In this interview, News-Medical spoke to Dr. Thierry Le Bihan, Principal Scientist for the REpAb™ project, about how polyclonal antibody sequencing can lead to the next generation of antibody discovery.
Can you give us a brief overview of why you think it’s beneficial to generate and de novo sequence polyclonal antibodies from a single animal?
From a production point of view, it is important to have polyclonal antibodies (pAbs) that are reproducible from batch-to-batch. The standard strategy consists of mixing several pAbs isolated from different animals (2-20 animals). The reason for this is they are able to generate an averaged product in large quantities. By doing this, the pAb is less sensitive to variability.
However, we are proposing a different approach from a sequencing point of view. The previously described pAb mixture is difficult to sequence as the level of complexity is significantly increased by pooling the different animals. Instead, it would be worthwhile to assess each individual animal and select the best producer based on the unique desired quality of the generated pAbs (i.e., immunoprecipitation, Western blot, ELISA, etc.). The benefit of this approach is that you can identify the most suitable pAbs from a given producer, decode it and then make several oligo forms using recombinant protein expression. This will thereby reduce batch variability and also allow working with a product that’s better than average.
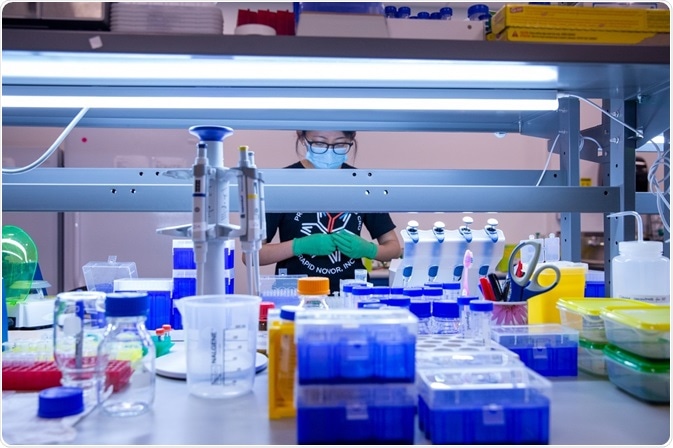

Image Credit: Rapid Novor Inc
What are the main challenges that arise when using polyclonal antibodies as a research tool?
Polyclonal antibodies (pAbs) are usually secreted by different B-cell clones, whereas monoclonal antibodies (mAbs) are produced by identical B-cells. What this means is that the heterogeneous nature of pAbs allows them to bind multiple epitopes of an antigen. Additionally, our native immune system generates complex pAbs in response to antigens and infection, this response is one that many scientists try to replicate in the lab. The way this response is replicated is through injecting animals with a specific antigen that will stimulate the immune response. The generated pAbs are then purified from the serum and are often pooled in order to reduce batch variability.
One of the biggest challenges of polyclonal-based development is the concept of the reproducibility crisis, which stems from batch-to-batch variability that can occur during production. The problem arises when different host animals are injected with the same antigen but then produce pAbs that have different specificities and affinities. This means that even if one was to use a new batch of antibodies, they cannot reproduce the exact same experimental results. Even more importantly, the same animal sometimes may have different immune responses to the same antigen and also exhibit time-dependent variation known as the affinity maturation process.
As antibodies are the most widely-used reagents, it's unsurprising that the majority of manufacturers produce pAbs. Yet, by the very inconsistent nature of pAbs, scientists have struggled with the reproducibility of the antibody reagents hindering their research progress. Thus, although pAbs are widely adopted in life science research, the challenges they can present are not always offset by their advantages.
Do you have a proposed solution to overcome these challenges associated with polyclonal development?
Yes. Although it may seem impossible to replicate the immune response, it will be possible to efficiently profile and capture the dominating forms of the immune repertoire; which is the first step. Our solution is to obtain the protein sequences of the most abundant antibodies that bind to the antigen in the polyclonal (pAb) population directly from blood or protein mixture.
Our current approach involves combining both a genomic and proteomic approach. By using additional B-cell sequencing in combination with protein sequencing, we will be able to assemble the complete heavy and light chains of an antibody. Through sequencing the pAbs from the serum of immunized animals, it will be possible to produce the most abundant mAbs binding to the antigen in vitro by recombinant synthesis with high reproducibility and predictable biological activity. With this approach, the previous batch variability in traditional pAb development will no longer pose a challenge. This advancement will also present the opportunity to reduce the use of laboratory animals and the time required to generate new lead mAb targets and “immortalize” a good product.
The combined approach of sequencing pAbs from serum is largely inspired by Rapid Novor’s existing monoclonal sequencing solution (REmAb®). This platform is a robust mass-spec based de novo solution that sequences mAbs with 100% accuracy and coverage. Around 13% of the antibodies our team sequenced had additional light chains. It was these kinds of insights that necessitated experimental processes that began to form the foundations of our current polyclonal sequencing platform, REpAb™.
