Introduction
Two major options are available that can be used for preparing chromatin during the chromatin immunoprecipitation (ChIP) process. These include using a chemical agent like formaldehyde (X-ChIP) to crosslink the chromatin, or using native ChIP (N-ChIP) for non-cross linking the chromatin.
Based on the scientist and scientific queries being inquired, Chromatrap has been shown to be entirely flexible. Table 1 shows the pros and cons of each technique - native ChIP and cross-linked ChIP. Figure 1 shows the Chromatrap® native ChIP protocol.
Table 1. Comparison of Chromatrap®’s native vs cross-linked ChIP kits
|
Native ChIP
|
Cross-linked ChIP
|
|
ADVANTAGES
|
DISADVANTAGES
|
ADVANTAGES
|
DISADVANTAGES
|
|
Suitable for investigating histone marks and abundant targets.
|
Slightly longer due to overnight dialysis.
|
ChIP carried out in 5 hours!
|
Chemical fixation required.
|
|
No chemical fixation of cells; cells remain in a more natural ‘native’ state.
|
Can only shear chromatin enzymatically.
|
Can shear chromatin by either sonication or enzymatically.
|
Cannot capture chromatin from cells in their ‘native’ state.
|
|
In some cases there is increased affinity of antibody binding to antigens on native chromatin as it is more accessible.
|
Some low abundant transcription factors will not be detected.
|
Can investigate a wide range of histone marks and transcription factors.
|
Cross-linked chromatin can occasionally mask epitopes of some antibodies, affecting antibody/ chromatin binding.
|
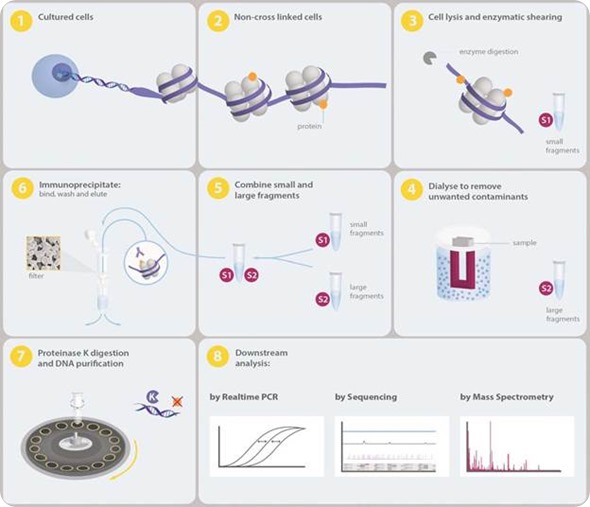
Figure 1. Overview of the Chromatrap® Native ChIP process
For this analysis, EZH2 and H3K27me3, were selected to emphasize the capability of Chromatrap®’s Native ChIP Kit and the Chromatrap® spin column ChIP kit for qPCR for selective enrichment of low abundant transcription factors and histone marks. EZH2 is a methyltransferase enzyme and is an enzymatic subunit of the Polycomb Repressive Complex 2 (PRC2) related to repressive gene transcription.
Methyltransferase is capable of regulating the methylation condition of the H3K27me3 histone mark. When EZH2 is abnormally expressed, it leads to increased methylation of H3K27me3 and this, in turn, is linked to different types of cancers (Yoo & Henninghausen, 2012).
Method
Chromatin preparation
Hec50 was used for preparing chromatin for both X-ChIP and N-ChIP methods, according to individual Chromatrap® process.
Experimental design
The X-ChIP as well as the N-ChIP method offers a powerful and sensitive platform for ChIP, and both can increase the signal for low abundant transcription factors and histone marks, thus emphasizing the Chromatrap® technology’s sensitivity. To show the reproducibility of the ChIP assay kits supplied by Chromatrap®, ChIP’s were performed in triplicate.
Antibody and gene targets
The EZH2 epigenetic mark is part of the PRC complex related to the repressive gene transcription. It contributes to the introduction of methyl groups to the H3K27me3 histone methylation mark, and this increased methylation on histone H3’s lysine 27 not only helps in suppressing transcription, but also aids in turning the genes off. Increased methylation levels in cancer can also turn off the tumor suppressor genes.
The Myelin Transcription Factor 1 or MYT1 gene is typically related to repressive transcription and heterochromatin antibodies, and is a suitable positive target for these types of antibodies. Actively expressed in all types of cells, glyceraldehyde-3- phosphate (GAPDH) is a valid negative gene target for these epigenetic marks. In order to show the low non-specific background binding acquired with the Chromatrap® technology, non-specific IgG antibody from the same species was also included along with the appropriate negative gene targets.
Immunoprecipitation
N-ChIP
Slurries were prepared with a 5:2 chromatin:antibody ratio. Inputs containing 5µg chromatin were prepared simultaneously and not subjected to ChIP enrichment but are used in the downstream analysis. Slurries were incubated on an end-to-end rotator, at 4°C temperature for 1 hour prior to the addition onto the column.
After removing unbound DNA through washing, the uniquely formulated Native ChIP Elution Buffer was used to elute selectively enriched DNA associated with the protein of interest. Before qPCR analysis, the samples were digested by the proteinase K enzyme.
X-ChIP
Slurries were prepared using a 2:1 chromatin:antibody ratio. Inputs were prepared concurrently comprising 1μg chromatin and were utilized for downstream analysis, but not enriched with ChIP. Slurries were loaded onto the Chromatrap® columns and incubated at 4°C temperature on a rocker for a period of 1 hour.
