Plasmid DNA isolation from bacteria is a vital molecular biology method used to create template DNA for preferred downstream reactions. Plasmid isolation approaches are basic; however, high-throughput plasmid DNA extraction has been challenging, with problems such as low yield and genomic DNA (gDNA) contamination.
An automated, high-throughput, multi-well-based plasmid extraction technique, with a yield suitable for downstream applications and free from gDNA, will significantly profit high-throughput laboratories, where plasmid extraction is frequently a bottleneck.
The 96-well, silica filter-plate plasmid extraction kits are available in the market. In these approaches, bacteria cells are harvested and lysed by alkaline lysis, the cellular debris is eliminated, and the extracted DNA is then gathered by binding to a silica membrane. Membranes are later washed, before the elution of purified DNA.
In this article, Analytik Jena describes a technique developed at SynbiCITE’s London DNA Foundry, which uses the CyBio® FeliX pipetting platform to automate a 96-well, silica filter-plate based plasmid DNA extraction protocol.
The use of the CyBio® FeliX pipetting platform (Figure 1) enables 96 samples to be processed at the same time, in approximately 1.5 hours, with the versatility to process numerous plates simultaneously. The compact platform minimizes the bench space required for an automated plasmid isolation system in a laboratory and, along with a bench top centrifuge and robotic arm, can enable a completely automated system for plasmid extraction. Using this technique, an average yield of 65 ng/μL plasmid DNA is separated from bacterial cultures, with low variability between the samples (±7.7 ng/μL SD) and in an elution volume of 50 μL. The plasmid DNA samples are also of high quality, ideal for downstream applications such as transformation or sequencing.

Figure 1. Integrated Cybio® FeliX pipetting platform in SynbiCITE’s London DNA Foundry laboratory at Imperial College London.
Challenge
The automation of isolation of plasmid DNA from bacterial cells.
Solution
A completely automated technique for the simultaneous isolation of plasmid DNA from 96 samples with the help of Cybio® FeliX.
Materials and Methods
Reagents and Instrumentation
Table 1. Reagents required for the method
| Reagent |
Manufacturer |
Part Number |
| PureLinkTM Pro Quick96 Plasmid Purification Kit |
ThermoFisher Scientific |
K211004A |
| Quant-iT™ Picogreen® dsDNA Assay Kit |
ThermoFisher Scientific |
P7589 |
| Terrific Broth |
Merk |
1016290500 |
Table 2. Instrumentation used in the method
| Instrument |
Manufacturer |
| CyBio® FeliX |
Analytik Jena |
| Pipetting Head R 96/250 μl |
Analytik Jena |
| CyBio® RoboTipTray 96-250 μl DW |
Analytik Jena |
| 5810 R Centrifuge |
Eppendorf |
Table 3. Consumables required for the method
| Item |
Manufacturer |
Part Number |
| 96 well, MASTERBLOCK®, 2 ml |
Greiner |
780271 |
| 96 well plates, deep well, 1 ml |
ThermoFisher Scientific |
11381555 |
Sample Preparation
- Grow bacterial clones, having plasmid DNA of interest
- Inoculate single clones into a 96-well, 2 mL, square-well plate holding 1.2 mL/well Terrific broth (TB) with suitable antibiotic(s)
- Grow cells overnight shaking, 37 °C
- It is recommended to use high copy number plasmid for higher yield
Method
- All buffers are prepared as instructed in the manufacturer’s guidelines and dispensed into 96-well, 1 mL plates in compliance with the volumes in Table 4 (volumes mentioned are appropriate for 1 x 96-well plasmid extraction)
- The plate holding overnight bacterial cultures is centrifuged at 2250 x g for 10 minutes
- The supernatant is removed to make sure that cell pellets remain intact
- RoboTipTrays (x5), buffer plates (x5), sample plate (holding cell pellets), and the “Clarification” plate (on top of a 96-well, 1 mL plate) are added to the deck of the CyBio® FeliX. For instance, as illustrated in Figure 2.
Table 4. Volumes of buffer to add to 96-well, 1 mL buffer plates for 1 x 96-well plasmid extraction.
| Buffer |
Volume (μl/well) |
| Resuspension |
350 |
| Lysis |
350 |
| Neutralization |
450 |
| Wash |
1000 |
| Elution |
150 |
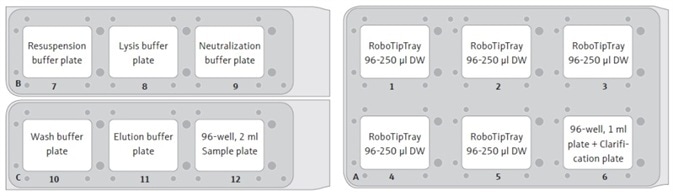
Figure 2. Suggested set-up positions for the CyBio® FeliX deck. Each RoboTipTray is used for a specific reagent(s), according to its position: (1) Resuspension buffer and sample handling (pre-clarification), (2) Lysis buffer, (3) Neutralization buffer, (4) Wash buffer and sample (post-clarification), and (5) Elution buffer. The sample plate will contain cell pellets, post-harvesting by centrifugation.
