IDPs are proteins that contain stretches of amino-acid sequence that are flexible and do not comprise stable structure in isolation. This is in contrast with a more traditional view of proteins as largely occupying a stable native structure that correlates with functions such as enzyme activity or binding.
Indeed, many prior studies on more complex systems worked with constructs containing truncations to remove disordered regions of the protein to facilitate crystallographic or other structural studies.
It is increasingly being recognized that in fact a sizable fraction of proteomes encode disorder, and that the special biophysics of these molecules plays important roles in cellular function and malfunction.
IDPs are involved in a variety of functions including as signaling hubs, multivalent interaction units, passive or active linkers, and entropic brushes. Recent work also shows that weak multivalent interactions by IDPs are important in the formation and dynamics of membraneless granules that are important in cellular function. They are also linked to diseases including neurodegenerative diseases (Parkinson’s and Alzheimer’s), cancer and diabetes.
The importance of these proteins has resulted in substantial interest from the biophysics and biology communities. Since the flexibility of these proteins also makes them hard to study using traditional structural biology methods, development and application of new methods or analyses (such as single-molecule experiments in our lab) is also an important line of work in the field.
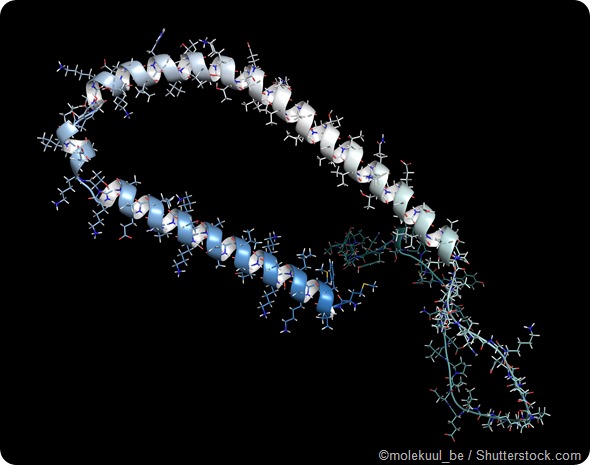
Is alpha-synuclein an important example of an IDP? How much was previously known about it?
Alpha-synuclein is an important example of an IDP that has been studied quite extensively for several years. It has been implicated in cellular functions, including involving synaptic and other vesicles, and has also been linked to Parkinson’s disease.
One of its interesting biophysical aspects is that it represents one of several IDPs that fold upon binding to partners such as membranes.
Partner associated folding is especially interesting with respect to switchable folding and consequent function or misfunction, with binding being able to even push the system into more than one folded form.
Can you please give an overview of your recent study that investigated how alpha-synuclein behaves when the membrane is crowded?
While a lot of work on protein biophysics is done in fairly dilute solution, cells have high concentrations of macromolecules that can take up room and “crowd” the solution. A lot of work has been done on crowding in three dimensions, with one idea being that crowding can result in stabilizing more compact forms of proteins.
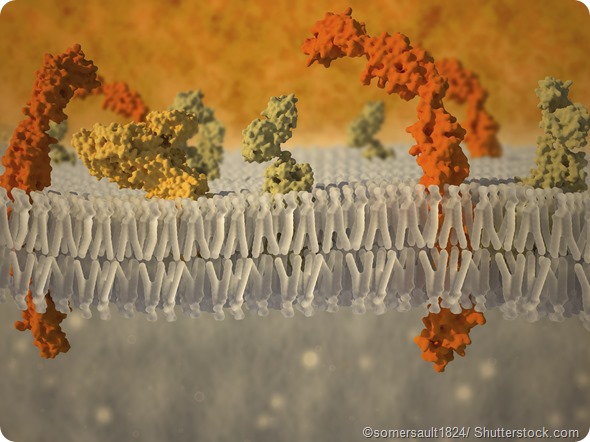
An interesting question that arises is what would happen if the protein had an additional dimension to “escape” to. We think we discovered just such a case while studying the influence of a small heat shock protein Hsp27 on the membrane binding and folding of alpha-synuclein.
Our results indicate that crowding on the (2-D) surface of a membrane can provide new layers of conformational complexity in membrane-binding IDPs. They also add an interesting illustration of the strengths of single-molecule methods for directly probing mechanistic complexity in biological systems.
What were your main findings and were you surprised by alpha-synuclein’s behaviour?
Dr. Priya Banerjee, a post-doc in the lab, was interested in the biophysics and function of a small heat shock protein Hsp27. Our initial set of findings for the system indicated that Hsp27 inhibited alpha-synuclein binding to membranes, by itself binding and blocking the membrane surface.
On closer examination though, the inhibition data surprisingly deviated from our expectations based on simulations of a simple mathematical model of the above process. Therefore, a graduate student Mahdi Moosa and Dr. Banerjee dug deeper into the system, and found that the simple model did in fact work, if we separately examined Hsp27 inhibition of the binding of two individual pieces of the alpha-synuclein helical region, which had very different individual binding affinities.
The results also indicated that combining effects of the two pieces in the full length protein was resulting in the observed deviation. Based on these observations, we predicted that for full length alpha-synuclein, Hsp27 could crowd the membrane and force the formation of a “hidden” state of alpha-synuclein, which is not very populated under uncrowded conditions. Hence, in the final key piece of the work, to test this model, we designed and carried out single-molecule FRET experiments, which directly revealed the hidden state.
