Metabolomics is a relatively new field of research and the youngest of the triad of systems biology that it is part of, alongside genomics and proteomics. The term “metabolome” was first introduced in 1998, and even up to 2010 metabolomics was regarded as an emerging field. This article will describe a brief overview of the history of metabolomics, from early conceptual ideas in ancient history, the development of technology, and breakthrough findings in fields in more recent times.
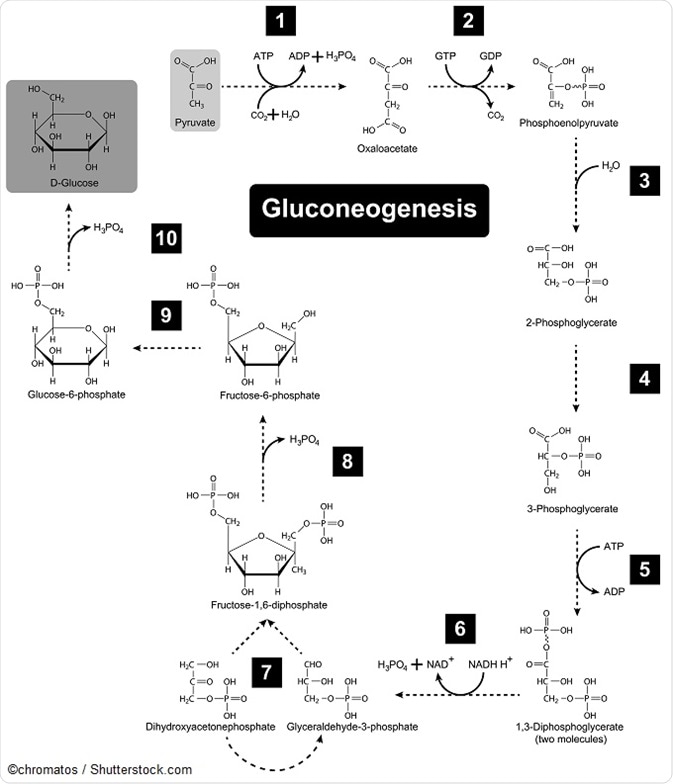
Image showing Gluconeogenesis metabolic pathway diagram.
Ancient history
The basic concept of biological fluids portraying information about the health of an organism has existed throughout history in various cultures.
For example, ancient Chinese doctors relied on ants to evaluate the concentration of glucose in the urine, which was a sign indicative of what is now known as diabetes mellitus. The color, taste and smell of urine were also linked to certain medical conditions in the Middle Ages, which are now known to be linked to the metabolomics of the organism.
In the thirteenth century, Arab physician, Ibn al-Nafis, noted the concept of metabolism with a statement about the body being in a continuous state of change due to dissolution and nourishment.
Development of technology
In the late 1940s, Roger Williams suggested that each individual may have a “metabolic profile” that was reflected in their biological fluids. He demonstrated his theory with paper chromatography to establish patterns in the metabolic components of bodily fluids, such as urine and saliva, of patients with schizophrenia.
Several decades later, the advancements in technology enable quantitative measurements of metabolites. In 1971, Horning and his team first coined the term “metabolic profile” when they showed that compounds present in urine or extracts from tissues could be measured with gas chromatography-mass spectrometry (GC-MS).
At the same time, nuclear magnetic resonance (NMR) technology began to be used to detect metabolites in raw biological samples. NMR was originally discovered in the in the 1940s, but began to be utilized in the study of metabolomics in the 1970s. With time, the sensitivity of this technology increased as a result of the use of stronger magnetic fields and magic angle spinning. In 1984, Professor Jeremy Nicholson demonstrated the possible use of NMR spectroscopy in the diagnosis of diabetes mellitus.
Analysis breakthroughs
The METLIN database, which categorized over 10,000 metabolites and their mass spectrometry data, was established in 2005. It now contains data of more than 240,000 metabolites.
The Human Metabolome Project finished the first draft of the human metabolome in 2007. The projects included a database of approximately 2500 metabolites, as well as 1200 drugs and 3500 nutritional components. Since this project, there have been several similar undertakings with plants species, such as for Medicago truncatula and Arabidopsis thaliana.
XCMS Institute: METLIN
Future of metabolomics
Significant progress has already been made, and metabolome profiling in real-time was demonstrated for the first time in history in 2015.
However, it is currently understood that continued progress in the field of metabolomics relies on improved methods and technology for mass spectrometry. At present, technical challenges in this area are slowing down the rate of progress and a breakthrough in this area is needed.
References
- http://www.biotechniques.com/BiotechniquesJournal/specialissues/2009/April/What-is-metabolomics-all-about/biotechniques-140692.html?pageNum=2
- https://www.ncbi.nlm.nih.gov/pubmed/23630115
- https://omics.pnl.gov/metabolomics-and-lipidomics
Further Reading
Last Updated: Feb 26, 2019