With the current state of knowledge regarding genetic testing, non-invasive prenatal testing (NIPT) is in favor for genetic testing in unborn babies, using fragments of DNA circulating in the mother’s blood.
Polymerase Chain Reaction (PCR) and Agarose gel electrophoresis is a method of gel electrophoresis used in biochemistry, molecular biology, genetics, and clinical chemistry in lab. Credit: Rattiya Thongdumhyu / Shutterstock
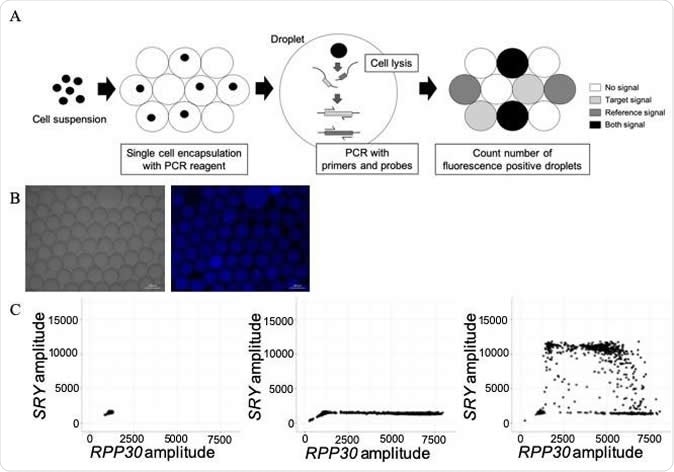
A) Schema of the single-cell-based droplet digital PCR (sc-ddPCR) system. Using the QX200 ddPCR system, each cell is simply encapsulated in one droplet. Up to 3,000 cells per well can be individually encapsulated. Subsequently, cell lysis and PCR with probes and primers are performed in each droplet. Finally, by quantifying the fluorescent droplets, single-cell genomic DNA is able to be assessed. B) Confirmation of single-cell encapsulation after generating droplets with a human B-cell line. Left, in bright field; right, with a Hoechst filter. C) Two-dimensional plot of signal amplitude of SRY and RPP30 probes in each droplet detected using our modified sc-ddPCR system. Signal amplitude of probes with 1× PBS as a blank control (left), a human female B-cell line (HEV0230) as a negative control (middle), and a human male B-cell line (HEV0057) as a positive control (right). Credit: Journal of Molecular Diagnostics
NIPTs – limitations
In contrast to invasive tests like amniocentesis which have the potential for fetal harm, the NIPTs in use today have many advantages including the absence of fetal risks, but they also have their limitations. One of the major shortcomings is that they sometimes yield non-useful data because they examine fragments of DNA, released from fetal or placental cells during their breakdown. These DNA bits are therefore called cell-free DNA or cfDNA, rather than the whole genome. This doesn’t always allow the analyst to know whether the DNA comes from the mother’s cells or the fetus. As a result, NIPT is a screening test and not a definitive diagnostic test.
Scientists are therefore always on the lookout for better NIPTs to make a conclusive determination of fetal genetic abnormalities. Most of these techniques depend on the use of cfDNA circulating in the mother’s blood at above 4 percent. Fetal cells in the mother’s blood are extremely rare due to the high effectiveness of the placental barrier, and this poses a formidable challenge to the analyst. To increase the chances of detection of the fetal DNA, many tests make use of the polymerase chain reaction (PCR). This is a method by which a single sample of DNA is multiplied into the millions, originally designed by American biochemist Kary Mullis, for which he was awarded the Nobel Prize.
The new technique
The current study used a modified form of the single cell droplet digital PCR (sc-ddPCR). Using some adjustments to the basic technique, they produced proof of concept that this can evaluate the genome of fetal cells found in extremely low concentrations in the maternal circulation, from a blood sample drawn from a peripheral vein. This could lead to the development of a test to assess the DNA of one living cell directly from maternal blood, avoiding the steps of cell fixation, cell staining and amplification of the whole genome.
Explains researcher Kenichiro Hata, “We have observed that in some cases NIPT results are discordant with the fetal genetic information that has been reported. This study serves as a proof of concept for non-invasive prenatal diagnosis using circulating fetal cells without any strict cell purification.”
The modification
With the sc-ddPCR, each droplet is in a well and contains up to 3,000 cells, all of which are analyzed at the same time. The original technique yields very specific and sensitive results from a single cell. The issue is that unless the cell suspension has a clear background, it doesn’t work well. To ensure this, the cell lines must be washed, or fluorescence-activated sorting of the cells performed.
The current modification obviates this need by providing a better PCR environment within the droplet, allowing higher sensitivity and specificity. This makes it possible to evaluate genetic information from a single cell based on a crudely purified sample containing nucleated cells.
The researchers took blood samples from 13 pregnant women and focused on using a crudely sorted cell suspension and sc-ddPCR techniques to identify the genomic DNA of a male fetus, from the fetal cells circulating in the mother’s blood. They found that they achieved high sensitivity in determining the presence of the SRY, or sex-determining Y gene, the chromosomal locus which regulates the beginning of sex determination in a male.
The proof
In other words, they found that the SRY gene was present only in the samples obtained from the 3/13 women who had male fetuses. This shows the high sensitivity and specificity of the modified sc-ddPCR system.
Implications
Once these findings are validated, by applying the technique to larger numbers of blood samples from pregnant women with respect to other genes, it could be extended to many other areas of application. Hata observes hopefully, “In the future, by optimizing cell sorting and encapsulation, as well as generating a more effective PCR environment in each droplet, this modified sc-ddPCR system may be a breakthrough analysis method that can be applied to various research realms and possibly to clinical diagnostic testing.”
Journal reference:
Direct Assessment of Single-Cell DNA Using Crudely Purified Live Cells: A Proof of Concept for Noninvasive Prenatal Definitive Diagnosis Sato, Taisuke et al. The Journal of Molecular Diagnostics, Volume 22, Issue 2, 132 - 140, https://jmd.amjpathol.org/article/S1525-1578(19)30431-3/fulltext