Our proof-of-concept study successfully derived the sequence of a specific polyclonal antibody from the serum of immunized rabbits with a high pAb background. We have further expanded our study to sequence antibodies from the blood of recovered COVID-19 patients as part of an antibody therapeutic discovery campaign. Our goal is to capture functional antibodies directly from blood with high throughput thereby facilitating the subsequent assessment of their therapeutic potential. By doing this the process of antibody discovery can be remarkably accelerated. We are currently inviting interested parties to participate in a royalty-free contracted antibody discovery campaign. At this time, our focus is primarily upon rabbits, mice, and humans.
Image Credit: Rapid Novor Inc
Why do you think polyclonal sequencing will allow for the next generation of antibody discovery?
Due to the flexibility and specificity of polyclonal antibodies (pAbs), they are much more suitable than monoclonal antibodies (mAbs) for applications where the conditions are less characterized such as clinical applications. These properties also allow pAbs to be more sensitive in certain assays against a variety of target proteins, cells, or organisms. However, replicating the immune system’s response to a myriad of foreign antigens using traditional experiments does not accurately quantify and categorize the large repertoire of naturally occurring antibodies produced by B-cells.
In particular, the native immune system generates polyclonal antibodies (pAbs) capable of binding multiple epitopes to mediate various immune responses. However, batch variability can occur when trying to replicate the immune response and generate new pAbs against a specific antigen. The current solution involves mixing a large variety of isolated pAbs from different animals to produce an averaged product. Yet, often what we can see is reduced affinity and specificity. By implementing a mass spectrometry-based solution, it will be possible to optimize the best production animal, isolate their pAbs, and then sequence the dominating antibody forms to generate recombinant antibodies.
Thus, a protein-sequencing-based approach to polyclonal development will help to eliminate the challenges associated with batch variability including reproducibility and project abandonment. Further, early results demonstrate that the mAbs derived from the polyclonal sera, have a higher affinity than the original pAbs mixture. This means immune response replication will be more attainable than ever.
What are the existing technologies that scientists have used to discover antibodies from blood?
Different technologies exist but scientists have mainly relied on DNA sequencing approaches for antibody discovery. The antibody proteins from the blood are often measured through next-generation sequencing (NGS) in order to capture and sequence the mRNA of B-cell receptors (BCR) from peripheral blood. This BCR sequencing technology produces large-scale sequence information of B-cell receptors which are indicative of the complete antibody repertoire. To further screen the useful sequence information, B-cell subpopulation is often enriched and characterized either by flow cytometry or immunoprecipitation against a specific antigen prior to sequencing.
Yet there are limitations when implementing a strict BCR-only approach. First, BCR sequencing is restricted to circulating B-cells, which only represents a minority of the total B-cell population. As B-cells mature in the bone marrow and migrate through the peripheral blood and enter into the spleen and lymph nodes, there are blind spots for BCR (unless you extract B-cells directly from the spleen and lymph node). Second, if no enrichment is performed, a large part of B-cells in peripheral blood are natural B-cells that are not directly related to the immune response. This can ultimately interfere with the data analysis and accuracy of the results. Lastly, only a small part of the BCRs on the surface of B-cells will be secreted into the serum to form soluble functional antibodies.
Although the genomic information of B-cell receptors is important to capture when discovering antibodies from the blood, alone it cannot accurately capture nor represent the circulating antibodies at a serological level. Thus, it is necessary, at the moment, to blend a genomic and proteomic approach in order to capture this information.
What are the advantages of REpAb™ polyclonal sequencing over B-cell sequencing?
B-cell sequencing is a good place to start as it provides large-scale genomic data, however without incorporating insights that are only capturable via protein sequencing (i.e., characterizing the real-time status of antibodies), you will have an incomplete and superficial understanding of the immune response.
As we are sequencing selected specific antibodies that have been isolated from blood using modern mass spec-based proteomics methods, we are able to sequence directly from convalescent plasma or serum proteins. Compared to a pure NGS-based sequencing approach, REpAb™ provides more accurate and specific antibody sequences as it’s based on relative abundance information. Additionally, we report on the end-point product (i.e., the IgG) not on an earlier intermediate stage (B-cell sequencing analysis).
By combining our proteomics approach with B-cell sequencing, we will be able to combine the advantages of the two methods. The benefits of blending both approaches are that the proteomics data can be interpreted in reference to a BCR sequencing database derived from the same donor. This will allow us to perform a more comprehensive and accurate analysis of immunity which will be an important step forward in antibody-based therapeutic discovery and development.
