With data obtained from the Virus Watch study in England and Wales, researchers found that the serial interval of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is broadly similar to estimates from previous studies and there is no evidence that the B.1.1.7 variant is associated with any change.
Serial interval is a key parameter in epidemiological studies of infectious diseases. It is an estimation of the spread of the disease and is often measured as the time interval between symptom onset in a primary case and the onset of symptoms in the secondary cases infected by the primary case. Serial interval is an important parameter used in disease transmission models and can help guide control strategies.
Apart from the serial interval, the doubling time of disease also depends on the reproduction number (R). Diseases that have a shorter serial interval but similar R values will have shorter doubling times. The serial interval for diseases varies widely, with it being 2.8 days for H1N1 influenza, 11.7 days for measles, 14 days for chickenpox, and 18 days for mumps.
For COVID-19, studies report the serial interval to be between 4.2 and 7.5 days. It seems that several new SARS-CoV-2 variants have increased transmissibility, possibly due to higher R or shorter serial intervals.
The B.1.1.7 strain, first detected in the United Kingdom, has rapidly spread around the globe. However, there is no understanding yet of how the serial interval for this strain differs from other circulating strains.
In a new study published on the medRxiv* preprint server, researchers analyzed data and explored if the emergence of the B.1.1.7 strain affected the serial interval of COVID-19.
Estimating serial interval
The team used data from the Virus Watch study, an online study following entire households in England and Wales during the COVID-19 pandemic. Participants completed daily symptom diaries and completed a survey every week reporting any symptoms and outcomes of any COVID-19 test. In households, pairs of patients who were initially infected and then infected the other were identified.
The team used an estimation of the percentage of B.1.1.7 cases in different geographical locations at different times, regardless of whether or not this strain was known to be involved.
The team found that about 25,000 illnesses were reported from about 10,000 households between June 2020 and March 2021. Of these, 915 tested positive for COVID-19, and the team identified 287 possible infector-infectee pairs.
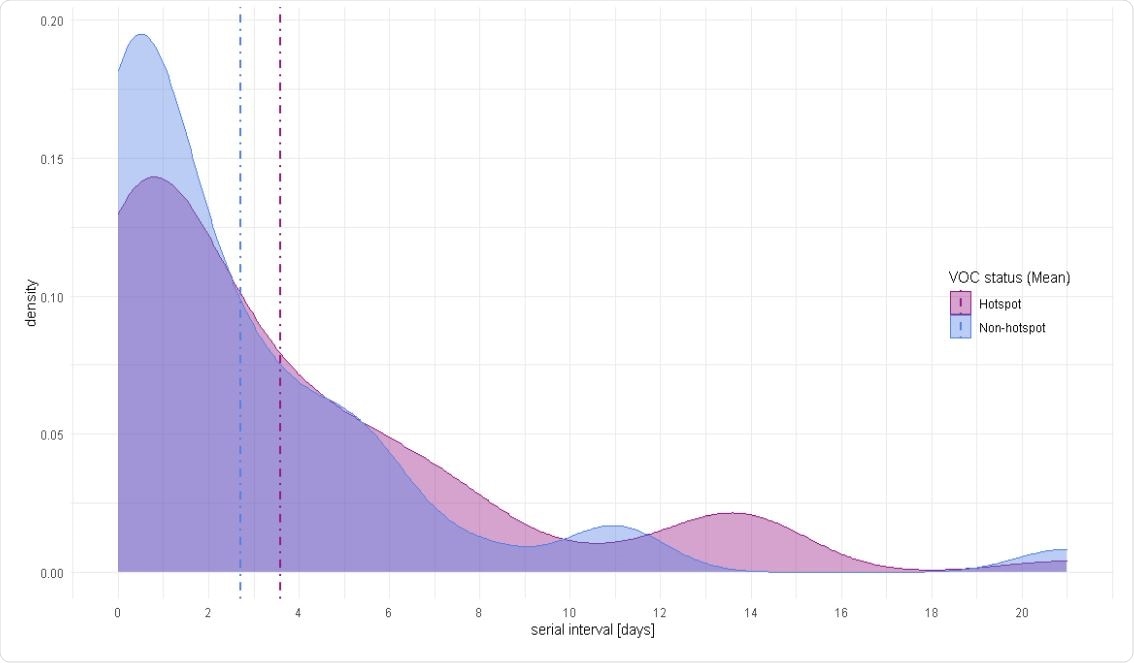
Serial interval density distribution for B.1.1.7 hotspot areas and non-hotspot areas (Mean).
From this data, the authors determined the majority of the serial interval to be between 0 and 8.5 days, with a peak at 0.5 days and a mean of about 3 days. There was no difference in the mean serial intervals between variant of concern hotspots and other non-hotspot regions. The shorter serial intervals may likely be because of close contact between household members, leading to faster infection and shorter serial interval times.
Thus, the serial interval determined here is within the ranges reported before, but a little lower than the estimates reported early in the pandemic. As the pandemic progressed, more and better preventative measures were implemented, thereby reducing the risk of infection.
The study had a large number of participants, but samples were taken during the symptomatic testing program, so some infections may have been missed.
“Our analysis does not provide evidence to suggest that changes in serial interval explain the rapid emergence of B.1.17,” writes the authors. There may likely be other explanations such as increased viral load or enhanced receptor binding that could explain the increased transmission and require further study.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Geismar, C. R. et al. (2021) Serial interval of COVID-19 and the effect of Variant B.1.1.7: analyses from a prospective community cohort study (Virus Watch). medRxiv, https://doi.org/10.1101/2021.05.17.21257223, https://www.medrxiv.org/content/10.1101/2021.05.17.21257223v1
- Peer reviewed and published scientific report.
Geismar, Cyril, Ellen Fragaszy, Vincent Nguyen, Wing Lam Erica Fong, Madhumita Shrotri, Sarah Beale, Alison Rodger, et al. 2021. “Household Serial Interval of COVID-19 and the Effect of Variant B.1.1.7: Analyses from Prospective Community Cohort Study (Virus Watch).” Wellcome Open Research 6 (December): 224. https://doi.org/10.12688/wellcomeopenres.16974.2. https://wellcomeopenresearch.org/articles/6-224/v2.