Over the past decade, culture-independent microbial investigation techniques have led to an improved understanding of the complete set of microbes that live on and within the human body. This set of human-associated microbes is collectively known as the microbiome. The human body is host to trillions of microbes and the gene set that makes up the microbiome is 360 times more abundant than that comprising the human genome.
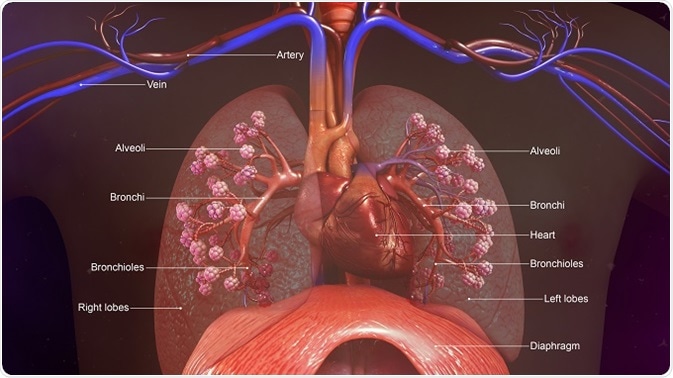
Historically, the lungs have been considered sterile in health, but culture-independent techniques have now shown that the lower respiratory tract is home to a diverse microbial community: the lung microbiome. Increasingly, evidence has emerged that supports the concept that a distinct microbiota is present in the lower respiratory tract in both healthy and disease states.
Research into the lung microbiome is rapidly growing and revealing important discoveries about the microbiome’s association with respiratory disease. It is hoped that continued study of this field will provide new insights into the pathogenesis of lung diseases as well as the role the lung microbiota play in respiratory health. Analysis of the lung microbiome is mainly performed using bronchoalveolar lavage (BAL), although spontaneously expectorated sputum is used to analyze cases of cystic fibrosis.
Details are given below regarding some of what is known so far about the lung microbiome.
The healthy lung
It is generally presumed that the fetal lungs are sterile and that infants acquire their lung microbiota after birth. Immediately after birth, the mother’s microbes rapidly populate the mucosal surfaces of the infant. At first, the colonization is uniform across the infant’s body sites, but the microbiota soon differentiates into site-specific communities in the days or weeks to come.
Many studies have characterized the microbiome of the adult lung using BAL samples taken from healthy human adults. These have shown that the most common phyla include Bacteroides, Firmicutes and Proteobacteria, with the most common genera being Streptococcus, Pseudomonas, Veillonella and Prevotella.
Lung Microbiome Project: Advancing Potential Treatments | UPMC
Cystic fibrosis
Very little is known about the lung microbiota in people with cystic fibrosis (CF) or how the microbial community of the airways contributes to lung disease in these individuals.
Molecular techniques have revealed a complexity in the airway microbiota among CF patients that had not previously been recognized. An early 2004 study, published by GB Rogers et al. in the Journal of Clinical Microbiology, showed that CF patients with bacterial infection of the airways had sputum samples containing vast amounts of various different bacterial species. Studies since then have also shown that the colonization of airways in CF is polymicrobial and includes many bacterial species that had not previously been recognized using culture-dependent analysis.
Studies of the lung microbiome in CF patients have also demonstrated the presence of previously unrecognized bacterial species that are potentially harmful and contributors to inflammation and destruction of the airways in CF.
Asthma and allergies
One important study, published in PLoS ONE in 2010 by Markus Hilty and colleagues, compared the lung microbiome of asthma patients with asthma-free controls. The authors reported an increase in Proteobacteriae and a decrease in Bacteroidetes among the asthma versus control patients. The bacterial species driving the relative increase in Proteobacteriae were Moraxella, Neisseria and Haemophilus.
Following on from this, YJ Huang et al. published a study in The Journal of Allergy and Clinical Immunology (2011), which investigated the differences in lung microbiota between asthmatics and controls. They reported a greater bacterial diversity and burden among the asthmatic versus control participants, as well as a greater relative abundance of Proteobacteriae. The presence of multiple species also correlated positively with the degree of bronchial hyper-responsiveness that asthma patients had.
Furthermore, in a study carried out by Hans Bigaard (University of Copenhagen), infants with lungs that contained pathogenic bacteria after they were born, were at an increased risk of developing asthma, compared with infants who were not born with the harmful bacteria in their lungs. In another study, Homer Boushey (University of California) found that adults with asthma had more bacteria in their lungs than non-asthmatic individuals. The severity of asthma also increased in cases with more diverse bacterial colonization.
Further Reading
Last Updated: Jul 19, 2023