These viruses are called bacteriophages – literally, eaters of bacteria – and phages for short. While phages have been around for a long time, these new phages are startling in terms of their size and biologically complex systems. Many of their genes are usually found in bacteria and are used to attack their host bacteria.
The process
Where did the scientists find these rare huge phages? They pored over a vast database of DNA sequences obtained from a broad and diverse range of environments on earth. These almost 30 microbiomes spanned the spectrum from a hot spring in Tibet, through bioreactors in South Africa, to the intestine of a woman carrying a baby in her womb. Other locations include underground holes, oceans, lakes, and the gut of a preterm baby.
Researcher Jill Banfield has been doing this for over a decade and a half. She works out the sequences of all the DNA bits in any sample she gets from a location on earth. She then fits the puzzle pieces together to get the genomic sequence.
In most cases, she gets a draft of the genome, but a few have been confirmed to be the genomes of entirely new microbes. Some of these microbes are regulated by tiny genomes which, indeed, seem incapable of sustaining life independently, and instead are entirely dependent on other life forms, namely, archaea and bacteria.
Large-genome phages
It was a year ago that she reported the finding of large phages, which she dubbed Lak phages, in the human gut and mouth. These phages eat bacteria in the saliva and the gut.
In the new paper, she reports the isolation of over 350 very large phages with genomes that are, at lengths of 200,000 bases, fourfold bigger than the average genetic sequence of any bacteriophage known so far (50 kb). Banfield comments, "We are exploring Earth's microbiomes, and sometimes unexpected things turn up."
The largest sequenced so far has a genome that spans an astonishing 735,000 base pairs, which is about 15 times the size of the average phage genome, and in fact, markedly larger than many bacterial genomes. But there could be larger still since they have only sequenced 175 of these large phages until now.
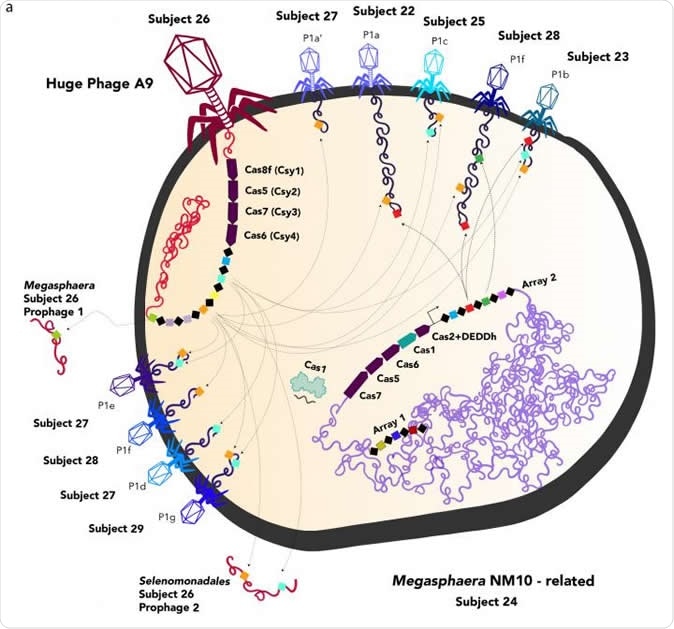
Depiction of huge phages (red, left) and normal phages infecting a bacterial cell. The huge phage injects its DNA into the host cell, where Cas proteins -- part of the CRISPR immune system typically found only in bacteria and archaea -- manipulate the host cell's response to other viruses. The UC Berkeley team has not yet photographed any huge phages, so all are depicted resembling the most common type of phage, T4. Image Credit: UC Berkeley image courtesy of Jill Banfield lab
A babble of phages
These large phages were classified into ten clades or groups. Each has been named "Big phage" though using different languages, or different words in the same language. The languages are those of the country of origin of the researchers. Thus there is a Mahaphage clade, a Biggiephage, a Judaphage, a Kaempephage, a Dakhmphage, a Jabbarphage and a Kabirphage, as well as an Enormephage, Whopperphage, and Kyodaiphage clade – in Sanskrit, Australian English, Chinese, Danish, Arabic (the next three), French, American English, and Japanese!