- With the help of the CyBio® FeliX, 250 μL/well resuspension buffer is added to the sample plate and blended well (with the same tips) for 3 minutes to resuspend the cell pellets (refer Table 5 for all pipetting speeds)
- 250 μL/well lysis buffer is added to the sample plate
- Using the sample tips, the buffer is blended for 4 minutes to lyse the cells
- 350 μL/well neutralization buffer is added to the sample plate to halt acidification of the DNA
- Blending is performed for 3 minutes using the sample tips
- The total volume of the lysed sample (850 μL/well) is shifted to the Clarification plate (given in kit), using the sample tips
- The Clarification plate is centrifuged on top of 1 mL, deep-well collection plate at 2250 x g, for 2 minutes. The sample will pass via the clarification filter, conveying the DNA to the collection plate below and eliminating the cell debris.
- The Clarification plate is disposed of and the 1 mL, deep-well collection plate is returned to the top deck of the FeliX (position 12, swapping the 96-well, 2-mL sample plate)
- The “Filter” plate (given in kit) on top of a clean 1 mL, deep-well collection plate is integrated to the bottom deck of the FeliX (position 6)
- The total volume of the clarified sample (850 μL/well) is shifted to the Filter plate, using clean sample tips
- The Filter plate is centrifuged on top of a 1 mL, deep-well collection plate at 2250 x g, for 2 minutes. The DNA will currently be bound to the silica membrane in the Filter plate.
- The flow-through solution is disposed of from the 1 mL, deep-well collection plate
- The Filter plate is positioned back on top of the 1 mL, deep-well collection plate on the bottom deck of the FeliX (position 6)
- 900 μL/well wash buffer is added to the Filter plate, using the sample tips (wash buffer plate should be prepared fresh every time)
- The Filter plate is centrifuged on top of the 1 mL, deep-well collection plate at 2250 x g, for 2 minutes. The DNA will stay attached to the silica membrane in the Filter plate.
- The flow-through solution is disposed of from the 1 mL, deep-well collection plate
- The Filter plate is centrifuged on top of the empty 1 mL, deep-well collection plate at 2250 x g, for 10 minutes, to eliminate any residual wash buffer. The DNA will stay bound to the silica membrane in the Filter plate.
- The Filter plate is positioned on top of the “Elution” plate (given in kit) on the bottom deck of the FeliX (position 6)
- 50 μL/well elution buffer is added to the Filter plate
- The plate is incubated at ambient temperature for 4 minutes
- The Filter plate is centrifuged on top of the Elution plate at 2250 x g, 2 minutes, to elute the DNA from the silica membrane
- The Elution plate holding the purified plasmid DNA samples is sealed with a foil seal and stored at 4 °C (short term) or −20 °C (long term)
Table 5. Pipetting speed for different solutions using the CyBio® FeliX pipetting platform
| Solution |
Pipetting speed |
| Resuspension buffer |
Default |
| Lysis buffer |
15 μl/s |
| Neutralization buffer |
20 μl/s |
| Lysed sample (pre-clarification) |
15 μl/s |
| Clarified sample |
Default |
| Wash buffer II |
Default |
| Elution buffer |
Default |
Results and Discussion
To realize a good yield from plasmid DNA isolation, it is vital to grow bacteria cells perfectly before attempting the method. Here, a Design of Experiments (DOE) approach was used to enhance the overnight growth of DH5α Escherichia coli (E. coli) cells at 37 °C. The quantity of cell growth was established by measuring the absorbance of 100 μL/well culture at 600 nm (Abs600). Using a custom-designed model, produced in the JMP® software, it was discovered that the ideal conditions for cell growth were to use a 2 mL, square-well, 96-well plate, holding 60% volume of media per well (1.2 mL/well) (Table 6 and Figure 3).
