Skip to
DNA-Protein interactions play an essential role in the regulation of transcription. Understanding how DNA binds to specific residues to mediate binding specificity alongside determining the recognition mechanism has presented a molecular and computational challenge to scientists. Nonetheless, the basis of the interaction of DNA and proteins has neem demonstrated through several computational and experimental methods.
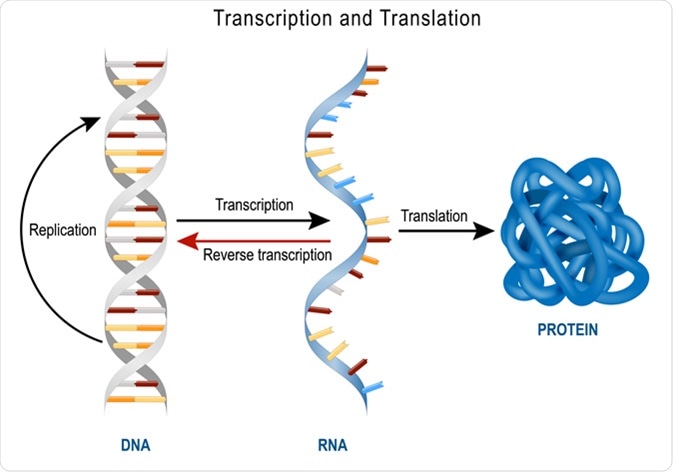
DNA Replication, Protein synthesis, Transcription and translation. Image Credit: Designua / Shutterstock
DNA-Protein interactions are mediated by one of two means; (i) direct contact between the base pairs of DNA and specific amino acids in the protein structure. This type of contact is intermolecular and termed a direct readout mechanism; (ii) indirect contact between the protein and DNA, mediated predominantly by water molecules and conformational changes in the structure of DNA. This type of contact is intramolecular and classified as an indirect readout mechanism.
Notably, the direct readout mechanism is both redundant and flexible; this suggests the interaction is not based on a simplistic code. Furthermore, mutational analysis of specific bases on DNA that not in direct contact with amino acids often affect binding affinity. These changes are thought to arise from changes in the structure of intramolecular water molecules and bridging amino acids and bases, conformational shifts in the DNA structure and/or its flexibility. This further supports the role of indirect readout and the significance of intramolecular interactions for DNA-Protein binding.
Structural analysis of DNA-Protein complexes
Three-dimensional structures have been used to understand the variables that affect and mediate the formation of DNA-Protein complexes. One such important factor are hydrogen bonds. Several studies have demonstrated how hydrogen bonds aid in recognition of cognate DNA-Protein interactions; analyses have demonstrated that the C5 of cytosine and C5-Met of thymine form weak hydrogen bonds with the amino acids Aspartate, Asparagine, Glutamine, Glutamate, Serine and Threonine. Furthermore, analysis of DNA-binding sites has revealed discontinuous sequence segments form hydrophilic surfaces which are prime sites of hydrogen binding interactions.
Energetic contributions
Analysis of the free energies of several forms of interactions including electrostatics, hydrogen bonds, Van der Waals and packing has demonstrated that Van der Waals contributions mediate the formation of DNA-Protein complexes whilst electrostatic forces are unfavorable. Despite this, basic residues, which carry net positive charges are found to mediate favorable contribution to binding despite a desolvation penalty which arises. This refers to the removal of ordered water molecules from the basic residue which shield the charge in order to allow direct electrostatic interaction with an oppositely charged species. Acidic and neutral residues were poor electrostatic contributors; since DNA is negatively charged due to the sugar-phosphate backbone, this concurs with the finding. Acidic residues carry a negative charge, which would result in energetically unfavorable interactions with DNA. Similarly, neutral bases offer no charge stabilization.
Another important mediator of DNA-Protein binding is cation–π interactions. These are a form of noncovalent interaction in which the face of an electron rich π system, typically a heterocyclic ring system such as those seen in the benzene or indole rings common to the aromatic amino acids, and an adjacent positively charged ion (cation). 73% of protein–DNA complexes were found to involve such interactions, and these were found to function over long ranges. Furthermore, of the six possible pairs of residues Arginine–Tyrosine was found to be the strongest.
The residues on the binding protein occur largely as a group of conserved residues which display high packing density; the solvent interface area ranges between 1120 and 5800 Å2 and the binding site is populated with positively charged groups predominantly contributed by Lysine and Arginine side chains and the phosphate groups of DNA molecules.
Water-mediated contacts and conformational changes of DNA
The conformational changes of DNA allow structural rearrangements to take place that are essential in mediating complex formation. Numerous studies have demonstrated that this conformational switching of DNA has also been found to mediate specificity. The conformational changes are evaluated by measuring six base step parameters that include shift, slide, rise, tilt, roll, and twist. In general, local variations particularly in the minor groove of the DNA alongside electrostatic interactions mediate the general mechanism for DNA binding specificity. Furthermore, water-mediated contacts control binding specificity.
Base–amino acid interaction
Computational simulations measuring the free energies of interactions between DNA base pairs and amino acid side chains have been computed and enabled systematic analyses of DNA-Protein interactions. In these methods, the side chain orientations are manipulated to generate a series of rotamers; for each positioning event, approximately 1 million conformations are generated which are subsequently averaged in order to calculate the thermodynamic parameters of free energy, enthalpy, and entropy. This generates free energy maps that show that the conformational entropy of side chain is instrumental in mediating specificity.
Mechanism for protein–DNA recognition
The aromatic residues and positively charged residues are the predominant interacting partners together with the phosphate groups of the DNA backbone in the formation of DNA-Protein interactions. Since these interactions are electrostatic in nature, it has been speculated that these underpin the specificity of the DNA-Protein complex formation. During the process of complex formation, hydrogen bonds between the polar amino acids and the atoms of the DNA molecule augment the affinity of binding; hydrogen bonds are also made between the main chain atoms of DNA and amino acid residues and hydrophobic interactions further mediate complex stability. The DNA itself also mediates the complexation process; as discussed earlier the conformational flexibility exhibited by the molecule affects binding specificity. This is particularly true as there exists as relationship between DNA stiffness and binding specificity of protein–DNA complexes.
The availability of protein–DNA complex structures has enabled the study of what mediates their formation. Of these noncovalent interactions, DNA conformation, and intermediate water molecules play an important part in the complexation process.
From DNA to protein - 3D
Further Reading
Last Updated: May 21, 2019