Flagella are primarily used for cell movement and are found in prokaryotes as well as some eukaryotes. The prokaryotic flagellum spins, creating forward movement by a corkscrew shaped filament. A prokaryote can have one or several flagella, localized to one pole or spread out around the cell.
Structure and Origin
The flagellum is comprised of a body at the base, which is embedded in the cell membrane, a filament or rod, which is the main corkscrew outside the cell, and a hook to connect the body and the filament. The flagellum also has its own export apparatus, as it can self-assemble. The hook has a set length that it grows to, whereas the filament is more variable. The set length is believed to be governed by the C ring, which measures the hook before exporting it. To facilitate self-assembly, the center of the flagellum is hollow, from the base all the way to the tip of the filament.
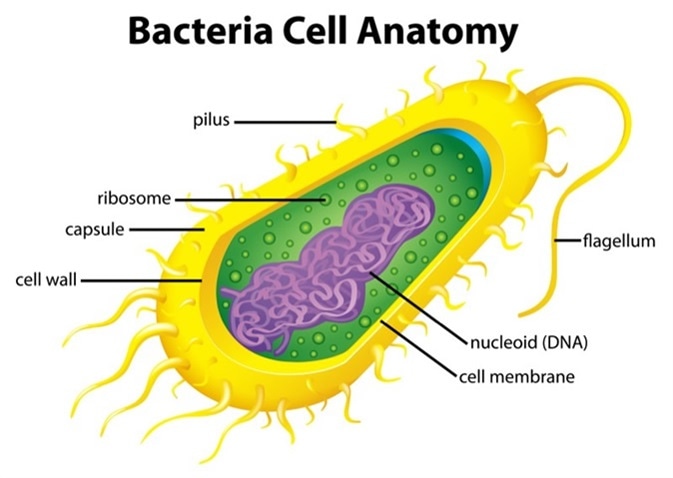
Illustration of the bacteria cell structure. Image Credit: BlueRingMedia / Shutterstock
The structure of the flagellum differs depending on if it is in prokaryotes or eukaryotes. In eukaryotes, the flagellum beats in a whip-like fashion, whereas in prokaryotes the flagellum is an unmoving cork-like entity, relying on the motor at its base for torque.
The structure of the flagellum is complex. At the base, the rotor sits inside the cell membrane. MotA and MotB proteins form rings located within and above the inner membrane, to combine the energy from proton movement across the membrane with torque generation. MotA and MotB are the static, nonrotating portion of the structure.
MS, P and L rings are protein rings that allow for the rotation of the main extracellular corkscrew rod through the membranes. The MS ring anchors the flagella to the cytoplasmic membrane, and it is here that the export system is likely located. The P ring facilitates rotation within the peptidoglycan, and the L ring within the outer membranes. In Gram positive bacteria, the P and L ring are lacking. The C ring is located intracellularly and regulates the direction of rotation. The direction of rotation is controlled by the chemotactic system, which the C ring responds to.
Role
The flagellum is mainly an organelle for movement. However, it can also participate in the formation of biofilms, export of proteins, and adhesion. Adhesion is important for many bacterial life cycles, and they have several mechanisms, such as fimbriae, pili, and other proteins to assist in this. The flagella and adhesive structures are typically not simultaneously expressed, but rather the bacteria switches from a moving to a stationary form.
Bacterial flagella can have an important role from a human perspective too. Flagella are usually needed for infection, and because of this they have pathogen-associated molecular patterns (PAMPs). PAMPs can be recognized by the human immune system through toll-like receptor 5 (TLR5). The N and C-termini of flagellin, which makes up the main rod, are highly conserved and are what TLR5 binds to. There are also polymeric flagellin forms, which do not bind TLR5.
Unlike the terminals, the central region of flagellin has higher sequence variation. This exposed region decides differences in adhesive function of flagellins from different strains or species of bacteria. Pseudomonas aeruginosa can cause urinary tract infections (UTIs) and other conditions, such as pneumonia. Its binding appears to be facilitated by flagellin (FliC) and/or FliD, proteins which cap the ends of the rod.
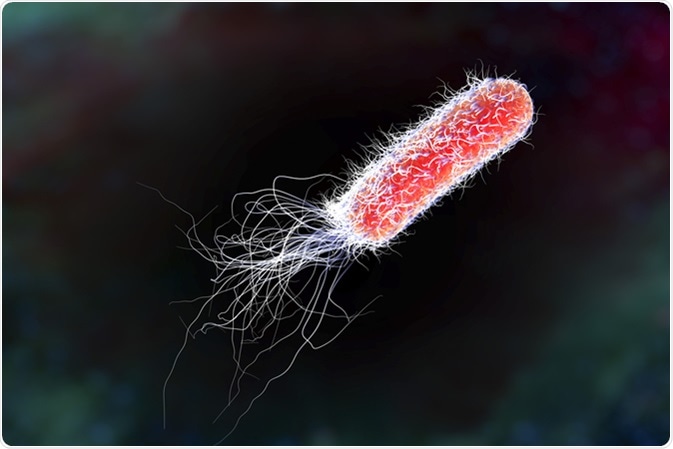
Bacterium Pseudomonas aeruginosa - antibiotic-resistant nosocomial bacterium. Illustration shows polar location of flagella and presence of pili on the bacterial surface. Image Credit: Kateryna Kon / Shutterstock
The role of flagella in producing biofilms is also important from a human perspective, due to dangerous biofilm formation on catheters and equipment. Studies on Escherichia coli have found that non-flagellated bacteria (i.e. lacking the main rod, without flagellin and FliD) showed reduced adhesion and thereby biofilm formation ability. In contrast, flagellated bacteria that were not able to move (i.e. lacking the motor, without MotB) were able to colonize as well as wild type strains.
Similarly, in Salmonella enterica serovar Typhi, flagella have a significant role in biofilm formation. Non-flagellated forms lacking flagellin were unable to form a biofilm. However, the flagellated, but not capable of movement, lacking MotA, mutant was able to form a biofilm on equal level as wild type strains. Here, flagellin in particular is crucial for biofilm formation.
Further Reading
Last Updated: Oct 29, 2018