Table 6. Design of experiment (DOE) model for optimizing overnight E. coli cell growth conditions, in a 96-well plate. Three factors including plate type, volume of media per well, and incubator shaking speed were evaluated using a custom-designed model generated with JMP® software. There were a total of 20 different runs in random orders from five whole plots. Each whole plot represents the same condition for shaking speed and was performed on separate days. Three plate types were tested: SW = standard well (maximum volume 200 μL); DW_1ml = deep, round well (maximum volume 1 mL); and DW_2ml = deep, square well (maximum volume 2 mL). Volume indicates the media volume expressed as a percentage of the maximum volume of each well. All cells were grown overnight at 37 °C in a shaking incubator (Kuhner ISF1-XC Climo-shaker) and the response measured for each run was the absorbance at 600 nm (Abs600) of 100 μL/well sample.
| Solution |
Pipetting speed |
Solution |
Solution |
Solution |
Solution |
| 1 |
1 |
SW |
100 |
250 |
0.242 |
| 2 |
1 |
DW_2ml |
75 |
250 |
0.548 |
| 3 |
1 |
DW_1ml |
100 |
250 |
0.218 |
| 4 |
1 |
DW_1ml |
50 |
250 |
0.465 |
| 5 |
2 |
DW_2ml |
100 |
300 |
0.195 |
| 6 |
2 |
DW_2ml |
50 |
300 |
0.594 |
| 7 |
2 |
SW |
75 |
300 |
0.552 |
| 8 |
2 |
DW_1ml |
75 |
300 |
0.172 |
| 9 |
3 |
SW |
50 |
250 |
0.484 |
| 10 |
3 |
DW_1ml |
50 |
250 |
0.24 |
| 11 |
3 |
DW_2ml |
75 |
250 |
0.49 |
| 12 |
3 |
DW_1ml |
100 |
250 |
0.117 |
| 13 |
4 |
SW |
75 |
200 |
0.465 |
| 14 |
4 |
DW_2ml |
50 |
200 |
0.265 |
| 15 |
4 |
DW_1ml |
75 |
200 |
0.182 |
| 16 |
4 |
DW_2ml |
100 |
200 |
0.127 |
| 17 |
5 |
DW_1ml |
75 |
250 |
0.176 |
| 18 |
5 |
SW |
50 |
250 |
0.507 |
| 19 |
5 |
SW |
100 |
250 |
0.37 |
| 20 |
5 |
DW_2ml |
75 |
250 |
0.46 |
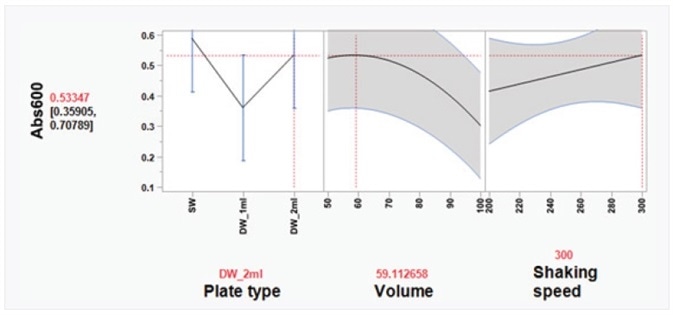
Figure 3. Optimal cell growth model visualized using the Prediction Profiler tool in JMP®. Using JMP® Design of Experiment (DOE) software, optimal growth conditions of E. coli cells in 96-well plates were determined, based on three variables: plate type, volume, and shaking speed. Three plate types were tested: SW = standard well (maximum volume 200 μL); DW_1ml = deep, round well (maximum volume 1 mL); and DW_2ml = deep, square well (maximum volume 2 mL). Volume indicates media volume expressed as percentage of maximum volume of well. DH5α E. coli cells were grown overnight at 37 °C in a shaking incubator (Kuhner ISF1-XC Climo-shaker). The absorbance at 600 nm (Abs600) of 100 μL/well sample was measured using a BioTek SynergyTM Mx Microplate Reader. Based on the JMP® DOE model, the statistically significant effects were plate type and volume of media. In a 2-mL deep-well plate, the Prediction Profiler tool in JMP® recommends using 60% volume of media per well (1.2 mL/well). Using these conditions, the predicted Abs600 is 0.533. These conditions were tested and the actual Abs600 measured was 0.572 (± 0.12 SD).
It should be kept in mind that high Abs600 values were also acquired when cells were grown in a typical depth, 96-well plate. However, because of the compact volume capacity of the wells in these plates, the total number of cells was not adequate for plasmid DNA extraction. It was learned that shaking speed did not have a major effect on cell growth. Moreover, it was established that the growth of cells in Terrific Broth (TB) enhanced the yield of isolated plasmid DNA, as compared to Lysogeny broth (LB) (Figure 4).
The growth of bacteria cells under enhanced conditions is vital for gaining a good yield of plasmid DNA. The yield achieved under the ideal conditions was more than 50 ng/μL (Figure 4), in an elution volume of 50 μL, which is adequate for downstream applications such as sequencing [1].