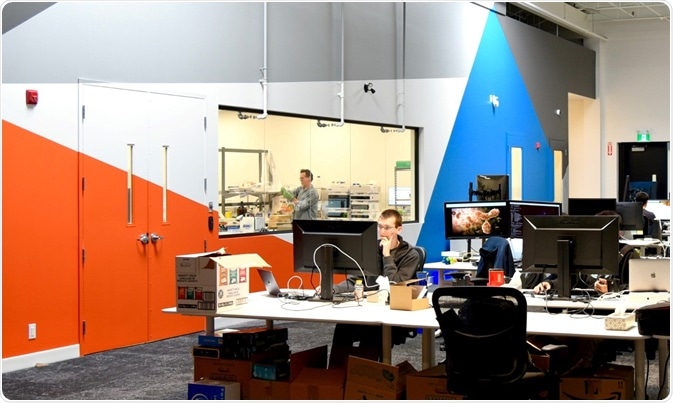

Image Credit: Rapid Novor Inc
How does your REpAb™ polyclonal sequencing solution combine both genomics and proteomics?
Our process begins with the plasma or serum of an immunized animal or convalescent plasma of a patient. By performing affinity purification with chromatography methods, we first extract and purify the antigen-specific pAb mixture from the plasma. In parallel, B-cells are also extracted from the blood and the RNAs are sequenced by NGS to output transcriptomics data.
The proteomics method involves several serial and parallel steps. We digest the pAb proteins into short peptides of overlapping sequences using multiple proteases. We then take the fragmented peptides and analyze them with liquid chromatography-tandem mass spectrometry (LC-MS/MS) to identify the sequence of each peptide. From there we implement our AI-based bioinformatic algorithms to assemble the short peptide pieces into longer peptides that belong to individual antibodies. In addition, we also separate the intact antibodies and perform protease digestion of those separated fractions. In other words, we gather information at a multiscale level from short peptides to intact proteins that are abundantly present in the mixture.
Once we have the proteomics data from mass spec, the transcriptomics data from NGS containing sequence information of large amounts of antibodies can be conveniently used for our bioinformatics software to further assemble and derive the complete amino acid sequences of the most abundant mAbs in the pAb mixture.
What are the future milestones for your polyclonal sequencing development?
While we are already able to independently determine the sequence of a monoclonal antibody (mAb), deriving sequences from a pAb mixture with full coverage still require NGS data due to the sequence complexity. We would like to develop a mass spec-based de novo sequencing solution that can pair the heavy and light chains of complex polyclonal antibodies without the assistance of other tools.
Additionally, our team’s current R&D is focused on sequencing dominant antibodies from convalescent patients. Our goal is to discover and identify therapeutic antibodies from human blood and directly use them in downstream pipeline development.
Image Credit: Rapid Novor Inc
What are the applications of this technology?
As a strategy to reduce the complexity of a pAb mixture, REpAb™ polyclonal sequencing will eliminate the reproducibility challenges that traditional antibody development methods suffer and will help to scale polyclonal development. By reducing the complexity of a pAb mixture into the dominating monoclonal forms, the sequences can then be recombinantly expressed. Early results have demonstrated that these monoclonal sequences have higher binding affinity compared to their original counterparts.
The applications of sequencing pAbs directly from serum or plasma will also give rise to a range of opportunities in the biotechnology and pharmaceutical industries. Notably, for disease diagnosis and monitoring, especially continuous monitoring. Further, as REpAb™ polyclonal sequencing identifies the antibody repertoire in response to a particular antigen or disease, this would allow for the identification of natural antibodies with higher neutralizing capabilities; this methodology has been applied against the Coronavirus in clinical trials.
Additionally, insights regarding the quantification and presence of specific antibodies over a specified period of time will have the potential to aid in the early detection and monitoring of specific diseases. Based on this method, we have developed EasyM™, which is a non-invasive clinical assay for early detection of relapse for Multiple Myeloma (MM) patients. For this particular application, as MM is cancer characterized by an increased level of specific antibodies in the blood, the individual-specific antibody can be sequenced from the MM patient's serum and monitored over time. The same methodology can be applied to other autoimmune diseases and cancers and used to develop further metrics in diagnostic and prognostic applications.
Further, if our current proof-of-concept comes to fruition and we are able to discover therapeutic antibodies directly from human blood, drug development processes will be significantly accelerated as failure rates will decrease and traditional antibody discovery processes can be bypassed. This will help decrease antibody therapeutic campaigns from years to months.
Where can our readers find more information about your company’s technologies?
About Dr. Thierry Le Bihan

Dr. Le Bihan received his Ph.D. in Chemistry/Biophysics from Université Laval. He has extensive proteomics experience through conducting research in globally recognized institutions such as The Campbell Family Institute for Breast Cancer Research and the University of Edinburgh. He joined Rapid Novor in 2018 and is now the principal scientist of the REpAb™ project.