Mathematical modeling suggests that the rate at which a patient’s immune system clears slow-growing variants of methicillin-resistant Staphylococcus aureus (MRSA) bacteria is a key determinant of whether antibiotics can cure the infection. Tsuyoshi Mikkaichi and Alexander Hoffmann of the University of California, Los Angeles, and the MRSA Systems Immunobiology Group present this work in PLOS Computational Biology.
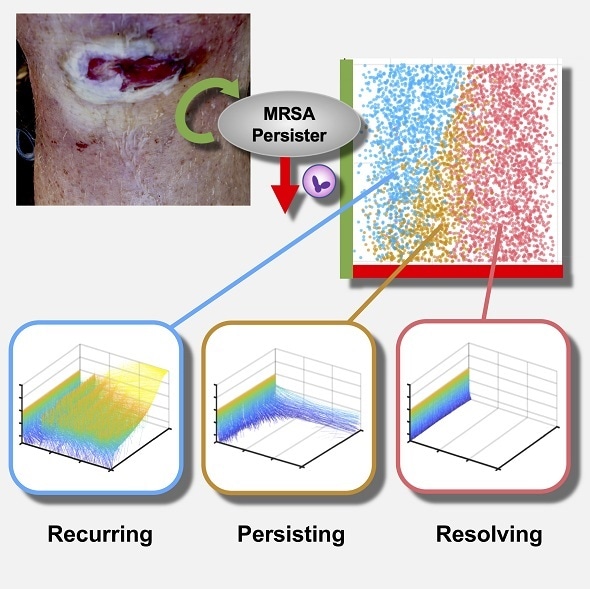
Credit: Mikkaichi et al.
MRSA infection can lead to a life-threatening condition known as persistent bacteremia, in which the bacteria are persistently present in the bloodstream. Researchers have proposed that when MRSA infects a person, it exists in two forms: normal bacteria and slow-growing variants that are less susceptible to antibiotics. It has been hypothesized that the rate at which normal bacteria switch to slow-growing variants influences whether an infection persists or can be cured.
To explore this question, Mikkaichi and colleagues constructed a mathematical model that simulates the dynamics of both normal and slow-growing bacterial populations during typical antibiotic treatments. The model effectively serves as a diverse group of virtual patients, some of whom are cured and others not, enabling the researchers to make precise predictions of why antibiotics fail for some.
The virtual analysis suggests that the rate at which a patient’s immune system clears slow-growing variants—not the rate at which they are produced—is a key determinant of whether drug treatment fails or succeeds. “Based on these findings, a drug that specifically kills the slow-growing variant may be the most effective treatment for persistent bacteremia,” Mikkaichi says.
The next step for this work is to improve understanding of interactions between slow-growing MRSA variants with the immune system, and why the immune system may be ineffective in killing these bacteria in patients with persistent infections.
These slow-growing variants may be hiding from the immune system by entering the patient’s tissue or immune cells and growing inside them.”
Tsuyoshi Mikkaichi, University of California, Los Angeles
Principal Investigator Alexander Hoffmann adds:
Training the immune system to recognize MRSA hiding in host cells could be an effective treatment strategy.”
Source:
Journal reference:
Mikkaichi, T. et al. (2019) Identifying determinants of persistent MRSA bacteremia using mathematical modeling. PLOS Computational Biology. doi.org/10.1371/journal.pcbi.1007087.