The ongoing COVID-19 pandemic has sparked many research projects to meet the challenge of the virus and come up with measures to fight its activity. A new study published in the journal Cell Host & Microbe in April 2020 describes a novel system to help scientists understand the virus, including mutations, diagnose infections, and also to develop and evaluate vaccines more quickly.
The team of scientists working at the University of Texas Medical Branch at Galveston has been working on engineering a reverse genetic system for the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) that is responsible for COVID-19 disease. This is among the most useful ways to uncover the working of a virus, because it allows the virus to be produced in the laboratory, in a petri dish culture where it can be studied in various ways.
.jpg)
Novel Coronavirus SARS-CoV-2 This transmission electron microscope image shows SARS-CoV-2—also known as 2019-nCoV, the virus that causes COVID-19. isolated from a patient in the U.S., emerging from the surface of cells cultured in the lab. Credit: NIAID-RML
Regular genetic vs. reverse genetics
RNA viruses can spread rapidly and cause severe or even fatal illness in animals and humans. The chief preventive weapon against such agents is effective immunization. To rapidly develop mutant viruses on which vaccines can be based, reverse genetics systems have developed, allowing viral genomes to be studied and manipulated.
The first application of reverse genetics to create an infectious viral particle was in 1981 when scientists generated the poliovirus from complementary DNA. Subsequently, all major families of viruses have been generated using this technology. They have also become the new face of vaccine design.
Vaccines are among the greatest contributions of science to human health. Traditional vaccine development techniques like live attenuation through repeated passaging or inactivation of viruses could be relatively inefficient at identifying promising candidates, however, in the face of a global threat. Advanced recombinant DNA methods coupled with reverse genetics, have thrown much light on RNA virus replication and pathogenesis, helping to accelerate vaccine development by allowing targeted modifications of the genome and directed attenuation.
Conventional genetics begins when a scientist is faced with an abnormal presentation and traces the gene responsible by genetic analysis, finally determining the DNA base sequence and the amino acid sequence and structure of the protein. With reverse genetics, recombinant DNA technology is brought into play, beginning with a protein or DNA and finding the mutant gene, ending up with the abnormal phenotype produced by the gene. For instance, a protein is translated backwards into the DNA sequence, which is then built up from the DNA bases in the laboratory. This is then used as a probe to detect the gene that is complementary to it. The gene is cloned, and various mutations introduced to detect the changes in the phenotype that result. Thus, reverse genetics informs the scientist about the normal role played by unknown DNA or protein sequences through in vitro mutagenesis and the resulting phenotypic changes.
The advantages of reverse genetics
The genetic system helps scientists to look at the way the novel coronavirus has changed its genome, which in turn determines its ability to cross-species from the bat to humans. It is still unclear whether this switch requires an intermediate host for the virus.
The use of the reverse engineering system has made it possible to obtain a form of the virus that is labeled with neon green so that any cell into which it gains entry glows green. This will help researchers to rapidly identify patients who have been infected by the new coronavirus.
The reverse genetics system can also assess the antibody response to a vaccine under development, specifically those antibodies that block viral entry into the cell. The level of antibody production in response to vaccination is the leading indicator of vaccine efficacy, according to the scientists. Being able to measure antibody concentration could thus speed up the process of vaccine testing.
Traditional methods of testing for viruses can run only a few samples at a time and require about a week to return a result. In contrast to this, says researcher Pei-Yong Shi says, "The neon green-labeled virus system allows us to test patients' samples in 12 hours in a high-throughput manner that tests many samples at once."
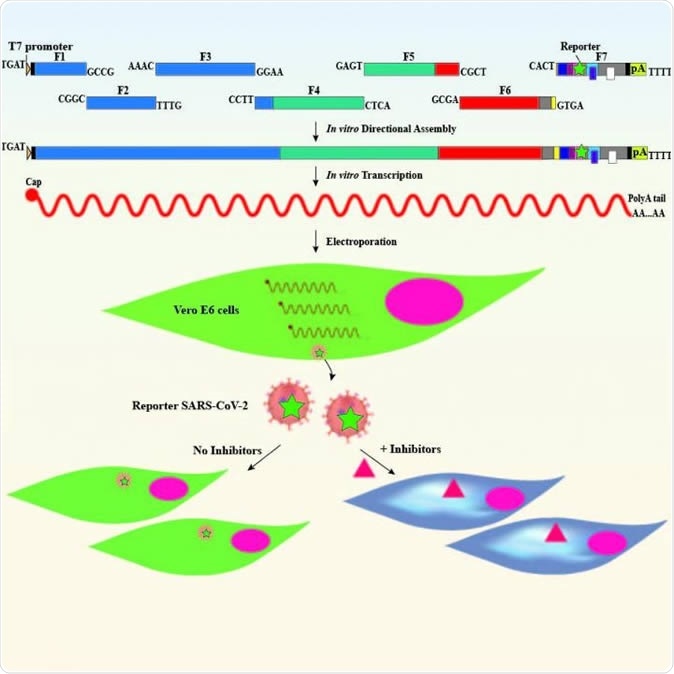
Graphic representation of the reverse genetic system for SARS-CoV-2. Image Credit: The University of Texas Medical Branch at Galveston
What does the technology mean?
The brain behind the reverse genetic system, Xuping Xie, comments on the advantage the developing technology gives to vaccine developers in terms of evaluating vaccines and cutting down on time to market. The UTMB researcher says, "UTMB will be very happy to make this technology widely available to both academia and industry researchers working to develop countermeasures quickly."
This is only one more instance of the importance of teamwork in fighting disease. Several teams of scientists pooled their expertise to develop the system. The researchers say they will expand the team science to areas of clinical care and patient diagnosis by developing serological testing, using the new technology.
While describing the reverse genetic system as a "critical tool for the research community", the researchers also say, "It's critically important to have a system that can be used for any new future or re-emerging viruses so that we can very quickly respond to the pathogens and protect peoples' health."
Journal reference:
An infectious cDNA clone of SARS-CoV-2, Cell Host & Microbe, Xuping Xie, Antonio Muruato, Kumari G. Lokugamage, Krishna Narayanan, Xianwen Zhang, Jing Zou, Jianying Liu, Craig Schindewolf, Nathen E. Bopp, Patricia V. Aguilar, Kenneth S. Plante, Scott C. Weaver, Shinji Makino, James W. LeDuc, Vineet D. Menachery, Pei-Yong Shi, Journal pre-proof, https://marlin-prod.literatumonline.com/pb-assets/products/coronavirus/CHOM_2291_s50_preproof.pdf