MicroRNAs (miRNAs) are small, short non-coding RNAs that bind to specific messenger RNAs (mRNAs) and regulate gene expression either by degradation or by translational inhibition of the mRNA.
miRNAs play an important fundamental role in the development of eukaryotic species as well as in many cellular and environmental processes. Although the variety of miRNAs in an organism is limited as compared to mRNAs, one miRNA can regulate a number of mRNA and affect the gene expression network.
miRNA profiling is a technique used to identify miRNAs that are instrumental in the regulation of a number of processes that are required for the development and maintenance of an organism.
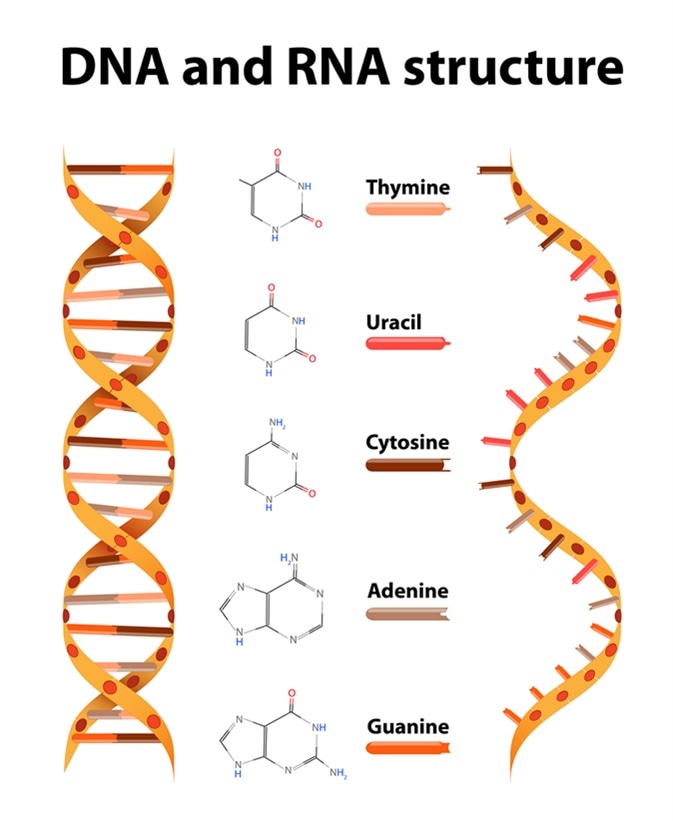
DNA and RNA structure and differences. Image Credit: Designua / Shutterstock
miRNA Biogenesis
The biogenesis of miRNA begins by cleavage of a primary transcript. The length of these primary transcripts (pri-miRNA) may be hundreds or thousands of nucleotides. Cleavage of the primary transcript will yield the precursor miRNA (pre-miRNA) that is approximately 70 to 100 nucleotides (nt) in length. Further cleavage produces the mature miRNAs that are usually 22 nt in length. miRNA have a lower molecular weight than mRNAs.
The mature miRNAs are finally integrated into the miRNA containing RNA-induced silencing complex (RISC) that can be used for various profiling applications.
Introduction to MicroRNA - Anna M. Krichevsky
Sample Selection and Purification for miRNA Profiling
Sample selection and RNA extraction methods are vital steps that affect the results of miRNA profiling. Samples used to extract miRNA may come from a range of sources, such as plasma, serum, culture cells, and fresh or fixed tissue or tumor.
Methods used for purification of miRNA include gel electrophoresis, immunoprecipitation, and laser capture microdissection. The quality of extracted miRNA is tested by spectrophotometry or automated capillary electrophoresis.
Advantages of miRNA over mRNA
miRNA are stable and intact in formalin-fixed paraffin embedded (FFPE) tissues whereas mRNAs are fragmented and not as reliable in FFPE. This advantage of miRNA makes it useful in clinical settings where FFPE tissues are the only samples available.
Microprofiling Methods
Quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) is a specific, inexpensive, and sensitive method that can identify novel miRNAs. However, the limitation of this method is that it is labor intensive if large experiments need to be carried out.
In addition, only one product can be amplified at a time and the short template length makes it difficult to distinguish miRNAs that have very few bases. Efforts have been made to mitigate the specificity issues by using locked nucleic acids as probes but this remains a challenge in larger trials. This method can be used for absolute quantification.
RNA sequencing (next generation sequencing) is also very sensitive and accurate and is used to detect novel RNAs. In this method, a small RNA cDNA library is prepared which is then sequenced in parallel with millions of individual cDNA molecules in one run. Analysis by bioinformatics identifies the new and already known miRNAs.
The advantage of RNA sequencing is that it can distinguish closely related miRNAs (differing even by a single nucleotide and iso-miRNAs) and platforms. However, data analysis requires an enormous amount of computational support and this method cannot quantify miRNA levels.
miRNA microarray has a lower specificity than RNA sequencing but can be used in large studies that cover multiple miRNA targets. The results from the microarray methods are usually validated using a second method such as qRT-PCR or northern blot. This method is the least expensive of all methods. Microarray probing is carried out by labeling the 3´ end of the miRNA with fluorophore-labeled nucleotides with the help of T4 RNA ligase.
These labeled miRNAs are then hybridized on to probes that are discretely arranged and finally arrayed in the form of spots onto a slide or onto beads. The changes in the RNA expression in the samples can be judged by the brightness of individual spots. As in RNA sequencing, this technique can also not be used for absolute quantification of miRNA.
Applications of miRNA Profiling
Apart from novel miRNA detection, miRNA profiling is used to study various gene regulatory network processes, protein interactions, developmental biology, as well as post-transcriptional modifications of miRNA.
For instance, a study using the microfluidics-based multiplexed qRT-PCR assay identified 288 miRNAs in 27 hematopoietic blood cells that are phenotypically distinct, which also included progenitor cells and stem cells.
miRNA profiles are also used as disease biomarkers to understand the differences between normal tissues and affected tissues in a particular disease. Some of the diseases where tissue-based miRNA biomarkers are used include Huntington’s disease, multiple sclerosis, schizophrenia, cancer, rheumatoid arthritis and so on.
Microarray studies have succeeded in identifying miRNA profiles that can classify cancers according to the organ from which it originates. The accuracy of these tests have been predicted to be anywhere between 89% and 100%. Non-invasive biomarkers such as circulatory miRNAs can be used to differentiate between esophageal cancer, hepatocellular carcinoma, and non-small cell lung cancer.
Of late, miRNAs have also been used in forensic settings by characterizing miRNAs from different body fluids.
Although there are hurdles with respect to biological variations between samples, miRNA profiling holds a promising future in molecular diagnostic testing.
Further Reading
Last Updated: Jan 18, 2023