Myc is one of the most frequently mutated, widely studied, and overexpressed genes implicated in cancer.
Most mutations in the myc gene lead to its constitutive expression. The myc gene codes for the myc protein which is a transcription factor. Myc transcription factor leads to upregulation of several genes which are involved in cell cycle progression, apoptosis, and cellular transformation, leading to increased cell number and thus, cancer. This gene is present on chromosome 8 and regulates almost 15% of all genes.
In mammals three proteins, c-Myc, N-Myc, and L-Myc have reasonable structural resemblance and are encoded by three different genes, and their over expression reportedly leads to tumor induction. Myc protein contains bHLH (basic helix-loop-helix) structural and LZ (leucine zipper) motifs.
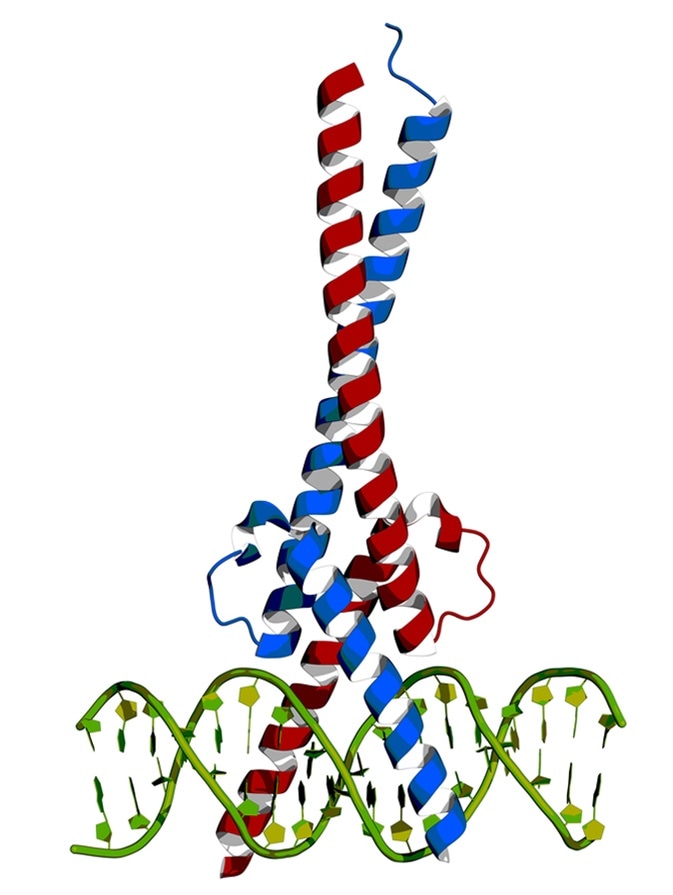
c-Myc and Max transcription factors bound to DNA. Image Credit: Petarg / Shutterstock
Hotspot Mutations in C-Myc
The coding sequence of c-myc bears several non-conserved hotspot mutations which can affect the function of the produced protein. For example, myc function can be inhibited by protein p107. However, mutations in the N-terminal region of myc can inhibit the p107 from performing its inhibitory effect, leading to activation of myc.
Another c-myc mutation hotspots is threonine-58 (Thr-58) which may either be phosphorylated or O-linked glycosylated. Phosphorylation of Thr-58 is essential for proteosome-mediated degradation of c-myc and mutation of Thr to Ala leads to myc protein stabilization. Activated RAS, which cooperates with c-myc during oncogenesis, inhibits the phosphorylation of Thr leading to stabilization of c-Myc protein. Thus, mutations at or around Thr-58 can lead to increase in c-myc levels and myc-mediated regulation of transcription.
Apart from mutations, c-myc gene amplification is also found in human breast, lung, ovarian and prostate carcinomas. Myc amplification is frequently observed in more aggressive breast cancers and correlates with poor prognosis and distant metastasis.
Chromosomal Translocations in Myc
Myc gene was first discovered in human Burkitt’s Lymphoma, a cancer of lymphatic system or B cells in particular. In 1983, Professor Jerry Adams and Professor Suzanne Cory, among others, discovered that Burkitts’s lymphoma involved chromosome translocations in the ‘myc’ gene, where c-myc moves from chromosome 8 to chromosome 14. The translocation region in chromosome 14 is near the enhancers of antibody heavy chain genes. Although this translocation into the immunoglobulin heavy chain locus occurs in about 80% of Burkitt's lymphomas, variant translocations into either the κ or λ light chain locus also occur at a frequency of about 10%. Translocation breakpoints can be divided into three classes in Burkitt's lymphomas: within the c-myc gene (class I), immediately 5′ to the c-myc gene (class II), and distant translocations (class III). Class III breakpoints are more than 100 kb from the c-myc gene. These classes are defined based on their location with respect to c-myc. The c-myc gene comprises three exons: exon 1 contains two promoters and is noncoding; exons 2 and 3 encode the myc protein. Class I translocation occurs within exon 1 and first intron, while the class II translocation occurs a few kilobases 5’ into the first exon. These translocations can disrupt the coding and regulation of c-myc.
SNPs in Myc
SNPs are one of the most frequent variations in the human genome, with >106 DNA variations between two individuals. Genome wide association studies (GWAS) is a common technique, used to map linkage between SNPs and cancer. For example, SNP rs13281615 is linked to breast cancer risk but not prostate or colon cancer. The 8q24 region of the genome (eighth chromosome, q arm) was found to contain more risk associated SNPs than anywhere in the genome. ‘Odds ratios’ are used to represent the risk associated with a disease where the ratio represents the odds of the disease occurrence in the presence of a factor compared to the odds of incurring the disease in its absence. The odds ratios of SNPs in the 8q24 region of the genome are 1.25-1.5, which means than risk of cancer is 25−50% higher in people with these SNPs. Although SNPs associated with cancer risk are present in both 5′ and 3′ region of myc, some SNPs have also been detected in regions distant to the myc gene (~2 Mb). There is evidence that such long-distance regulatory interactions can be mediated by chromatin loops.
References
Further Reading
Last Updated: Feb 26, 2019