification, as well as specific CRISPR detection into a one-pot reaction system that separates the individual reactions into ten thousand compartments on a commercial high-density chip.
Such a streamlined one-pot reaction combines CRISPR-based detection and nucleic acid amplification into a single step in a closed tube, significantly reducing the risk of sample cross-contamination during batch processing.
For validation purposes, DNA containing the nucleoprotein gene of SARS-CoV-2 was used, which showed linear signal-to-input response. Furthermore, RADICA was compared to digital PCR for quantifying synthetic SARS-CoV-2 DNA and Epstein-Barr (EBV) viral DNA.
The researchers have also tested for possible inhibitory effects of background DNA on reactions that were carried out in small partitions. The isothermal feature of RADICA-based detection assay utilizes a simple constant-temperature heat bath, which can enable rapid viral detection that can be utilized even in low-resource areas.
Compounding effect of precision and speed
In this study, the use of RADICA method resulted in sensitivity and detection limits comparable with those of real-time PCR and other isothermal methods, with the ability of absolute quantification. Furthermore, a significant advantage of RADICA observed in this study was its speed.
The aforementioned absolute quantification of DNA showed a dynamic range from 0.6 to 2027 copies per microliter, without any cross-reactivity on similar viruses or human background DNA. This study's findings also suggest that background DNA will not inhibit the RADICA sample reaction within the dynamic range that will be used for testing purposes.
Moreover, when speed is concerned, RADICA can precisely detect and quantify nucleic acid in one hour without thermal cycling, which is four times faster alternative when compared to digital PCR-based virus detection.
Potential uses of breakthrough technology
"We have established and characterized RADICA, which combines the speed and sensitivity of isothermal amplification, the specificity of CRISPR-based detection, and the ability to obtain absolute quantification by sample partitioning" study authors summarize their methodological breakthrough.
The absolute quantification approach for viral detection may not only support easier clinical processing but could also be exploited for research purposes. And since viral loads can be linked to transmission risk and disease severity (as is the case with SARS-CoV-2), this may prove pivotal in tackling this pandemic as well.
Future work will concentrate on expanding the applications of RADICA to scientific areas such as the analysis of gene expression, detection of rare mutants, sequencing library quantification and copy number variation – with significant benefits for clinical, pharmaceutical, ecological and public health fields.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Wu, X. et al. (2020). A Digital CRISPR-based Method for the Rapid Detection and Absolute Quantification of Viral Nucleic Acids. medRxiv. https://doi.org/10.1101/2020.11.03.20223602, https://www.medrxiv.org/content/10.1101/2020.11.03.20223602v1
- Peer reviewed and published scientific report.
Wu, Xiaolin, Joshua K. Tay, Chuan Keng Goh, Cheryl Chan, Yie Hou Lee, Stacy L. Springs, De Yun Wang, Kwok Seng Loh, Timothy K. Lu, and Hanry Yu. 2021. “Digital CRISPR-Based Method for the Rapid Detection and Absolute Quantification of Nucleic Acids.” Biomaterials 274 (July): 120876. https://doi.org/10.1016/j.biomaterials.2021.120876. https://www.sciencedirect.com/science/article/abs/pii/S0142961221002325?via%3Dihub.