Biofilms are found in many different settings, providing advantages in industry and disadvantages in human health. As these structures are so complex and found in many different environments, there are lots of different tools and techniques used to monitor and analyze them.
What are Biofilms?
Biofilms are composed of many microorganisms which are permanently adhered to each other and to the surface, forming one large mass which is held together with an extracellular matrix of extracellular polymeric substances (EPS). There are 4 stages of biofilm formation: initial attachment, followed by irreversible attachment, maturation, and dispersion. The matrix is made by the microorganisms within the biofilm throughout development.
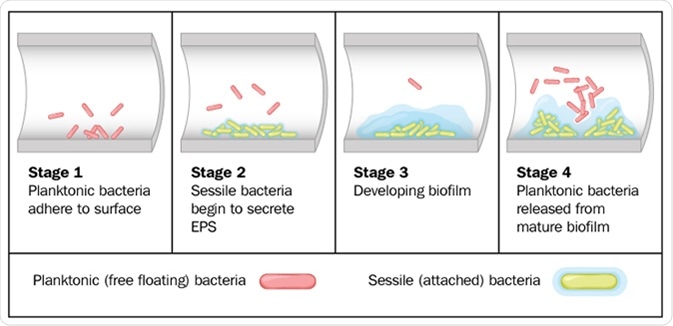
Biofilm formation by free-floating bacteria in the lumen of a tube. Image Credit: Blamb / Shutterstock
Biofilms are quite pervasive and their usage varies. For example, they are often used in sewage treatments to remove organic compounds within the water, but they can also be found naturally in dental plaques and on hip replacements. Tools and techniques to monitor and analyse them include staining, metabolic, genetic and physical assays.
Staining Assays
One of the first staining assays used in biofilm analysis was the crystal violet assay. In this method, the negatively charged molecules within a biofilm (i.e. polysaccharides of the EPS and negatively charged bacterium) are stained with a crystal violet basic dye. Following this staining process, the crystal violet dye is removed and quantified. The amount present can then be used to indicate the size of the biofilm.
However, this method has numerous limitations. For example, this staining process has exceedingly low reproducibility and is not able to distinguish between living and dead cells.
Therefore, several different staining techniques have been developed to overcome these issues, including the 1,9-dimethyl methylene blue (DMMB) assay which binds to Staphylococcus aureus with extremely high specificity, the fluoresceine-di-acetate (FDA) assay which is only taken up by live bacterium and fluoresces yellow, and the LIVE/DEAD BacLight assay which can distinguish between live and dead bacterial cells.
Metabolic Assays
There are several types of metabolic assay available for biofilm analysis. For example, the resazurin assay involves resazurin which is taken up by cells and converted to a pink-fluorescent resorufin which can be analyzed through spectrophotometric analysis. This, therefore, allows the analysis of viable cell number within a biofilm.
Another example is the BioTimer assay, which is taken up by bacteria and converted from red to yellow following fermentative metabolism. The time taken for color change can then be used to analyze the size of the biofilm.
Genetic Assays
Real-time quantitive-reverse transcription PCR (qRT-PCR) is another method frequently used to analyze biofilms. This process monitors a PCR reaction throughout the amplification process, allowing the collection of real-time data. The fluorescent signal after each cycle can be used to indicate the amount of bacterial genetic information and the viability of all cells within the biofilm.
Another genetic assay used to monitor biofilms is fluorescence in situ hybridization, which involves the labeling of probes with fluorescent dyes. These probes can then be used to bind to a target within the biofilm – allowing identification of different species when paired with confocal laser scanning microscopy.
Physical Assays
There are many different microscopy techniques available to analyze biofilms, including, mass spectroscopy, confocal laser scanning microscopy, electron microscopy, x-ray microscopy, and scanning probe microscopy. These methods allow the biofilm to be imaged and therefore monitor biofilm processes in real time, as well as the components of different biofilms.
Overall, there are many different available tools for the analysis of biofilms. Each approach has different advantages and limitations and, therefore, utilizing a combination is the best approach to minimize errors and maximize the amount of information obtained from these complex structures.
Further Reading
Last Updated: Jan 23, 2019