What functions do you think this behaviour could perform?
The alternative structural state that we discovered could have altered binding and downstream functionality. In particular, the segment of alpha-synuclein that flips off the surface could interact with other proteins in solution, bringing them closer to the membrane for further functions.
More generally, such alterations of structure by crowding on membranes could result in altered interactions and other functions both on the membrane and in solution.
Are other IDPs that are bound to the cell membrane likely to also behave in this way?
The conceptual basis and mathematics of the 2-D crowding model are general. It is known that cell membranes are crowded objects. Additionally, binding of many IDPs is found to be modular. Hence, such crowding effects are likely to be important in other IDPs that bind to membranes and to result in new mechanistic bases for their tunable functions.
Did your research reveal any information about the role of alpha-synuclein in Parkinson’s disease?
The conformational properties and interactions (including aggregation and with membranes) of alpha-synuclein have been linked to Parkinson’s disease. Therefore, while not part of our current paper, our new understanding of factors that can tune complexity in alpha-synuclein biophysics could be relevant for its role in PD.
What are the next steps in your research?
Our results open up a few lines of research.
In the alpha-synuclein-membrane system, one next step is to understand how the observed crowding effects are altered by changes in lipid composition of the membranes or post-translational modifications, both of which are biologically important.
In addition, the possibility of binding of an additional partner to the N-terminal region in the hidden alpha-synuclein state and downstream function can be explored. Also, we would be interested in exploring crowding effects in other membrane-binding IDPs.
Considering the three-body problem of physics, extension from two to three interacting bodies enormously increases the complexity. Similarly, our current paper reveals how a three-component interacting system in biology can have interesting and complicated consequences. This idea can be extended to additional interacting components at molecular and larger scales, both of which occur in cells.
On the methodology side, we’ve been working on extensions of single-molecule FRET to multiple colors, useful for studying such multicomponent interactions. On a larger scale, we’ve recently used single-molecule FRET in initial studies aimed to understand protein structural features in protein droplets. These droplets are formed by phase separation, a process now believed to be important in the formation of several membraneless cellular organelles such as the nucleolus and nuclear speckles. Studies of these droplets and organelles is an area of high current interest.
Finally, we are also interested in other aspects of biological complexity. These include the response of protein systems to complicated input stimuli, important given that cells are non-equilibrium systems.
We recently explored the low-pass behavior of a model folding system (see Polinkovsky et al. below). We are interested in understanding how such complex stimuli affect multicomponent and crowded systems, again with relevance to the physics of the cell.
Where can readers find more information?
- Our just-published article: Banerjee PB, Moosa, MM & Deniz AA, “Two-dimensional crowding uncovers a hidden conformation of alpha-synuclein”, Angew Chem Int Ed Engl (2016) 55:12789; http://dx.doi.org/10.1002/anie.201606963; PMID: 27612332
- Lab website: http://www.scripps.edu/deniz
- General IDP review: Wright PE & Dyson HJ, “Intrinsically disordered proteins in cellular signalling and regulation”, Nat Rev Mol Cell Biol (2015) 16:18; http://dx.doi.org/10.1038/nrm3920; PMID: 25531225
- Single-molecule studies of IDPs and protein complexity: Lee T, Moran-Gutierrez CR & Deniz AA, “Probing protein disorder and complexity at single-molecule resolution”, Semin Cell Dev Biol (2015) 37:26; http://dx.doi.org/10.1016/j.semcdb.2014.09.027
- This paper discusses the molecular low-pass filter behaviour mentioned above: Polinkovsky ME, Gambin Y, Banerjee PR, Erickstad MJ, Groisman A & Deniz AA, “Ultrafast cooling revels microsecond-scale biomolecular dynamics” Nat Commun (2014) 17:5737; http://dx.doi.org/10.1038/ncomms6737
About Associate Professor Ashok Deniz
 Prof. Ashok Deniz earned his Ph.D. degree in (Physical Organic) Chemistry from the University of Chicago. He used sound and spectroscopic methods to study thermodynamic and kinetic features of mechanistically-important but very short-lived organic molecules.
Prof. Ashok Deniz earned his Ph.D. degree in (Physical Organic) Chemistry from the University of Chicago. He used sound and spectroscopic methods to study thermodynamic and kinetic features of mechanistically-important but very short-lived organic molecules.
He then began post-doctoral work at the University of California, Berkeley, where he was involved in early developments in the single-molecule FRET field. He joined the faculty of The Scripps Research Institute in 2000, where he established a single-molecule biophysics program.
Ashok and his lab have been involved in the development and biophysical applications of single-molecule tools, with a focus on FRET and combinations with microfluidics and chemical biology. Applications have included mechanistic studies of protein folding and the biophysics of protein disorder.
Recent directions of interest include probing complex non-equilibrium molecular dynamics, and extending studies to larger molecular assemblies such as during phase separation (of relevance to membrane-less cellular organelles).
Overall, the general goal of the Deniz Lab is to use novel tools to discover and understand predictive physical and chemical principles that underlie the biology of proteins and other biomolecules.