The world is currently undergoing a significant transformation in biomarker science. Biomarker signatures are no longer academic luxury or exploratory afterthoughts; they have matured into strategic assets – instruments that actively de-risk decision making throughout the drug development process.
From preclinical discovery to late-phase clinical trials, the correct signature can accelerate go/no-go decisions, improve patient classification, and confirm mechanisms of action in real time.
This transition has been fueled by two converging forces: the growth of high-dimensional, high-throughput platforms, ranging from proteomics and transcriptomics to multiplexed imaging and single-cell technologies, and the incorporation of AI and sophisticated analytics into translational processes.
Together, they've made it possible to uncover signatures that are not only biologically rich but also actionable, predictive, and modular.
Despite the increasing availability of omics technologies and analytical platforms, many teams still struggle with a seemingly basic question: what type of signature is required?
Start with the clinical question, not the technology
Too frequently, the platform comes before the issue. A client might say, “We want to do proteomics on this study,” without explaining why. What choice will be made based on this information?
Is the goal to identify patients who are more likely to respond? Is there a requirement to demonstrate targeted engagement? Is there a need to explain variation in response or illness progression? Each objective has unique constraints for time, sample type, and resolution.
Each strategy requires compromise: RNA provides pathway insight but is less direct in its function; proteins are closer to the phenotype but can be limited by available detection reagents; and cell-based assays provide depth but are often logistically complex.
There is also a tendency to have “one marker to rule them all.” However, single-analyte indicators seldom function in isolation. Biology is noisy, and redundancy is a benefit, not a disadvantage. That is why well-designed signatures are composite by definition, integrating several weak signals into a robust, understandable readout.
So, how to pick the right set of markers?
- Begin with biological plausibility: Incorporate markers that are mechanistically sensible
- Ensure technical feasibility: Are these indicators consistently quantifiable in the selected sample matrix, in a clinical context, and at the required volume and stability?
- Finally, ask if they scale: Can these indicators be validated across cohorts, regions, and analytical platforms?
The true art is in selecting markers that are biologically relevant, clinically accessible, and analytically stable.
Platform convergence: When technologies derisk each other
In translational science, there is a seductive idea that a single tool, platform, or dataset will tell all that is needed to know. But biology rarely cooperates at that level of simplicity. The most robust biomarker signatures do not arise from a single source of truth; rather, they emerge through convergence.
Platform convergence is the idea that multiple technologies, when used together, can eliminate ambiguity, compensate for each other’s flaws, and boost confidence in a biological signal. When used wisely, technologies do not merely complement one another; they also reduce risk.
Assume a transcriptomic panel identifies a cluster of genes upregulated in responders to immunotherapy. That is intriguing, but it is just theoretical. If the same patients also exhibit:
- an elevation in serum inflammatory proteins
- alterations in T cell subsets as determined by flow cytometry, and
- activation markers observable through multiplexed tissue imaging
And then it is no longer simply a theory. Users will view a moving mechanism with three distinct modes. Each platform builds trust in the others, reducing the risk of overinterpreting noise. This is how multi-omic signatures gain traction: not by being clever, but by being coherent.
Strengths and weaknesses: Why no single platform is enough
Source: Synexa Life Sciences
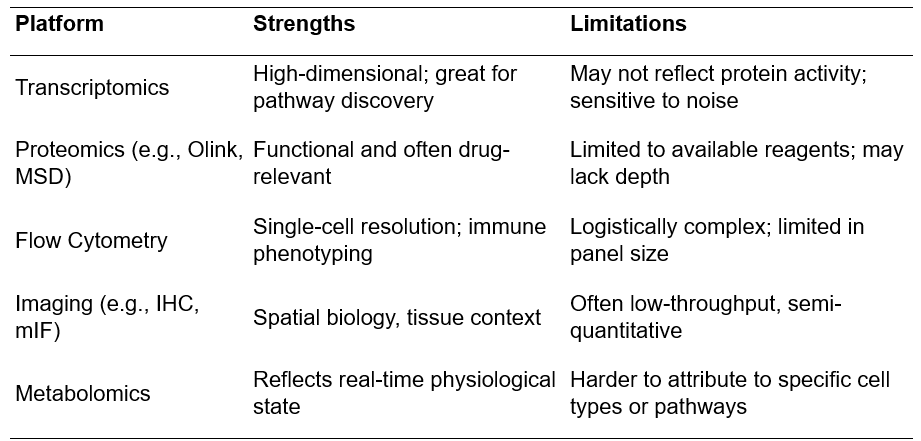
Platform design should mirror biological redundancy
Biology is inherently redundant. Cytokine signaling, for example, often involves multiple molecules and feedback loops. So, why expect a single biomarker, or even a single technology, to capture such complexity?
A resilient signature is similar to this biological architecture: it is layered, versatile, and can generate a signal under varying conditions and across different sample qualities.
This does not imply more noise; rather, it indicates deliberate overlap, in which many markers or modalities address the same biological event from various perspectives. If IL-6 levels rise in a serum proteomics panel, look for changes in STAT3 pathway activity in transcriptomics or increased myeloid populations in flow cytometry. When these signals align, confidence rises. When they deviate, they encourage more inquiry, occasionally revealing novel biology.
A redundant design also helps to future-proof a signature. Clinical studies may be messy: samples get lost, locations change, and reagents shift. A signature based on a single fragile signal is brittle.
However, one created with overlapping layers can still provide insight even if some of the data is absent or noisy. This is especially important in trials involving several regions, suppliers, or extensive durations.
The redundant design provides multiple entry points for later assay simplification. During early development, users may start with a high-dimensional, multi-platform signature and iteratively refine it down to a handful of proteins or transcripts that still reflect the original biology, since they've mapped the redundancy and know what works.
