Chromatrap® offers a one-of-a-kind solid state patented technology, which is characterized by unmatched sensitivity. This makes it possible for users to perform Chromatin immunoprecipitation (ChIP) assays using only 1000 cells per immunoprecipitation.
In the past, ChIP techniques needed a large number of cells, which was difficult when there was limited starting material. Sometimes samples are small, cell types can be hard to come by or are difficult because they offer only a low chromatin yield. Such situations have prompted the development of several techniques which are able to utilize these sources of chromatin.
A large number of available techniques are complicated and time consuming. In order to make ChIP efficient from lower cell numbers, some of the techniques include spiking of samples with chromatin from another source, or DNA amplification before downstream analysis.
Using the ChIP-on-chip method and without amplification or spiking, MicroChIP (µChIP) from 10,000 cells has been carried out (Acevedo et al., 2007). The µChIP technique can be changed so that it uses qPCR as the downstream process, which was successfully performed (Collas, 2011). However, this type of ChIP process is still lengthy and demanding.
There are alternative methods which have had success with ChIP using only a few cells. These methods are specifically used for targets which are high abundant, and require more than three days (Dahl and Collas, 2007; Sachs et al., 2013).
We will now show Chromatrap®’s exceptional sensitivity and wide dynamic range, which makes it possible to carry out ChIP using only 1000 cells. Excellent real signal is obtained, regardless of starting cell number and with no difference in terms of enrichment.
Methods
Chromatin preparation
For the study the human endometrial carcinoma cell line Hec50 (Holinka et al., 1996) was utilized for all extracted chromatin. Standard protocols were used to culture cells, which were allowed to reach appropriate numbers.
Three types of plates were used: 96-well plates for 1000 cell chromatin, 12-well plates for 10,000 cells, and 6-well plates for 100,000 cells. A TC-10 Automated cell counter (BioRad) was used to count cells on representative wells of all culture vessels.
A suitable amount of 1% formaldehyde was used to fix cells in. For 1000 cells a quantity of 100 µl was used; for 10,000 cells 1 ml, and for 100,000 cells 3 ml. The formaldehyde was then decanted and 0.65M glycine added, thus quenching each monolayer of cells.
Afterwards the cells were scraped into ice-cold PBS ranging in size: 1 ml, 400 µl or 100 µl corresponding to 6-, 12- and 96-well plates respectively. Using centrifugation, the cell pellets were collected. Cell membranes were lysed in 100 µl Chromatrap® Hypotonic Buffer followed by nuclear lysis in 100 µl Chromatrap® Lysis Buffer. A Qsonica sonicator on high setting was used to shear the chromatin stock for 15min: 30 seconds on, 30 seconds off to generate 100-500bp fragments.
Chromatin immunoprecipitation
The Chromatrap® ChIP-seq Protein A kit (Cat no 500189) was used for ChIP in accordance with the protocol supplied in the kit. Slurries were prepared for immunoprecipitation using 100 µl chromatin stock from each extraction, depending on the number of cells: 1000, 10,000, 100,000.
Enrichment was carried out using 2 µg H3K4me3 antibody(Chromatrap® Cat. No. 700010) or 2 µg rabbit IgG (Chromatrap® Cat. No. 700014), the latter was used to ascertain nonspecific background binding . As an input for downstream processes, an equivalent amount of chromatin for each IP was set aside.
After reverse crosslinking and Proteinase K digestion, both the ChIP samples and inputs were purified using the Chromatrap® DNA purification columns, which are supplied in the Chromatrap® ChIP-seq kit.
To check the effectiveness of the precipitation of two gene loci, qPCR was utilized. Chromatrap® primer sets were used: for a positive gene target, GAPDH (Chromatrap® Cat. No. 800000), and for a negative gene, ß-globin (Chromatrap® Cat. No. 800010).
Results
For all cell numbers used in the test, excellent signal was obtained in terms of occupancy of the histone mark H3K4me3 onto the target gene GAPDH (Figure 1).
