Introduction
The Discovery of Giant Viruses of Amoeba
Diversity and Distribution of Giant Viruses Across the World
Location and Morphology of Giant Viruses
The Life Cycle of Giant Viruses
References
Giant viruses are a part of a group of nucleocytoplasmic large DNA viruses (NCLVD) whose viral particle magnitude, structure, genome length, and complexity are larger than the standard virus family. In the original NCLDV family, the group consisted of the viral families; Poxviridae, Asfarviridae, Iridoviridae, and Phycodnaviridae.
These families are composed of viruses that contain sizeable double-stranded DNA genomes, characterized by replicating entirely inside the host cytoplasm, and all share a hypothetical common origin. They are therefore referred to as a monophyletic group. Although they were known before 2003, increased interest was garnered due to discovering the first giant virus of amoebae named Acanthamoeba polyphaga mimivirus (APMV) in the same year.
AMPV is part of the NCLDV group, occupying its isolated branch in the phylogenetic tree, as this was associated with unique morphological features. Consequently, this led to a new family, mimiviridae, being established. The discovery of mimiviruses was subsequently followed by the isolation of additional giant viruses, which coincided with the development of improved isolation techniques previously constrained by particle-based definitions of viruses.
After that, new and NCLDV members were discovered, which included the following: cedratviruses, faustoviruses, kaumoebavirus, marseilleviruses, mollivirus, orpheovirus, pacmanvirus, pandoraviruses, and pithoviruses. As more giant viruses are being discovered, definitions are being debated.
Despite this, these viruses exhibit distinctive features alongside common characteristics that define them as part of the NCLDV group.
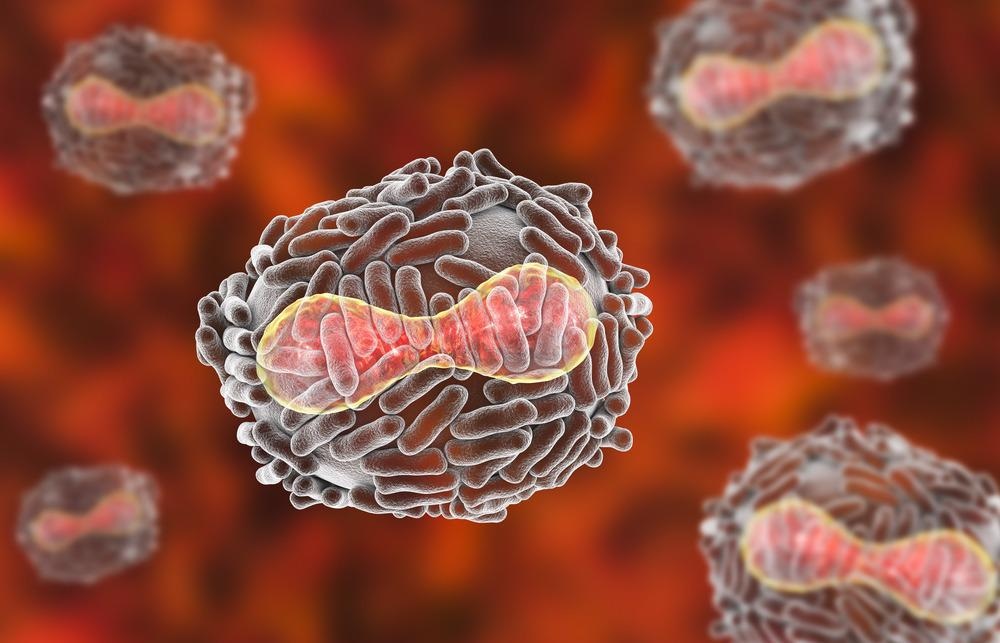
Image Credit: Kateryna Kon/Shutterstock.com
Mimivirus was accidentally discovered due to studying amoeba-associated pathogens from water samples during a pneumonia outbreak in Bradford, England. Initially, the virus was considered a bacterium that infected amoeba, as this stained gram-positively when first encountered in 1992, which it was referred to as the Bradford coccus. When this sample was taken to Aix Marseilles University in France, it was characterized and described in 2003 as the largest known virus at the time.