Unbound DNA was removed by washing and subsequently eluted by means of a ChIP Elution Buffer. Before qPCR analysis, the samples were subjected to the reverse cross-linking process and digested by the proteinase K enzyme.
qPCR Analysis
qPCR was applied for examining GAPDH and MYT1 specific DNA fragments related to immunoprecipitated antibodies, H3K27me3 and EZH2. Each precipitation assay was completed using an equivalent loading of IgG antibody as a negative control. The percentage of the input chromatin was normalized using the signal generated by non-specific binding of unspecific IgG. Error bars represent the standard error of the mean of the triplicate ChIPs. Both primer sets were used at an optimum annealing temperature of 60°C and at a working concentration of 4mM.
Results
It was observed that H3K27me3 enrichment on the MYT1 positive gene locus is similar for X-ChIP (Figure 2) as well as for N-ChIP (Figure 4). Through low enrichment for the GAPDH negative gene target (Figures 2 and 4) and the non-specific IgG antibody (Figure 3), good signal to noise is respectively shown for the H3K27me3 and the EZH2 low abundant transcription factor.

Figure 2. X-ChIP H3K27me3 onto Myt1/GAPDH: Low signal is produced by the negative gene target GAPDH when H3K27me3 is enriched using the qPCR Standard Chromatrap® Spin Column Kit.
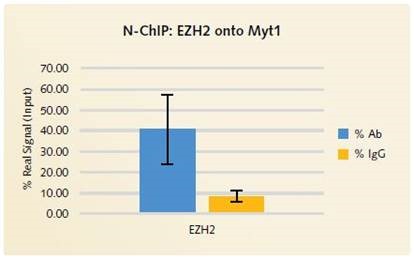
Figure 3. N-ChIP EZH2 onto Myt1: % Ab vs % IgG: Excellent signal to noise is demonstrated with the Chromatrap® Native ChIP Kit when specific antibody enrichment is compared to the enrichment of non-specific IgG.
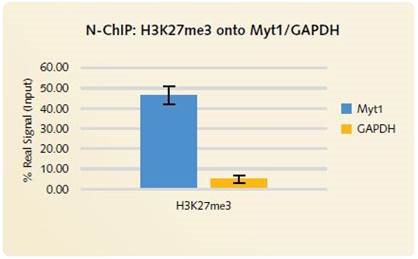

Figure 4. N-ChIP H3K27me3 onto Myt1/GAPDH: High levels of enrichment is produced using the Chromatrap® Native ChIP Kit onto the positive gene target MYT1. Low signal is produced for the negative gene target GAPDH, highlighting the highly specific enrichment of H3K27me3 onto the positive gene loci.
Using the Chromatrap® technology, low background signal was again observed with the cross-linked ChIP (Figure 2). Figure 4, highlights high signal for the positive MYT green targets compared to low signal for the negative GAPDH gene target.
Conclusion
This article has shown the variation between cross-linked and native ChIP, thus enabling users to choose the most appropriate method for their studies. In comparison with the non-specific IgG enrichment and the negative gene target for the qPCR standard Chromatrap® Spin Column Kit and the Native ChIP Kit, excellent positive enrichment has been shown clearly. With the aid of the Chromatrap® Native ChIP Kit, chromatin can be prepared without any necessity for aggressive chemical fixation.
Thanks to the Chromatrap® technology, it is now possible to use native chromatin for analyzing certain transcription factors and histone modifications, including the low abundant transcription factor EZH2 as illustrated in this analysis. This was accomplished with a low chromatin concentration, showing the high sensitivity and excellent specificity of the Chromatrap® Native ChIP Kit.
Chromatrap® has expanded its product line with the inclusion of the Native ChIP Kit. The products provide a variety of options through which scientific users can select the most optimum method for preparing their respective chromatins.
References
- Yoo, K. H. & Henninghausen, L. (2012). EZH2 Methyltransferase and H3K27 Methylation in Breast Cancer. Internation Journal of Biological Sciences. 8(1): 59-65.
About Chromatrap®
 Chromatrap® is a product of Porvair Sciences, a wholly owned subsidiary of Porvair plc. We are one of the largest manufacturers of Ultra-Clean microplates, 96 well well filtration plates and Microplate handling equipment for life science and synthetic chemistry. With offices and Class VIII clean room manufacturing located in the UK, combined with a world-wide network of distributors and dedicated distribution hub in the USA, we pride ourselves on our continuous innovation, research and flexibility to meet customer demands. We offer OEM production and contract manufacturing through our North Wales facility.
Chromatrap® is a product of Porvair Sciences, a wholly owned subsidiary of Porvair plc. We are one of the largest manufacturers of Ultra-Clean microplates, 96 well well filtration plates and Microplate handling equipment for life science and synthetic chemistry. With offices and Class VIII clean room manufacturing located in the UK, combined with a world-wide network of distributors and dedicated distribution hub in the USA, we pride ourselves on our continuous innovation, research and flexibility to meet customer demands. We offer OEM production and contract manufacturing through our North Wales facility.
Our porous polymeric material, BioVyon™, whose chemical functionalisation can endow it with internal surface properties individually configured to capture and separate target species out of difficult mixtures, has opened up many possibilities in the field of BioSciences where molecules of interest such as DNA, RNA, proteins etc can be selectively pulled out of complex mixtures of biological origin. The materials have proven to be a remarkably good substrate for accepting novel chemistries such as the organically bound Protein A and Protein G in Chromatrap®.
Using our 25 years experience of microplate manufacturing, Porvair Sciences has now developed a high-throughput bead-free ChIP assay based on our filtration plates containing our Chromatrap chemistry. Chromatrap-96 enables large scale epigenetic screening to become a reality in many laboratories and eliminates many of the long and laborious steps previously undertaken in such work.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.Net which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.