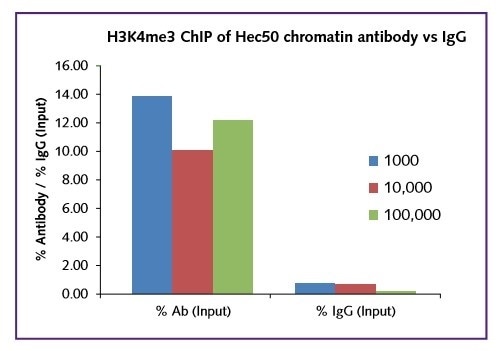
Figure 1: GAPDH positive gene target only, H3K4me3 vs IgG.
Regardless of the starting cell number (1000, 10,000, 100,000), the real signal was comparable within the range 10-14% for this target. In all cases a low background IgG in was recorded. The low binding of H3K4me3 to the negative gene target ß-globin further shows the selectivity of the assay (Figure 2).
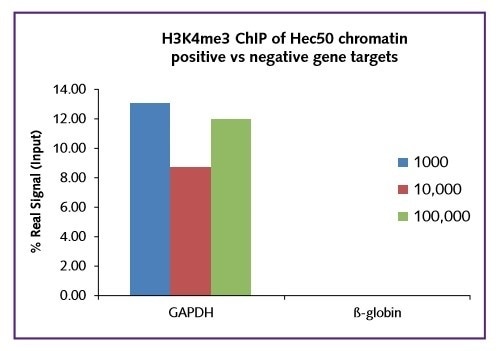
Figure 2: GAPDH positive and β-globin negative.
Conclusion
The Chromatrap® ChIP-seq Protein A kit is sensitive, specific and flexible, which enables robust and reproducible target enrichment using as few as 1000 cells per IP. The signal strength achieved is superb regardless of the number of cells used per IP, while background noise is reproducibly low.
References
- Acevedo, L. G., Iniguez, A. L., Holster, H. L., Zhang, X., Green, R., and Farnham, P. J., (2007). Genome Scale ChIP-chip analysis using 10,000 human cells. Biotechniques. 43, 791-797.
- Collas, P., (2011) A chromatin immunoprecipitation protocol for small cell numbers. Methods in molecular biology (Clifton, N.J.) 791: 179-193.
- Dahl, J.A and Collas, P, (2007) Q2ChIP, a quick and quantitative chromatin immunoprecipitation assay, unravels epigenetic dynamics of developmentally regulated genes in human carcinoma cells. Stem Cells. 25(4):1037-46.
- Holinka, C. F., Hata, H., Kuramoto, H., Gurpide, E., (1986). Responses to estradiol in human endometrial adenocarcinoma cell line (Ishikawa). J. Steroid Biochem. 24:85-89.
- O’Neill, L. P., Vermilyea, M. D. And Turner, B. M. (2006) Epigenetic characterisation of the early embryo with a chromatin immunoprecipitation protocol applicable to small cell populations. Nat. Genet. 38, 835-841.
- Sachs, M., C. Onodera, K. Blaschke, K.T. Ebata, J.S. Song & M. Ramalho-Santos, (2013) Bivalent Chromatin Marks Developmental Regulatory Genes in the Mouse Embryonic Germline In Vivo. Cell Reports 3: 1777-1784.
About Chromatrap
Chromatrap® is a product of Porvair Sciences, a wholly owned subsidiary of Porvair plc. We are one of the largest manufacturers of Ultra-Clean microplates, 96 well well filtration plates and Microplate handling equipment for life science and synthetic chemistry.
With offices and Class VIII clean room manufacturing located in the UK, combined with a world-wide network of distributors and dedicated distribution hub in the USA, we pride ourselves on our continuous innovation, research and flexibility to meet customer demands. We offer OEM production and contract manufacturing through our North Wales facility.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.Net which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.