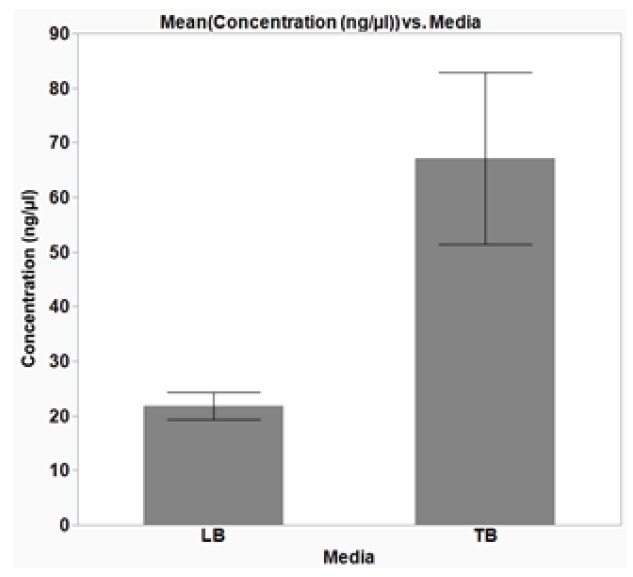
Figure 4. Optimal growth media for plasmid DNA isolation from E. coli cells. E. coli DH5α cells were grown overnight, shaking at 37 °C, in a 2-mL, square-well, 96-well plate in either Lysogeny broth (LB) or Terrific Broth (TB), 1200 μL/well. Plasmid DNA (p15a ORI) was extracted from the cells and the concentration of the isolated DNA was determined using a NanoDrop (Thermo). The average data from three replicates are plotted with error bars representing the standard deviation (SD). The isolated DNA yield from cells grown overnight in TB media was greater than when cells were grown in LB. Furthermore, the average yield from cells grown in TB was greater than 50 ng/μL in a 50 μL elution volume, which is sufficient for sequencing. The average 260/280 ratio was 1.9 (±0.07 SD) and 1.97 (±0.07 SD) for cells grown in LB and TB, respectively, indicating good DNA purity.
Analytik Jena has demonstrated that the plasmid isolation technique defined in this article offers a reproducible yield from samples processed concurrently in a 96-well plate (Figure 5). High-quality DNA, with an average yield of 65.3 ng/μL (±7.7 SD), in a 50 μL volume is acquired from a low copy number plasmid (p15a ORI), which is enough for downstream applications such as sequencing or transformation into bacteria cells.
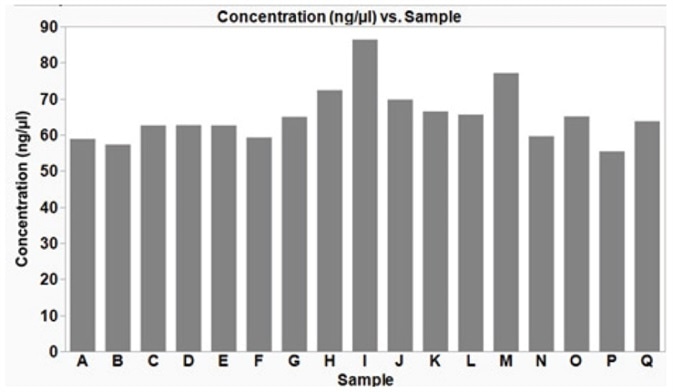
Figure 5. Plasmid DNA extracted from multiple bacterial E. coli samples simultaneously, using an automated method on the CyBio® FeliX. Using the automated method described here, plasmid DNA (p15a ORI) was isolated from E. coli DH5α cells, grown under optimized conditions. The isolated plasmid DNA yield was reproducible across the samples with an average yield of 65.3 ng/μL (± 7.7 ng/μL SD). The average 260/280 ratio was 2.0 (± 0.05 SD), indicative of good DNA quality.
As anticipated, the use of a higher copy number plasmid produces a better yield of isolated plasmid DNA (Figure 6). Consequently, the use of high copy number plasmids provides an option for higher yield of plasmid DNA, if necessary.