Bacterial weapons within viruses?
The bacterial genes within these phages contain some CRISPR elements, gene editing protein encoding segments, that are used by bacteria to resist viral attack. The scientists think these elements are meant to become part of the host's CRISPR system, empowering the bacteria to resist other viruses so that these large phages can enjoy their meal uninterrupted. Another researcher, Basem Al-Shayeb, explains that these phages use this system for their own benefit, "to fuel warfare between these viruses."
Another fascinating element in one of these novel phages is a protein that plays the same role as the Cas-9 protein, an integral part of the widely used CRISPR-Cas9 gene-editing tool first introduced by Doudna and Charpentier. This new protein has been called CasØ – the letter Ø, or phi (in Greek), is the symbol for a phage. This is part of the Cas-12 family, but these phages also have other bacterial CRISPR elements like Cas-9, Cas-X, and Cas-Y proteins.
Some of these phages also have very large CRISPR arrays. This refers to the presence of arrays composed of viral DNA fragments, memory arrays that allow the rapid recognition of a repeat attack by any of these viruses. This, in turn, triggers instant Cas reactions, enabling specific targeting of these viruses.
Life or non-life?
Another startling finding was the occurrence of genes encoding ribosomal proteins. The ribosome is the cell organelle that reads mRNA and uses it to make specific proteins. This is among the first occurrences of ribosomal genes in viruses.
They also found genes that code for transfer RNAs, which are the molecules that pick up the right amino acids to be incorporated into the protein sequence. There are regulatory genes for tRNAs, genes to switch on the protein synthesis process, and even a few ribosomal segments. These genes are found only in life forms since they have to do with the manufacture of protein components – which is unlike what any other virus can do, and which is a capability that spells life.
Investigator Rohan Sachdeva says this is "one of the major defining features that separate viruses and bacteria, non-life and life." He calls this "blurring the line a bit."
Why are these genes there? The researchers think they might be used to hijack the ribosomes to start replicating viral rather than host bacterial proteins. An additional finding is the presence of alternative genetic codes, or the use of more than one genetic code to denote the same amino acid. This feature could well throw the bacterial ribosome off track and fool it into decoding the viral RNA rather than the host's own RNA.
Implications
The discovery of new tools used in the fight between bacteria and viruses offers a lot of possibilities for finding new gene-editing tools. Many of the new genes are still to be explored for their functions, and it could well be that these new proteins will be helpful in various applications in industry, the medical sciences, or in agriculture.
There are dangers, too. Viruses transfer genes from one bacterium to another – some of which could be responsible for antibiotic resistance or for virulence (the capability to cause disease). This property of the newly discovered phages could mean that there is a risk of transferring some of these harmful genes into the human microbiome since phages occur alongside bacteria.
Since the number of genes carried by these large phages is markedly higher than ordinary phages, they have a larger capability to move around genes that can injure other cells. In turn, this increases the risk that some of these genes will be acquired by genes in the human environment.
Conclusion
Altogether, the researchers are fascinated by the presence of phages with huge genomes in a variety of microbiomes across the earth. The relationship between these large phages is interpreted as meaning they come of an ancient family of large-genome viruses.
Says Banfield, "Having large genomes is one successful strategy for existence. These viruses of bacteria are a part of biology, of replicating entities, that we know very little about."
True enough. These phages are of a size that makes them fit somewhere in the gap between the archaea (the earliest relatives of bacteria), and ordinary phages that look like non-living things. Banfield describes them as "successful strategies of existence that are hybrids between what we think of as traditional viruses and traditional living organisms."
Journal reference:
Al-Shayeb, B., Sachdeva, R., Chen, L. et al. Clades of huge phages from across Earth’s ecosystems. Nature (2020). https://doi.org/10.1038/s41586-020-2007-4