Protein biomarkers have tremendous potential for improving the diagnosis, prognosis, and treatment of neurological disorders such as Alzheimer's disease, Parkinson's disease, and multiple sclerosis.
Understanding the complicated processes of neurological diseases, on the other hand, requires more population-level research into the human genome and proteome.
The demand for highly sensitive and specific assays has further hampered the development of trustworthy biomarkers. Overcoming these barriers is critical to advancing precision medicine in neurology.
The pilot phase of the UK Biobank-Pharma Proteomics Project (UKB-PPP), a pioneering endeavor that combined large-scale proteome profiling using the Olink® Explore 3072 platform with genetic and clinical data from over 50,000 UK Biobank participants, marked a significant advancement.
This unique dataset has enabled groundbreaking discoveries at the interface of genomics and proteomics, significantly advancing our understanding of complex diseases and paving the way for more personalized medical interventions.
The UK Biobank's vastness and richness make it an unrivaled resource for academics, uncovering previously unachievable discoveries.
Following the success of the trial phase, in January 2025, it was announced that Olink® Explore HT (which covers over 5,400 proteins) would be used to profile all 500,000 UK Biobank members.
This extended dataset is expected to reveal considerably more about the complex relationships between proteins and disease, providing the framework for tailored medicines and improved diagnostic tools.
Neurology is one of the numerous therapeutic areas where the UKB-PPP has led to significant advancements.
With around 15,000 neurological disorders represented (Table) and approximately 50 related research papers published to date using UKB-PPP data, the discipline has already made substantial advances in biomarker discovery for early diagnosis, patient classification, and disease pathogenesis.
The following sections will discuss how the UKB-PPP has facilitated transformative research in neurological science.
Table. Number of cases of different neurological diseases in the UKB-PPP. Source: Olink®- Part of Thermo Fisher Scientific
| Disease / Disorder |
Number of cases (UKB-PPP) |
| Depression |
5,446 |
| Pain |
3,477 |
| Sleep disorders |
1,872 |
| Stroke |
893 |
| Epilepsy |
807 |
| Alzheimer’s disease & related dementias |
690 |
| Parkinson’s disease |
342 |
| Motor neuronal disease |
232 |
| Multiple sclerosis |
217 |
| Schizophrenia |
130 |
| Total |
∼15,000 |
Prediction of future dementia in healthy individuals
Early prediction and identification of neurodegenerative diseases offers the best chance for prevention and treatment. Large-scale population health research enabled by the UKB-PPP has improved dementia prediction in healthy people up to 10 years before diagnosis.1
Researchers at Shanghai Medical College used their access to UKB-PPP data to conduct a significant in silico study to uncover predictive risk indicators and signatures for all-cause dementia (ACD), Alzheimer's disease (AD), and vascular dementia (VaD).
After identifying 4-11 significant predictive analytes for those disorders, combining GFAP or GDF15 with demographic factors yielded reliable predictions of ACD, AD, and VaD.
When evaluating the time to diagnosis from a blood draw, integrating GFAP with demographic data provided an accurate prediction of future dementia over a 10-year period for ACD and AD. GFAP and NEFL levels began to change considerably at least 10 years before incident dementia was identified.
“Utilizing a data-driven proteomics strategy, we innovatively identified important plasma biomarkers for future dementia prediction from the largest prospective community-based cohort with long-term follow-up to date. These findings are poised to yield significant implications for screening people at high risk for dementia and for early intervention.”- Guo et al. 2025
Understanding the pathology of Parkinson’s disease with multi-cohort validation
Another study used Mendelian randomization to uncover predictive and causal proteins associated with Parkinson's disease (PD). UKB data identified 38 proteins associated with Parkinson's disease incidence over a 14.5-year follow-up period.2
Six of the top ten most significant proteins were completely replicated in a Parkinson's Progression Markers Initiative (PPMI) validation cohort. ITGAV, HNMT, and ITGAM all had significant relationships with incidence PD across three baseline-to-diagnosis intervals.
The top 16 related proteins and demographic characteristics predicted incident Parkinson's disease in subgroup analyses of up to 5 years and over 5 years, which were validated in the PPMI cohort.
“By characterizing the temporal evolution of sporadic PD, we address key gaps in understanding early-stage pathology, aiding biomarker and therapy development.” - Gan et al. 2025
The largest plasma proteomic screen of multiple sclerosis – support for known associations and novel biomarkers
Blood-based protein biomarkers are promising for the diagnosis, monitoring, and prognosis of multiple sclerosis (MS); yet, they are not widely used in MS research or care.
Markers of MS risk and disease severity were assessed in 407 prevalent MS cases from the UKB with well-documented clinical data.3 Seventy-two proteins were linked to MS, several of which had previously been discovered.
Granzyme A (GZMA), one of the downregulated proteins, was discovered as an MS biomarker for the first time.
Pathway analysis revealed enrichment of cytokines, cytokine receptors, and lysosomal processing proteins in MS, as well as a decrease in proteins involved in leukocyte migration, extracellular matrix interaction, actin cytoskeleton regulation, and cell-cell adhesion (Figure).