Examples of redundant design in practice:
- Immune gene expression patterns, such as interferon-γ-driven profiles, are widely used in checkpoint blockade studies to reinforce flow-based T cell activation and serum CXCL9/10 levels
- In fibrosis investigations, protein-level indicators of extracellular matrix remodelling (e.g., MMPs, TIMPs) are cross-validated with transcriptome signatures from FFPE tissue and histological grading, establishing a relationship between molecular change and tissue outcome
- CAR-T programs can use cytokine surges (Olink/MSD), immune cell dynamics (flow), and transcriptome stress signatures (Nanostring or bulk RNA-seq) to create a sophisticated safety and response monitoring system
Of course, not every researcher can afford all modalities at the outset. This is when tiered platform strategies come in. A realistic method may look like this:
- Discovery: Employ high-dimensional platforms (such as RNAseq, Olink Explore, scRNAseq) on a limited subset
- Refinement: Determine key features and narrow down to targeted panels (for example, flow, Olink Target 96)
- Validation: Implement refined, cost-effective assays across the wider trial population
This method not only saves money but also protects scientific integrity by ensuring that every stage of biomarker development is supported by orthogonal evidence.
AI, complexity, and the clinically grounded signature
The world is now approaching a period of biomarker research in which interpretation, rather than facts, is the limiting element. High-throughput technologies typically generate datasets including tens of thousands of characteristics from dozens or hundreds of samples. AI and machine learning have proven important in navigating this complexity and identifying candidate signatures.
However, while these techniques can speed up discovery, they do not replace biological intuition. And they surely do not address the difficulty of clinical translation. AI can identify patterns, but it cannot tell what is important.
The appeal of AI lies in its ability to uncover subtle, nonlinear patterns that traditional statistical methods would miss. Random forests, LASSO regression, and support vector machines are all algorithms capable of identifying multidimensional biomarker combinations that correlate with response, toxicity, or disease progression.
However, there is one important caveat: models are only as good as the input data, training logic, and interpretability. No matter how sophisticated the algorithm, a model based on incorrectly normalized NPX values or inconsistently gated flow data will produce inaccurate results.
Even more important is clinical significance. A signature consisting of 50 transcriptome characteristics that require frozen biopsies and a GPU cluster for analysis may perform admirably in a discovery dataset, but it fails miserably in a real-world trial setting.
Interpretable, actionable, and portable
That is why the next generation of biomarker signatures must reach a higher standard.
They should be:
- Interpretable: It is essential for clinicians and regulators to comprehend the significance of the signature
- Actionable: The signature should facilitate a decision - whether to stratify a patient, adjust a dosage, or escalate care
- Portable: The assay must be practical under clinical trial conditions (for instance, in FFPE, serum, plasma; requiring a low sample volume; applicable across various geographies and laboratory vendors)
This is where AI and subject expertise come into play: human-guided feature selection paired with automated learning can yield simpler, more robust signatures.
A five-protein panel developed from an Olink research may eventually surpass a 200-gene expression model if it is more repeatable, easier to evaluate, and better matched with clinical objectives.
The operational litmus test: Is your signature deployable?
In the years of assisting biomarker development across several therapeutic areas and trial phases, Synexa has seen how even the most biologically convincing signatures can fail if operational practicality isn't considered from the start. This is not a failure of science; rather, it is a mismatch of aim and infrastructure.
The idea is to plan for the practical limitations of clinical research. That requires looking beyond the discovery dataset to verify that the signature is applicable across geographies, sample types, and clinical time scales.
Some guiding questions:
- Is the sample type (e.g., serum, CSF, FFPE) readily available in the study protocol?
- Can the assay be scaled to large, multi-site trials?
- Is there redundancy in the signature in case of missing data?
- Can it be translated into a regulatory-grade assay later?
These are not small matters; they frequently represent the distinction between exploratory data and a validated biomarker strategy.
The signature is only as good as the system that supports it
An intelligent biomarker signature is not only an output; it is a component of a system. It must be traceable, reproducible, and survive GCP and regulatory examination. That is why integrating AI and high-throughput data must be built into clinical trial processes rather than operating alongside them.
Synexa believes that companies that embrace biomarker strategy as a fundamental development discipline rather than a downstream deliverable will have the greatest success in translational medicine. Synexa sees biomarker signatures as active design tools that shape trials, explain decisions, and eventually drive improved patient outcomes.
Synexa collaborates closely with its partners to anticipate these key elements early on, ensuring that signatures are not only physiologically relevant but also technically and logistically feasible. Whether supporting early-phase mechanism-of-action research or a worldwide Phase 3 trial, Synexa emphasizes that a successful signature can withstand real-world interactions while still delivering.
About Synexa Life Sciences
Synexa Life Sciences is a biomarker and bioanalytical lab CRO, specializing in the development, validation and delivery of a wide range of complex and custom-designed assays.
With a team of over 200 staff across three global laboratory locations; Manchester, Turku (Finland) and Cape Town, we provide innovative solutions to support our customers to achieve their clinical milestones.
Our main areas of expertise include biomarker identification and development, large and small molecule clinical bioanalysis, (soluble) biomarker analysis (utilizing MSD, LC-MS/MS, ELISA, RIA, fluorescence and luminescence-based technologies), cell biology (including flow cytometry, ELISpot and Fluorospot) and genomic services to support clinical trials and translational studies.
We pride ourselves on our deep scientific expertise and ability to tackle complex problems, translating them into robust and reliable assays to support clinical trial sample analysis.
Since 2019, Synexa has been backed by Gilde Healthcare, a specialized healthcare investor. Synexa, improving the quality of human health through innovative biomarker and bioanalytical solutions.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.net, which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.