Lung cancer is the most common and deadly cancer, accounting for about 1.76 million deaths globally. Molecular and histological profiling of lung cancer is important to identify molecular and genetic targets for personalized medicine.
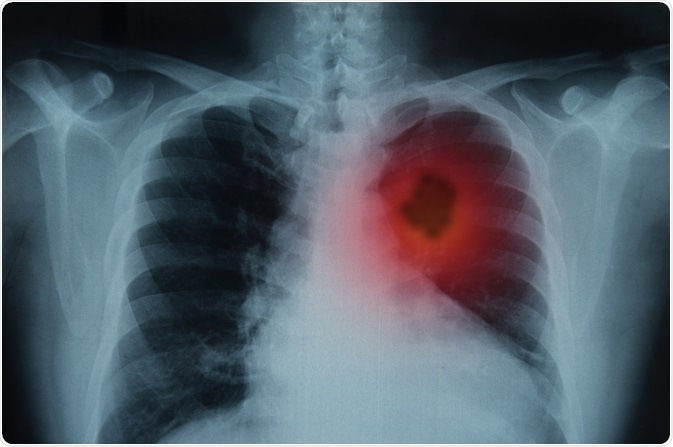
Image Credit: create jobs 51/Shutterstock.com
What is lung cancer?
According to the histological classification, lung cancer is broadly divided into two types: small cell lung carcinoma (SCLC) and non-small cell lung carcinoma (NSCLC). NSCLCs that account for about 85% of all lung cancers are further subdivided into three types: adenocarcinoma, squamous cell carcinoma, and large cell carcinoma.
Lung cancer is associated with a very poor prognosis because of its highly heterogeneous nature. Moreover, lung tumors belonging to the same histological subtype often show molecular and histological heterogeneity.
Therefore, molecular characterization of each subtype in terms of genetic mutations, DNA methylation, mRNA and microRNA expressions, and protein expression is important for understanding the molecular pathology of lung cancers.
Besides heterogeneity, poor cancer-related prognosis is associated with the metastatic potential of lung cancer cells, which is defined as the ability of lung cancer cells to spread from the site of origin (primary tumor) to other distant organs (secondary tumor).
What is the WHO classification of lung cancers?
Adenocarcinoma
Based on the metastatic ability, the World Health Organization (WHO) has categorized adenocarcinomas into three subtypes: adenocarcinoma in situ (preinvasive tumors with a diameter of ≤3 cm), minimally invasive adenocarcinoma (tumor diameter: ≤3 cm; invasion size: ≤5 mm), and invasive carcinoma.
In general, adenocarcinoma is characterized by the presence of a lepidic pattern, which means the growth of cancer cells along the alveolar lining without vascular or stromal invasion. For accurately diagnosing minimally invasive adenocarcinoma, the presence of lymphatic, vascular, or pleural invasion or necrotic tumor can be regarded as exclusion criteria.
Invasive adenocarcinoma, on the other hand, is categorized based on five main features, including lepidic, papillary, acinar, micropapillary, and solid adenocarcinoma. Invasive adenocarcinomas are broadly categorized into two forms: lepidic-predominant adenocarcinoma (invasion size: >5 mm) and invasive mucinous adenocarcinoma.
Squamous cell carcinoma
According to the updated version (2015) of the WHO classification, squamous cell carcinomas are of three types: keratinizing, non-keratinizing, and basaloid.
Small cell lung carcinoma
Small cell lung carcinoma is the most common type of high-grade neuroendocrine tumors, which also include large cell neuroendocrine carcinoma and carcinoid tumor.
What are the molecular characteristics of lung cancers?
The Cancer Genome Atlas has done extensive molecular characterization of lung cancers to identify therapeutic targets.
Adenocarcinoma
According to the Cancer Genome Atlas data, the rate of somatic mutations is high in lung adenocarcinomas. In adenocarcinoma patients, about 18 statistically significant genetic mutations have been found, including TP53, KRAS, KEAP1, STK11, EGFR, NF1, and BRAF.
Of these mutations, EGFR, KRAS, and BRAF are the most common driver genetic mutations that increase the signaling through RTK/RAS/RAF pathway. Besides driver genetic mutations, amplification of two oncogenes namely ERBB2 and MET are responsible for increased RTK/RAS/RAF signaling.
A subgroup of adenocarcinomas with wild-type EGFR and KRAS mutations harbor a genetic fusion in which echinoderm microtubule-associated protein-like 4 gene is fused with anaplastic lymphoma kinase gene (EML4-ALK fusion).
Based on the mRNA characterization, lung adenocarcinomas are categorized into new transcriptional subtypes, including terminal respiratory units (EGFR mutations and kinase fusion), proximal inflammatory (NF1 and TP53 co-mutations), and proximal proliferative (KRAS mutation and STK11 inactivation) subtypes.
Based on DNA methylation characterization, adenocarcinomas are subdivided into CpG island methylator phenotype (CIMP)-high, CIMP-intermediate, CIMP-low subtypes. The CIMP-high subtype is often characterized by MYC overexpression and hypermethylation of WNT signaling pathway genes.
In addition, lung adenocarcinomas can be subdivided into 6 groups depending on the protein expression characterization. The most common proteins that are differentially expressed in these groups include HER2, β-catenin, Smad4, Cyclin D1, mTOR, and Rad50.
Squamous cell carcinoma
In squamous cell carcinoma patients, about 11 statistically significant mutations have been found, including TP53, CDKN2A, PTEN, PIK3CA, KEAP1, NOTCH1, and RB1. About 90% of these mutations occur in the TP53 gene. A novel loss-of-function HLA-A mutation has also been observed.
Cellular signaling pathways that are frequently affected by these mutations include oxidative stress pathway, PI3K/AKT pathway, and squamous cell differentiation pathway.
Alike adenocarcinoma, mRNA characterization divides squamous cell carcinomas into four subgroups: classical (KEAP1, PTEN, and NFE2L2 alterations), basal (NF1 alteration), secretory, and primitive (RB1 and PTEN alterations).
MicroRNA characterization divides squamous cell carcinomas into four subgroups, which partially overlap with mRNA-based subgroups. Based on DNA methylation, squamous cell carcinomas are divided into four subgroups: methylation clusters 1 to 4, in which clusters 2 and 4 have the lowest and highest rates of DNA hypermethylation, respectively.
Small cell lung carcinoma
The most frequent genetic alterations observed in small cell lung cancer include SOX2 amplification and RLF-MYCL1 fusion. In addition, mutations in genes involved in histone modification (CREBBP, EP300, MLL) are commonly observed. Because of the heavy smoking status associated with small cell lung carcinoma, C: G>A: T transversions are found in about 30% of the mutations.
Based on mRNA characterization, small cell lung carcinoma is divided into two subgroups: group 1 (17%) and group 2 (83%). In group 2, increased expressions of CHGA, GRP, ASCL1, and DLK1 are observed; whereas, in group 1, these genes are found to have a lower level of expression.
Further Reading
Last Updated: Aug 13, 2020