Nascent polypeptide chains or polypeptidyl-tRNAs (pep-tRNAs) occur transiently during protein synthesis. The potential to study these intermediates and better understand their role in processes like gene regulation has been greatly enhanced by the development of a process termed PETEOS—short for peptidyl-tRNA enrichment using organic extraction and silica adsorption. This method, developed by scientists at Tokyo Tech, allows for the large-scale harvesting, processing, and identification of pep-tRNA polypeptide moieties.
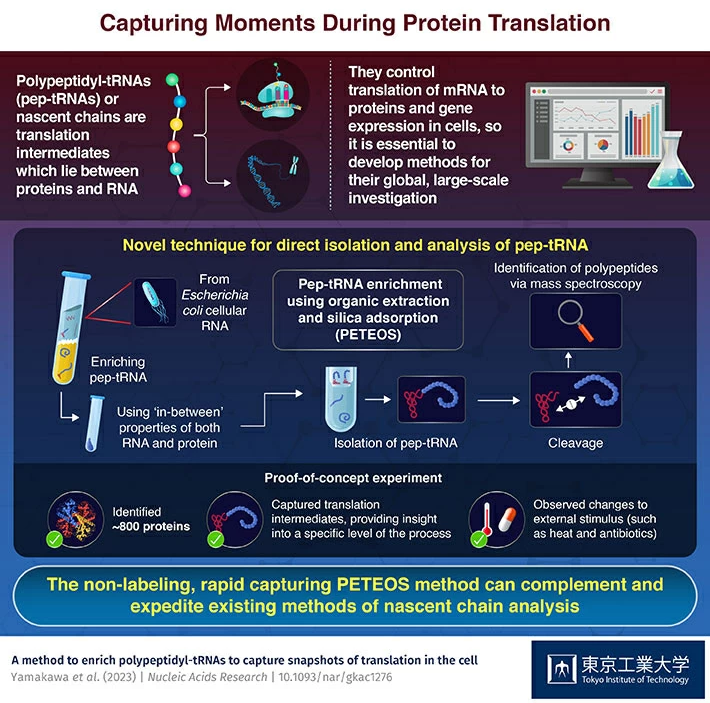
Image Credit: Tokyo Institute of Technology
Advances in molecular biology have revealed that pep-tRNAs—nascent polypeptides inside the ribosome that are covalently attached to transfer RNA—are involved in a myriad of cell functions, including gene expression. All proteins exist as pep-tRNAs at some point and studying these translation intermediates is vital as they possess properties of both RNA and protein and can help researchers better understand the specifics of translation. Depending on stimuli and/or stresses, translational regulation is very rapid and spans initiation, elongation, and elongation pausing. Garnering deeper insights into the process of translation therefore requires a suitable method to process pep-tRNAs in large quantities. These nuances have fueled the development of molecular tools to investigate cellular translation.
Presently, the two primary approaches being used to survey cellular translation leverage vastly different strategies. The first, ribosome profiling, a powerful deep-sequencing technology, monitors translation by targeting ribosome-protected mRNA fragments produced from RNase digestion. Unfortunately, this approach is inherently unable to capture pep-tRNAs, is time consuming, expensive, and data intensive. The alternative relies on proteomics with liquid chromatography-tandem mass spectroscopy (LC-MS/MS) and requires using unusual amino acids to label polypeptides. However, such proteomics approaches are time intensive, and, most importantly, do not capture the translation elongation status of the cell, as pep-tRNAs aren't directly targeted.
To navigate around these limitations in 'pep-tRNA-ome' technology, a team of scientists from Tokyo Institute of Technology (Tokyo Tech), led by Prof. Hideki Taguchi, developed PETEOS. Their findings have now been published in Nucleic Acids Research. Regarding the main advantages of their work, Prof. Taguchi explains, "This non-labeling methodology is unique in that it specifically enriches and rapidly captures pep-tRNAs while still complimenting conventional translatome analyses platforms. "
PETEOS has four steps. First, RNA is enriched using organic solvent extraction and then isolated on a silica column. Next, the target polypeptides in the isolated pep-tRNAs are separated from their ribonucleotides using a combination of high pH and temperature. Then, the liberated polypeptides are subjected to enzymatic digestion and cleaved. And lastly, these polypeptides are identified using shotgun LC-MS/MS proteomics. "As a proof-of-concept, we were able to identify nearly 800 E. coli proteins that were derived from pep-tRNA using PETEOS. Importantly, these proteins were N-termini enriched which underscores that the method really does capture the intermediate pep-tRNA pool during translation. ", says Prof. Taguchi when quizzed on the significance of their new method.
The team was also able to show that PETEOS could capture "screenshots" of the nascentome—the pool of nascent pep-tRNAs present at any given time—when E. coli cells were subjected to heat shock and antibiotic treatments. PETEOS outperformed conventional methods in terms of the capacity to capture acute protein expression under heat stress, and the rearrangement of the nascentome following exposure to different antibiotics.
PETEOS does have a few limitations, including a difficulty in analyzing low abundance pep-tRNAs using LC-MS/MS, limited quantitative capacity because of the acidic phenol/chloroform step, no codon-level resolution, and difficulty capturing the C-terminal end of the growing polypeptide. However, some of these issues exist for conventional approaches as well and can be overcome by improving the C-terminal peptide enrichment efficiency of this process.
The team believes there is tremendous potential to optimize their protocol moving forward.
Despite the limitations, we're confident PETEOS has potential as it enables analyses that current translatome methods simply cannot. Most importantly, it can be applied to any organism! "
Prof. Hideki Taguchi, Tokyo Tech
With more research in this field, we will deepen our understanding of protein synthesis.
Source:
Journal reference:
Yamakawa, A., et al. (2023) A method to enrich polypeptidyl-tRNAs to capture snapshots of translation in the cell. Nucleic Acids Research. doi.org/10.1093/nar/gkac1276.