Copying the Recipe of Life
DNA, or deoxyribonucleic acid, is the biological molecule which contains the information needed to create a living organism. As the cell divides to become two, the DNA has to be copied so that both cells contain the necessary genetic information. The synthesis, or making of new DNA strands in living cells is referred to as “DNA replication”.
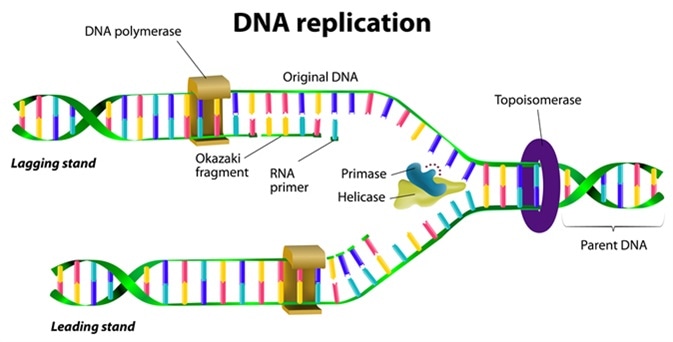
DNA replication. Image Credit: Designua / Shutterstock
Making a New Strand of DNA
DNA is found as a double helix, where two strands of DNA are bound together into two helices. The process of DNA replication begins when the two strands of DNA separate. An enzyme called helicase unwinds and separates the bonds between the two DNA strands, and both these separated strands act as templates from which new DNA is made.
DNA replication - 3D
DNA polymerases are a group of enzymes which make new DNA. But, for this enzyme to work, it needs a primer – a short nucleotide sequence which is attached to one of the singe DNA strands. During DNA replication, the primer is usually a short ribonucleic acid (RNA) sequence, which is later degraded and replaced by DNA.
The primer provides a 3’ hydroxyl group onto which the DNA polymerase adds the precursors of DNA, the nucleotides. When nucleotides are added to the 3’ end of the primer or the new DNA strand, a bond is formed between the 3’ hydroxyl group of the primer/new DNA and the 5’ phosphate group of the nucleotide.
There are four types of DNA nucleotides, which each have different nitrogenous bases; adenine (A), cytosine (C), guanine (G) and thymine (T). These are always found in pairs, A-T and C-G. This is referred to as “Watson-Crick base pairing”, and thus if the template has an “A” nucleotide, then a “T” nucleotide will be added to the growing DNA strand. If it is a “G” nucleotide, then a “C” nucleotide will be added to the growing strand.
Directionality of DNA, and How it Affects DNA Synthesis
DNA has directionality, with one strand going from 5’ to 3’ and the other going from 3’ to 5’. In the 5’-3’ strand, the 3’ end is exposed during the synthesis of new DNA. This means that DNA polymerase is able to make new DNA in the direction of the template DNA.
However, when it comes to the 3’-5’ stand, this would leave the 5’ end exposed; how does DNA polymerase deal with this? In the 3’-5’ strand, new DNA is made by the DNA polymerase making short 5’-3’ fragments, called Okazaki fragments. Then a different enzyme, DNA ligase, connects the Okazaki fragments together to form the new 3’-5’ DNA strand.
Proof-Reading
Sometimes, errors are made when adding nucleotides onto the template DNA strand. Some DNA polymerases have what is called “3’-5’ exonuclease activity”. The 3’-5’ exonuclease activity, working against the polymerase or synthesis activity, cuts away nucleotides which do not match the template. This provides a proof-reading so that the new DNA strand is as accurate as possible.
How is this Exploited in Molecular Biology?
Properties of DNA synthesis have been used in various molecular biology techniques; polymerase chain reaction (PCR) and DNA sequencing.
Polymerase Chain Reaction
PCR is a technique developed in the 1980s to amplify a specific part of DNA. DNA primers are designed to cover the region of interest. Heat is used to separate the two DNA strands, and the temperature is then lowered to allow the primers to attach to the DNA. A heat-stable DNA polymerase then makes new DNA strands matching the region of interest by adding nucleotides to the 3’ end of the primers.
Polymerase Chain Reaction
DNA Sequencing
DNA sequencing is working out the order of the bases (A, C, G, T) found in DNA. Sanger Sequencing is one of the early DNA sequencing techniques, where DNA synthesis is halted. This is made possible by removing the hydroxyl group from the 3’ end of nucleotides, therefore when these “dideoxy nucleotides” are incorporated into the DNA strand the DNA polymerase cannot add the next nucleotide. These dideoxy nucleotides are mixed with nucleotides, therefore fragments of varying lengths are created. By having dideoxy-A, dideoxy-C, dideoxy-G and dideoxy-T in separate reactions, the last nucleotide of the fragment can be determined. Once separated by size, the DNA sequence can then be determined by looking at what the last nucleotide of the fragments are according to size.
Although modern DNA sequencing methods are automated and less laborious than Sanger Sequencing, there are some which is based on Sanger Sequencing; for example, different fluorescent dyes can be added to each dideoxy-nucleotide and the differences detected by different fluorescence colors.
Further Reading
Last Updated: Feb 26, 2019