Aneuploidy is defined as a chromosomal aberration involving either an addition or deletion of chromosomes compared to the ‘normal' number of 46 chromosomes. Flow cytometry is a useful tool for detecting anueploidy.
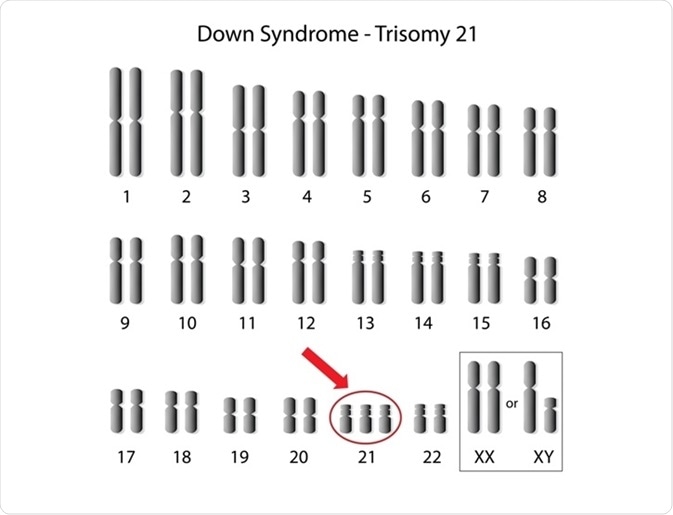 Alila Medical Media | Shutterstock
Alila Medical Media | Shutterstock
Why flow cytometry?
Traditionally, aneuploidy was diagnosed by counting the metaphase chromosomes under the microscope. However, in many cases, and primarily due to the high or lower number of chromosomes, it was difficult to obtain a precise count. In addition, the method involves a lot of time and handling of strong acids, which makes it unsuitable for large-scale use.
Flow cytometry surpasses the aforementioned disadvantages since it quantifies the amount of DNA rather the counting the number of chromosomes. Also, due to the self-contained nature of the method, there is no interaction with hazardous solvents.
Flow cytometry is an accurate, reliable technique
The accuracy of flow cytometry is subject to following conditions: low level of variation coefficient of the DNA peaks, an equal proportion of cells in each peak, and a small difference in DNA of the sample and internal reference.
For example, if the DNA peak in the internal reference peaks at 100, the peak of the DNA from the samples should lie in the range of +/- 50 from the reference peak. The mean positions should be defined for both reference and sample peaks. The standard ratio is constant and refers to the ratio of the sample peak and reference peak.
Flow cytometry to detect aneuploidy in plant samples
In plant samples, the midrib of an open leaf is cut using a razor in the petri dish that consists of citric acid and Tween 20. The sample can be filtered using a nylon mesh. The internal reference standard, such as red blood cell (RBC) nuclei and nuclear stain are added to the buffer.
DNA content is determined by comparing the nuclei of the internal reference and the sample nuclei, and around 5000-10000 nuclei are analyzed to get an average. However, in certain cases, all chromosomes may not have the same number, which can potentially lead to complications in the flow cytometry analysis.
Flow cytometry to detect aneuploidy in cancers
Flow cytometry has also been performed to detect aberrations in the chromosome number in human small cell lung carcinoma. In this case, two or three tissue sections that were approximately 50 micrometers wide were dewaxed in xylene, subjected to rehydration in ethanol, and then washed in distilled water.
The tissues were treated with citrate and pepsin to obtain single-cell suspensions. These were then filtered in nylon mesh and washed. Dissociated nuclei were stained for nuclei using propidium iodide. These nuclei were then subjected to flow cytometry and analyzed using the ModFit software.
Analyzing the data
In cases where the coefficient of variation exceeds 10%, the histograms are discarded and not used for the final analysis. The sample is considered diploid in cases where there is a single and separate peak in G0/G1 phase. The histograms that were tetraploid or multiploid are classified as aneuploid.
Further Reading
Last Updated: Mar 28, 2019