Although DNA was first isolated in 1869, it was not until more than a century later that the first genomes were sequenced, making genomics a relatively new discipline. Genomics, as we now know it, truly began in the 1970s, although there were several significant milestones that shaped the field in the preceding century.
Early history (1971 – 1910): Discovery of DNA
Friedrich Miescher published a paper in 1871 that identified the presence of nuclein and associated proteins in the nucleus of cells. This is what we now refer to as DNA and forms the base of the genomics field.
In 1904, Walter Sutton and Theodor Boveri found that chromosomes occur in matched pairs, with one inherited from each of the mother and the father. They called this the Chromosome Theory of Heredity.
Albert Kossel was awarded the Nobel Prize in Physiology and Medicine in 1910 for the discovery of the five nucleotide bases, which are:
- Adenine (A)
- Cytosine (C)
- Guanine (G)
- Thymine (T)
- Uracil (U)
1950-1968: Development of knowledge about DNA
Erwin Chargaff developed the understanding of the pairing of the adenosine, cytosine, guanine and thymine nucleotide bases in 1950. He discovered that there were always equal amounts of thymine and adenine or cytosine and guanine in DNA samples. As a result, he deduced that adenosine forms chromosome pairs with thymine and cytosine forms chromosome pairs with guanine.
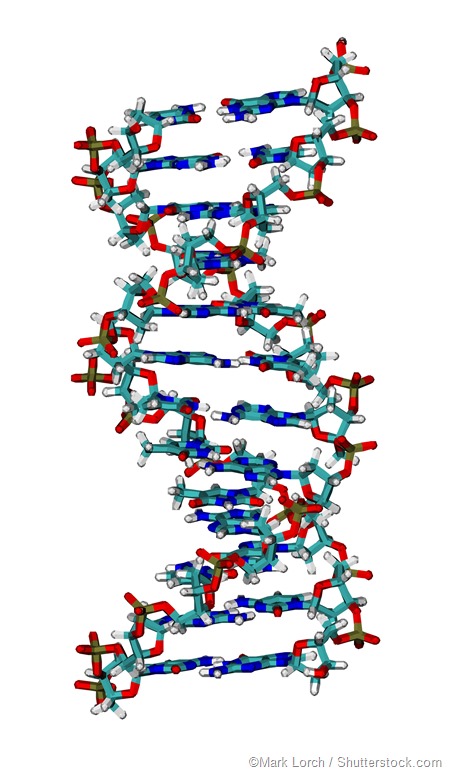
In 1952, Alfred Hershey and Martha Chase demonstrated with a series of experiments that DNA, not protein, is responsible for carrying genetic traits that may be inherited. James Watson and Francis Crick discovered the double helix structure of DNA in 1953.
In 1961, Marshall Nirenberg and Har Gobind Khorana led the research team that identified what is now known as the “codon” of DNA. This is the block of three letters in DNA that correlates to a specific amino acid. It is when amino acids combine, during translation, that particular proteins are made. They won the Nobel Prize for Physiology and Medicine in 1968 for cracking the genetic code, alongside Robert Holley, who sequenced the first transfer RNA (tRNA) molecule.
1977 – Present: Sequencing of DNA
In 1977, Frederick Sanger developed a sequencing technique for DNA to sequence the first complete genome, called phiX174 virus, which opened the doorway to the possibility in the field of genomics. In 1980, he won the Nobel Prize with Wally Gilbert and Paul Berg for pioneering DNA sequencing methods.
In 1983, James Gusella identified the location of the gene associated with causing Huntington’s disease. In the same year, Dr. Kary Mullis developed a technique used to amplify DNA, known as the polymerase chain reaction (PCR). In 1985, Alec Jeffreys developed a method to profile DNA by counting the number of repeating DNA sequences at specific regions of the genome.
In 1992, techniques for testing embryos in the womb for genetic diseases were developed. This enabled expectant parents to find out if their unborn child was likely to be affected by cystic fibrosis of hemophilia.
In 1995, the first bacterium genome sequence, Haemophilus influenza, was completed. One year later, the genome of yeast, Saccharomyces cerevisiae, was completed. Dolly the Sheep, who was the first cloned animal was also born at this time. In 1998, John Sulston and Bob Waterston published the genome of the nematode worm, Caenorhabditis elegans.
The Human Genome Project was launched in 1990 with the aim to sequence all 3 billion letters of a human genome. Chromosome 22 was the first chromosome to be sequenced as a part of this project in 1999. The project was completed in 2003, and confirmed that humans have 20,000-25,000 genes.
The International Mouse Genome Sequencing Consortium sequenced the first full genome sequence of a mammal in 2002. The genome of the mouse is smaller that the human genome, the largely similar to the human genome. The International HapMap Project was also launched in this year to create a catalog of common variations in the human genome.
In 2007, there was a breakthrough in the technology used to sequence DNA, which led to a 70-fold increase in the output of DNA sequencing in one year. This led to the launch of the 1000 Genes Project in 2008, which aimed to sequence the genomes of a large population group of 2500. The results of this project were published in 2010.
References
- http://www.yourgenome.org/facts/timeline-history-of-genomics
- http://assets.cambridge.org/97805218/00228/excerpt/9780521800228_excerpt.pdf
- https://www.genomic.org.uk/
- https://www.ndsu.edu/pubweb/~mcclean/plsc411/History-of-Genetics-and-Genomics-narrative-and-overheads.pdf
Further Reading
Last Updated: Jul 19, 2023