Acute infectious gastroenteritis represents a major cause of infant mortality in developing areas of the world and infant morbidity in developed countries. The condition can be caused by a myriad of different microorganisms, but viruses are increasingly recognized as predominant causative factors. Among them, rotavirus is the leading cause of acute gastroenteritis in infants and young children worldwide.
The name rotavirus stems from the characteristic wheel-like appearance of the virus when observed by electron microscopy, which is derived from the Latin word “rota” meaning “wheel”. This pathogen was initially discovered in diarrheic cattle, mice and monkeys, and finally in infants and young children in 1973 by Bishop and Flewett.
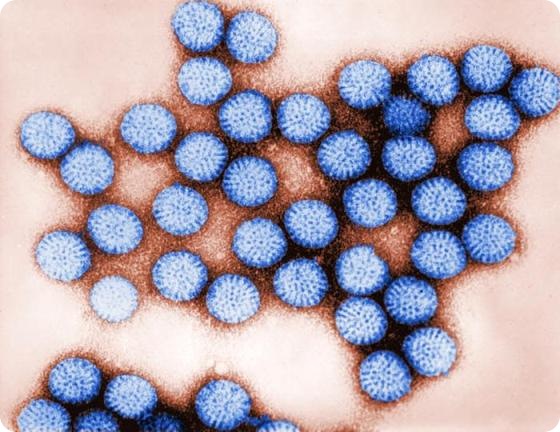
Transmission electron micrograph of intact rotavirus particles, double-shelled. Distinctive rim of radiating capsomeres.
Structure of the virus
The genome of rotavirus contains 11 double-stranded RNA segments with 18,555 base pairs. Each of these segments is a gene, numbered from 1 to 11 by decreasing size. Every genome segment codes for one protein required for the viral lifecycle, except for genome segment 11 that encodes for two proteins.
Such segmented nature of the rotavirus genome is a fertile ground for high frequency of genetic reassortment during mixed infections – a feature that is able to generate new, possibly more dangerous virus strains. Nevertheless, the amount of gene reassortment occurring in nature is not completely known, as only a few rotavirus genomes have been sequenced to date.
The genetic material is found inside a complex 70-nanometre viral nucleocapsid with three concentric shells: an inner core, an internal capsid and an outer capsid. Sixty spikes between 10 and 12 nm in length protrude from the outer capsid.
The viral proteins VP1, VP2, VP3, VP4, VP6 and VP7 are structural proteins that form the virion. The non-structural proteins (produced by the cell infected by rotavirus) are NSP1, NSP2, NSP3, NSP4, NSP5 and NSP6. At least six of the twelve aforementioned proteins bind RNA, with functions important in rotavirus replication.
Classification
Rotaviruses are part of the genus Rotavirus, which is one of the 15 genera of Reoviridae family, subdivided into the sub-families of the Sedoreovirinae (with genera Rotavirus, Orbivirus, Phytoreovirus, Cardoreovirus, Mimoreovirus and Seadornavirus) and the Spinareovirinae (with genera Coltivirus, Orthoreovirus, Aquareovirus, Oryzavirus, Dinovernavirus, Cypovirus, Fijivirus, Mycoreovirus and Idnoreovirus).
According to the International Committee on Taxonomy of Viruses (ICTV), rotavirus can be classified into 7 distinct groups (from A to G), as well as 4 specific subgroups within the group A. Groups A–C can be found in both humans and animals, while rotaviruses of groups D–G are limited exclusively to animals.
As group A is the most important for human infection and disease, it has been classified further using various approaches. Two outer capsid proteins, VP4 (the protease-cleaved protein or P protein) and VP7 (the glycoprotein or G protein) are the determinants of the viral serotype classification, known as P-serotypes and G-serotypes.
Furthermore, this group is also classified according to the migration pattern of the RNA genome segments during polyacrylamide gel electrophoresis, whole genome RNA hybridization patterns (genogroups), as well as nucleotide sequence analyses or genotypes. In 2008, a nucleotide sequence-based, complete genome classification system was developed for strains in this group.
Further Reading
Last Updated: May 13, 2019