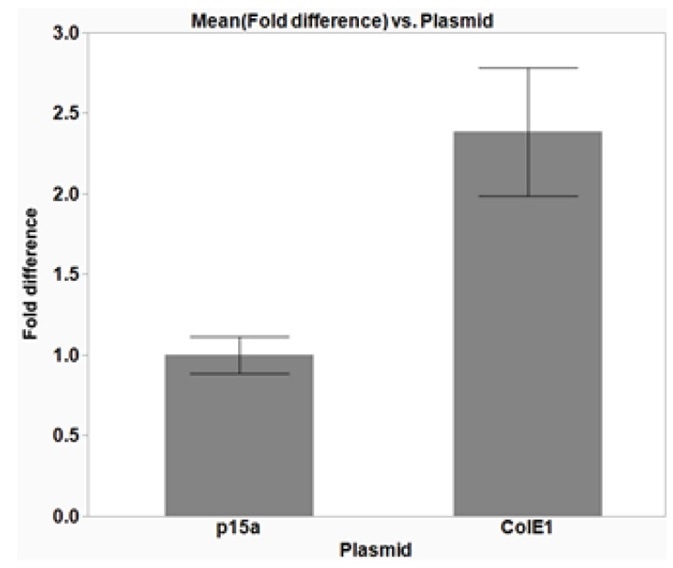
Figure 6. The effect of plasmid copy number on isolated DNA yield. DH5α E. coli cells were transformed with cloning vectors with different origins of replication (ORI) to investigate the effect of copy number on the yield of isolated plasmid DNA. p15α plasmids have a low copy number, while ColE1 plasmids have a higher copy number. The average data from three replicates are plotted with error bars representing the standard deviation (SD). The use of the higher copy number plasmid, ColE1, increases the isolated plasmid DNA yield by more than two-fold, as compared to the p15a plasmid. The average 260/280 ratios were 1.91 (± 0.08 SD) and 1.89 (± 0.03 SD) for p15a and ColE1 plasmids, respectively, indicating good DNA quality.
The technique explained here can be used to process a multi-well plate of 96 samples concurrently, using the CyBio® FeliX pipetting system. If used along with a robotic arm and an incorporated centrifuge with adequate depth for the filter plates, this technique can be completely automated and adapted to process numerous 96-well sample plates at a single time.
Conclusion
With the increase in higher throughput demand in molecular biology laboratories, there is a need for implementing essential methods, like plasmid DNA isolation, on a high-throughput, automated scale.
In this case, Analytik Jena aimed to formulate a technique for the high-throughput isolation of plasmid DNA from bacterial cell cultures, which can be 100% automated. It was shown that this technique isolates a lot of high-quality, plasmid DNA from bacterial cultures, at yields ideal for downstream applications. The DNA yield is extremely dependent on the quality of the overnight growth of bacterial cultures; hence, Analytik Jena has drawn enhanced growth conditions that are vital for the success of this technique. The application of the compact CyBio® FeliX pipetting platform for high-throughput plasmid DNA isolation delivers an option to allow molecular biology laboratories to match the demand and to overcome a typical bottleneck in laboratory workflow.
References
[1] https://eurofinsgenomics.eu/
About Analytik Jena US
Analytik Jena is a provider of instruments and products in the areas of analytical measuring technology and life science. Its portfolio includes the most modern analytical technology and complete systems for bioanalytical applications in the life science area.
Comprehensive laboratory software management and information systems (LIMS), service offerings, as well as device-specific consumables and disposables, such as reagents or plastic articles, complete the Group’s extensive range of products.
About Life Science
The Life Science product area demonstrates the biotechnological competence of Analytik Jena AG. We provide a wide product spectrum for automated total, as well as individual solutions for molecular diagnostics. Our products are focused to offer you a quality and the reproducibility of your laboratory results.
This will surely ease your daily work and speed up your work processes in a certain way. All together we support you through the complete process of the lab work. Besides we offer customized solutions and are able to adapt our products to your needs. Automated high-throughput screening systems for the pharmaceutical sector are also part of this segment’s extensive portfolio.
About Analytical Instrumentation
Analytik Jena has a long tradition in developing high-performance precision analytical systems which dates back to the inventions made by Ernst Abbe and Carl Zeiss. We have grown to become one of the most innovative manufacturers of analytical measuring technology worldwide.
Our business unit Analytical Instrumentation offers excellent competencies in the fields of optical spectroscopy, sum parameters and elemental analysis. Being proud of our core competency we grant all our customers a long-term warranty of 10 years for our high-performance optics.
About Lab Automation
With more than 25 years of market experience, Analytik Jena with its CyBio® Product Line is a leading provider for high quality liquid handling and automation technologies. In the pharmaceutical and life science industries, our products enjoy the highest reputation for precision, reliability, robustness and simplicity.
Moreover, the Automation Team designs, produces and installs fully automated systems tailored to our clients' application, throughput and capacity requirements. From stand-alone CyBio® Well up to fully customized robotic systems we handle your compounds, biomolecules and cells with great care.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.Net which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.