The data was directionally concordant with MS for approximately 83 % of overlapped proteins reported in a previous Olink MS investigation.4
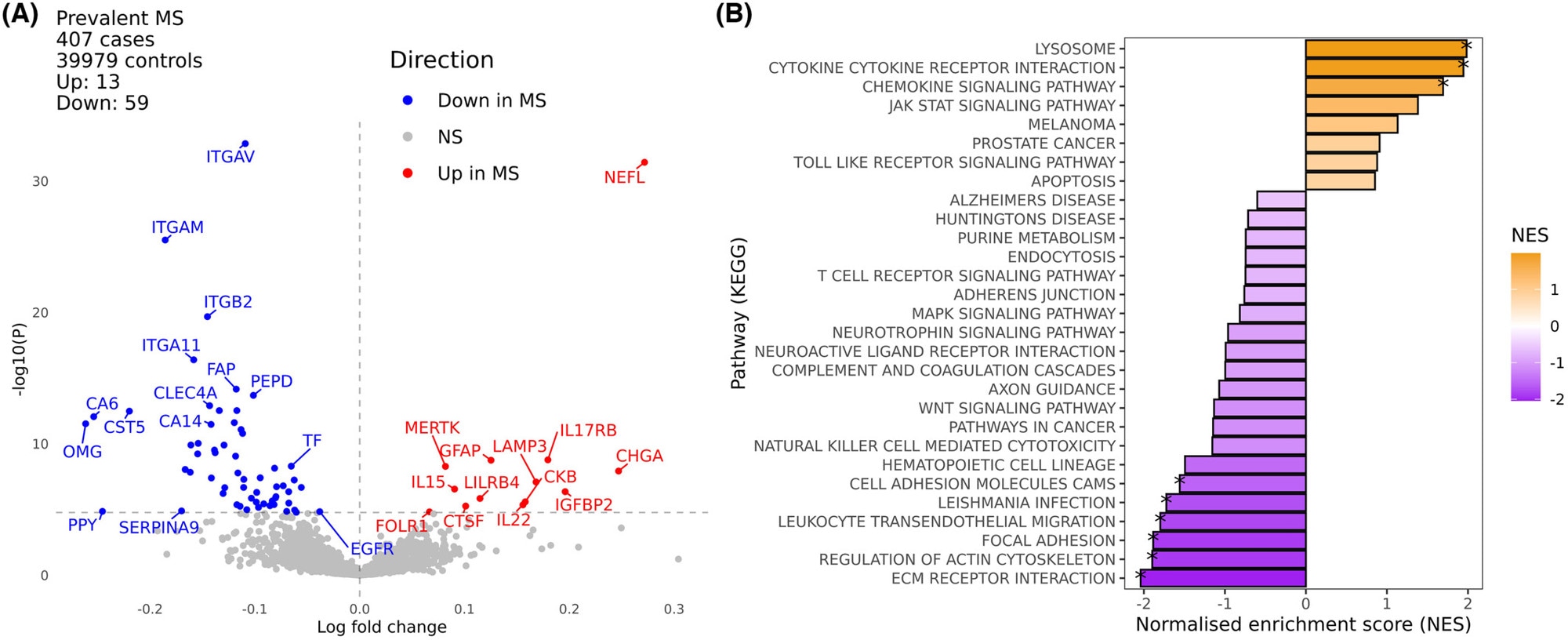
Plasma proteomic analysis of multiple sclerosis. (A) volcano plot of differences in plasma proteins in UK Biobank participants with (n = 407) and without (n = 39,979) MS at the time of sampling. (B) Gene set enrichment analysis of pathway-level differences in the plasma proteome between MS and healthy controls. Image Credit: Jacobs et al.3
“[The findings] demonstrate the power of biobank-scale datasets for discovering how the plasma proteome is altered in multiple sclerosis. Ultimately, this avenue of research could yield new drug targets, new insights into disease biology, and provide an adjunct to existing methods for individual-level prognosis in MS.” - Jacobs et al. 2024
References
- Guo, Y., et al. (2024). Plasma proteomic profiles predict future dementia in healthy adults. Nature Aging. DOI: 10.1038/s43587-023-00565-0. https://www.nature.com/articles/s43587-023-00565-0.
- Gan, Y.-H., et al. (2025). Large-scale proteomic analyses of incident Parkinson’s disease reveal new pathophysiological insights and potential biomarkers. Nature Aging. (online) DOI: 10.1038/s43587-025-00818-0. https://www.nature.com/articles/s43587-025-00818-0
- Jacobs, B.M., et al. (2024). Plasma proteomic profiles of UK Biobank participants with multiple sclerosis. Annals of Clinical and Translational Neurology, (online) 11(3), pp.698–709. DOI: 10.1002/acn3.51990. https://onlinelibrary.wiley.com/doi/10.1002/acn3.51990.
- Åkesson, J., et al. (2023). Proteomics reveal biomarkers for diagnosis, disease activity and long-term disability outcomes in multiple sclerosis. Nature Communications, (online) 14(1). DOI: 10.1038/s41467-023-42682-9. https://www.nature.com/articles/s41467-023-42682-9.
About Olink®- Part of Thermo Fisher Scientific
Olink’s mission is to accelerate proteomics together with the scientific community, to understand real-time biology and gain actionable insights into human health and disease. Our innovative solutions deliver highly sensitive and accurate protein quantification, giving scientists the power to investigate complex biological processes with precision.
One platform. Endless possibilities.
Explore up to 5,400 proteins with high specificity, transparent data, and the flexibility to answer any research question. Meet the next-generation proteomics platform trusted by the scientific community, from small academic research teams through to leading pharma companies.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.net, which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.