It subsequently underwent reclassification as a virus due to observations of an eclipse phase during its replication and viewing isometric properties by transmission electron microscopy (TEM); the isometric particle shape is traditional of viruses. Its name is derived from its ability to mimic microbes. They measure approximately 750 nanometers in diameter and are characterized by a protein capsid. This protein capsid contains an internal lipid membrane surrounded by a dense glycoprotein fibril layer that mediates viral attachments to host cells and other microorganisms.
The capsid has a modified vertex instead of the typical icosahedral symmetry. This shape is referred to as stargate, denoting its starfish shape and its characteristic viral genome released into the host cytoplasm. The genome measures 1.2 Mbp in length and encodes >1000 genes. These genes are involved in different functions, such as DNA replication, recombination, and repair, transcription, and translation.
After discovering mini viruses, other giant viruses of amoeba have been described over the past ten years. These have revealed a complexity of viruses with great structural diversity and possible the different sizes and shapes from ~200 nm icosahedral capsids to spherical, ovoid, or tail-containing species that are>1000nm in length.
Marseilleviruses, from the family Marseilleviridae, were the second group of amoeba viruses to be discovered and were associated with Acanthamoeba cells in water samples from Paris, France, and are composed of smaller viruses that present icosahedral capsids of 175–250 nm.
Other isolates described since are composed of five different lineages, with viruses discovered in different regions, including Brazil, Australia, Tunisia, and Japan. These groups are closely related to the African swine fever virus, a mammalian parasite, despite their ability to infect amoeba cells.
In-depth structural analysis of the giant virus, possibles are rare; those that are available have revealed that some viruses display exclusive features. Among them, one type of giant virus, the faustovirus, is formed by double concentric icosahedral capsids that connect by protrusions of the internal caps on their surface, which is not observed for other types of giant viruses.
Since the discovery of the Mimivirus, the number of new isolates has grown every year. These viruses have been isolated using samples on each continent across several conditions, including polluted environments and extreme conditions of temperature and pH. Due to the ever-expanding group of giant viruses, the notion that these viruses are ubiquitous is reinforced. Metagenomic studies have contributed to this reinforcement, indicating that many amoeba-related viruses are yet to be discovered.
An extensive metagenomic analysis of soil from the Harvard Forest in Massachusetts, USA, revealed the complete genome of 16 giant viruses in the family of klosneuvirus and other members of the mimiviridae. This indicates a large diversity of giant soil viruses and suggests several more remain to be characterized.
Image Credit: nobeastsofierce/Shutterstock.com
The replication cycle of giant viruses starts with entry into host cells. Giant viruses enter the host cell by phagocytosis or endocytosis. The genome is released into the host cytoplasm, where a viral factory is established. Here, the genome is replicated, and the morphogenesis of new particles occurs. Nuclear morphological changes occur during the morphogenesis of particular giant viruses (mollivirus and pandoravirus). The cycle of replication ends when a viral progeny is released. This can occur by exocytosis or following lysis of the cell.
Giant viruses and the parasites that carry them are in new fields in virology. Since the discovery of the giant mimivirus, new viral groups have been described, and there is a constant expansion of that complexity and the complexity of the entire virosphere. The advancement of understanding has come from new isolation techniques for viral characterization associated with metagenomic techniques. To fully understand new features, a better understanding of their biology is necessary - this is particularly true due to their large genomic and structural variety.
- Schulz F, Roux S, Paez-Espino D, et al. (2020) Giant virus diversity and host interactions through global metagenomics. Nature. doi:10.1038/s41586-020-1957-x.
- Schulz F, Yutin N, Ivanova NN, et al. (2017) Giant viruses with an expanded complement of translation system components. Science. doi:10.1126/science.aal4657.
- Brandes N, Linial M. (2019) Giant Viruses-Big Surprises. Viruses. doi:10.3390/v11050404.
Further Reading
Last Updated: May 18, 2022