Identification of microbes joins together the discipline of microbiology with the study of infectious diseases. Methods of reliable and accurate microbial identification are valuable to a wide range of scientific fields, some of which pertain to life-threatening health situations.
 Image Credit: Parilov / Shutterstock.com
Image Credit: Parilov / Shutterstock.com
The analytical techniques used to carry out microbial identification have, therefore, become essential to a number of applications. A range of platforms have been developed to carry out microbial identification. Here, we cover the various biochemical tests that are used for this task.
Identifying bacteria via analyzing their enzymatic profile
Biochemical reactions can reveal the vital information necessary for accurately identifying the genera of various bacteria within a sample. By their nature, bacteria produce large volumes of enzymes, and it is these enzymes that allow for their identification via biochemical methods. The type of enzymes produced by a bacterium can usually be used to classify its species given that bacteria have distinct enzymatic profiles.
Each species of bacteria has specific metabolic needs and relies on different enzymes to fuel those unique needs. The presence of catalase, gelatinase, oxidase, urease, for example, can be used to identify the species of bacteria.
Biochemical reactions used in biochemical tests depend on the presence of such bacteria. Such biochemical tests have been designed to measure the levels of bacterial enzymes which can be interpreted to accurately identify the species of bacteria they have been produced by.
These biochemical tests have become commonplace in the fields of healthcare, where they are relied upon for assisting with disease diagnosis; epidemiology, for the tracking and tracing of disease outbreaks; pharmaceutical, for the analysis of environmental microbes that may have health implications; and forensic science, where microbial investigation can assist in the investigation of bioterrorism threats.
Below, we discuss the various biochemical tests that have been established for these various applications.
Traditional biochemical tests for microbial identification
Current tests for microbial identification can be split into traditional methods and modern methods. Simple biochemical tests such as catalase testing, oxidase testing, and substrate utilization tests fit under the category of traditional tests, alongside staining and microscopy methods such as gram staining, endospore staining, and Ziehl-Neelsen staining.
Catalase activity is specific to certain bacterial strains such as Staphylococci, Micrococci, E. coli and the other Enterobacteriacaea, and Salmonella spp. Others are known not to cause this activity, such as Streptococcus and Enterococcus bacteria. Catalase testing involves inducing catalase activity by adding hydrogen peroxide to a bacterial scraping placed on a microscope slide. Bubbles that appear on the slide indicates catalase activity and suggests the presence of catalase-positive bacteria.
Bacteria with cytochrome c oxidase activity (CCO) can be identified with oxidase testing. When present, the COO enzyme, which forms part of the bacteria’s electron transport chain, oxidizes tetramethyl-p-phenylenediamine (used as a reagent). This oxidation reaction causes the reagent to turn purple in color, therefore, the presence of this enzyme can be confirmed visually, as when the enzyme is not present, the reagent does not change color.
The limitation of oxidation tests is that they are vulnerable to inaccurate results given that while oxidase-positive bacteria are aerobic, some are capable of anaerobic respiration also. Additionally, false-negative results can be induced if the bacterial species under investigation possesses an oxidase that does not react with the reagent.
Finally, a range of substrate utilization tests is commercially available for microbial identification. These tests involve the addition of unknown bacterial species to a panel of substrates. Scientists can then identify the bacteria based on the color changes induced in the separate panels, demonstrating unique patterns associated with specific bacterial species. These tests are often conducted alongside catalase and/or oxidase tests in order to increase accuracy and reliability.
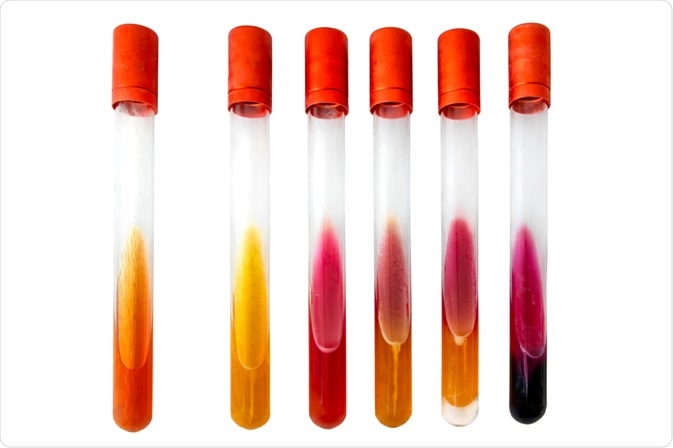 Image Credit: eczserapyilmaz / Shutterstock.com
Image Credit: eczserapyilmaz / Shutterstock.com
Modern biochemical tests for microbial identification
In addition to traditional methods, recent years have seen new, modern methods emerge that have been adapted for microbial identification. Polymerase chain reaction (PCR), and immunological tests such as ELISA are among these modern methods. Newer biochemical tests that have emerged include fatty acid profiling and metabolic/chemo profiling.
Fatty acids are vital to the construction of the cell membranes of bacterial cells. As with enzymes, fatty acid profiles are unique to specific bacterial species. Therefore, fatty acids obtained from a species of unknown bacteria can be used to identify it. The methods of gas chromatography and mass spectrometry are most commonly used to conduct fatty acid profiling.
Finally, metabolic/chemo profiling is used to detect the unique secondary metabolic profiles of bacteria. All microbes produce metabolites, primary metabolites such as ATP that are associated with basic functioning, and secondary metabolites, such as antibiotics and immunosuppressive compounds that are more specific to the species of bacteria.
These secondary profiles can be measured by methods such as high-performance liquid chromatography (HPLC) and mass spectrometry to identify the species of bacteria that they are associated with.
Sources
Aslanzadeh, J., n.d. Biochemical Profile-Based Microbial Identification Systems. Advanced Techniques in Diagnostic Microbiology, pp.84-116. https://link.springer.com/chapter/10.1007/0-387-32892-0_6
Janda, J. and Abbott, S., 2002. Bacterial Identification for Publication: When Is Enough Enough?. Journal of Clinical Microbiology, 40(6), pp.1887-1891. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC130684/
Sandle, T., 2016. Microbial identification. Pharmaceutical Microbiology, pp.103-113. https://www.sciencedirect.com/science/article/pii/B9780081000229000098
Further Reading
Last Updated: Jan 14, 2021