Automated patch clamp techniques offer an advantage over conventional patch clamping as they remove the time constraint and limited throughput associated with manual (conventional) patch clamping.
 Image Credit: Kateryna Kon / Shutterstock
Image Credit: Kateryna Kon / Shutterstock
Manual patch clamping requires considerable skill and experience due to the technically challenging setup requirements, and as such, only allows recording of a small sample of cells (approximately 8-10 per day).
High-throughput patch clamping is particularly pertinent in the field of drug discovery. Screening of a large library of drug compounds forms the basis of drug design. Drug toxicity also requires high-throughput to determine the safety profile of a compound.
Other areas within pharmacology that depend on high-throughput clamping include the recent emergence of therapeutic channel activators or blockers.
Automated patch clamp for cells in solution
The medium in which the cells are applied determines the method of automation. Using a microplate in place of a micropipette to create gigaseal is appropriate for cells suspended in a liquid medium.
Automated patch clamp techniques, therefore, offer a means to screen hundreds of cells each day. Automated patch clamping accesses plates or chips using a “bottom-up” approach rather than “top-down” access to a single cell in conventional patch clamping.
In the conventional patch clamp, the electrode micropipette approaches the cells that are plated on a dish from above, forming a gigaseal with a portion of the membrane. This occurs in a sequential cell by cell manner.
Contrastingly, automated patch clamping places cells on a multi-well plate or chip, which is made of a dielectric material. This produces a surface, which is a poor conductor of electric current, but can separate, or polarize charge, thereby enabling the support of a static electric field within the material.
The dielectric material partitions two solutions and the surface is etched with micrometer holes that emulate the micropipette openings. Single or multiple cells are ‘seeded’ (placed) in each of the wells.
When the external pressure is lowered, a suction force is generated, which captures cells, thereby forming multiple gigaseals. This suction force or agents that produce pores, such as antibiotics, can be used to disrupt the portion of membrane in the hole.
Access to the cell interior enables recordings of current to be taken in either whole-cell or in perforated-patch configurations. This is illustrated in the figure below:
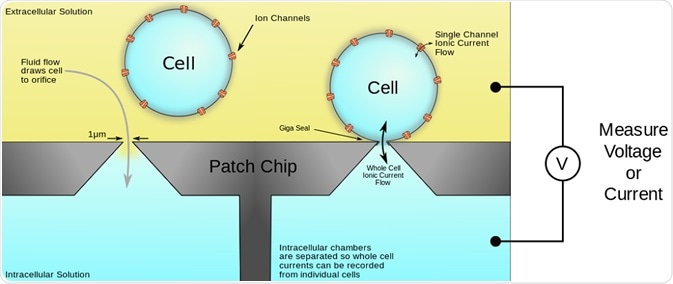
Figure 1. Schematic of the automated patch-clamp system performed on a microplate. The plates have multiple wells, each with either a single hole or multiple holes, which mimic the hole of micropipette used in conventional patch clamping. The cells are seeded in the wells and a pulling force generates suction that pulls the membrane through the hole. Membrane disruption is achieved by pore-forming agents or further suction to allow access to the cell interior for measurement of ion currents.
Automated systems possess additional features, such as fluidic control, suction mechanism and software, which increases throughput, and improves execution of the patch clamping technique.
Automated patch clamp for cells in vivo
When the test cells are in vivo, or in a live organism or tissue, an alternative automated patch clamp technique can be used. An example of in vivo automated patch clamping protocol is provided by Kodandaramaiah, et al. (2012), where recordings from a live mouse cortex and hippocampus were taken.
This protocol employs a robot arm to perform the process of manual patch clamping. This circumvents the need for skilled practitioners, and enables recordings in ‘whole-cell’ configuration to be taken. In this method, the generation and control of three pressure settings is achieved via system of valves.
The three settings are high, low and vacuum, and represent the pressure changes needed to carry out the process. High pressure prevents clogging of the valve by unwanted cells, while low pressure enables the capture of target cells. The vacuum facilitates gigaseal formation with the target cells.
Switching between these pressure differentials is achieved by a computer. The positioning of the micropipette is similarly computerised. The micropipette is lowered by 2-3μm in a stepwise fashion until contact with a cell is made. The accuracy and repeatability of this processes is improved compared to conventional patch clamping techniques where the operator would perform this manually.
The entire protocol is governed by an algorithm that completes the following steps:
- The pipette is lowered under high pressure to prevent clogging into the in vivo medium;
- The pipette at low pressure searches for and captures target cells;
- The pipette under vacuum pressure forms a gigaseal;
- Application of a voltage ‘zap’ to enable membrane rupture, and access to the cytoplasm.
The selection of cells is achieved by monitoring of the electrical resistance when the pipette contacts cells. If a cell is successfully captured, the cell membrane will block a voltage signal sent down the micropipette, and suction is subsequently applied to form the gigaseal.
In conventional patch clamping, the operator would need to decide whether the cell encountered is a target or unwanted cell – the computerization improves the precision of this decision making. A video demonstrating this process can be viewed here:
Automated whole-cell patch clamp electrophysiology of neurons in vivo
The steps performed in automated patch clamping are identical to those involved in manual patch clamping. However, the accuracy and precision of the technique are greatly enhanced, as are the speed at which recordings are taken. Furthermore, human error is removed due to computerized control.
Subsequently, automation allows the manual patch clamping method to be applied broadly in laboratories, which would otherwise require significant labor and training of practitioners.
The production and installation of micropipettes automatically eliminates the need for operators. The system is controlled by a single person monitoring many setups simultaneously, thereby increasing throughput. High throughput methods are sought after in many laboratories, because of the accurate and precise data that can be obtained.
Further Reading
Last Updated: Oct 23, 2018