Please can you give an introduction to whole-mount scanning of breast tissue? How does this differ from conventional methods?
We actually normally refer to this as whole-specimen imaging of breast tissue. What we mean is that when tissue is removed from the breast, which could be in the form of a lumpectomy – a breast-conserving surgery – or a mastectomy, the piece of tissue removed is relatively large.
In pathology, that tissue is initially sliced at perhaps 3 to 5 mm intervals. This process is called “bread-loafing”.
In conventional practice, an assistant then identifies smaller areas from the tissue that would represent the areas of interest and from these the tissue sections are produced on conventional small microscope slides of dimensions 25 mm x 75 mm.
For whole-specimen imaging, we actually produce tissue sections that have the same area as the entire bread loaf slice. The slides can be quite large; for a mastectomy they could have dimensions as great as 125 mm x 175 mm. Because of the greater amount of tissue involved we have had to develop special techniques to prepare those sections.
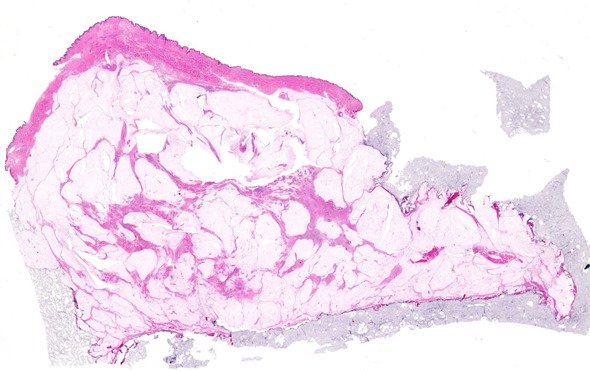
Whole Mount Breast - 5" x 7" at 20X. Image credit: Huron Digital Pathology.
The basic advantage is that by viewing the entire tissue section as one enormous slide, much better sampling of tissue would be accomplished and the pathologist could much more easily appreciate the relationships between tissue types to assess margins, heterogeneity of disease and focality.
For example, when you cut such a large lump of tissue, it tends to spread out or slump and this can create a false measurement of the tumour margins. In my lab we have developed a special gel to encase the tissue so that the lump of tissue is supported during bread-loaf slicing so that it more closely maintains the conformation that it would have had within the breast.
Once these large bread loaf slices are produced, we have to treat the tissue in different ways so that we can do the processing quickly enough. Since we're working with larger amounts of tissue, more time is required for processing chemicals to diffuse into and fix the tissue as well as for performing the other procedures of slide preparation. We use specialized techniques, including microwave processing of the tissue to minimize the time that it takes to produce these large sections.
We also use a special horizontal sledge microtome which allows us to cut serial 4μm sections from these very large bread loaf slices. With a conventional microtome you could really only produce something slightly larger than the conventional microscope slide, but this microtome allows us to accommodate the entire area of the specimen. Finally, we digitize the stained slide using a high resolution electronic scanning system.
What research have you carried out to determine whether whole mount scanning of breast tissue could lead to better clinical outcomes?
We looked at three areas and we've performed research studies using both mastectomies and lumpectomies to assess the difference between the information that we would get using the whole-specimen technique, as opposed to the conventional technique.
The first question was whether there are differences in the assessment of tumor margins using the techniques. We studied over 60 lumpectomies and looked at a comparison of the margins, as assessed by the conventional approach and the whole mount approach.
We found considerable, frequent differences, where samples that had been diagnosed as having acceptable margins with the conventional technique, in fact, turned out to not be acceptable when we looked at the whole-specimen slides.
We used the same tissue for the experiment. We produced the whole-specimen bread loaf slices and the pathology assistant identified from those slices, the tissue that would be sampled using the conventional technique. Then, after digitization, we produced two types of digital slide images – the whole-mount digital image and a set of smaller conventional size images, which would represent what would be done with the conventional technique That way, we could make a fair comparison using the same tissue.
Indeed, we found that there were disagreements between the methods and we believe that the whole-specimen result is more likely to be correct. In some case a margin that was found to be acceptable with the conventional technique was found to be not acceptable with the whole mount technique. However, in some cases, we also found the opposite, where a margin that was considered as not being adequate with the conventional technique was shown to be adequate using the larger slides.
The second question we looked at was regarding the assessment of focality; whether the disease had a single focus or multiple foci. We used the same general approach and again found differences between the two methods. Those differences could go in either direction, so the disease that was deemed to be in a single focus with the conventional approach was often found to be multi-focal with the whole mount and vice versa.
Finally, we've been looking at assessment of extent of disease or tumor burden, comparing the conventional RECIST measurement in one dimension of the size of the cancer, versus a volumetric assessment that we can perform from the whole mount images. We're seeing differences, but this work is still in progress and the results are still being analyzed.
Based on your research, has one technique been shown to be more accurate than the other or is further research needed to confirm?
Our sense is that, because the whole mount provides a more complete picture of what's happening in the tissue and the pathologist doesn't have to piece together bits of information in their mind, it actually provides a more accurate picture of what's going on in the tissue.
Our inclination is to believe that the whole mount is providing better information, but I think the fact that there is a difference is very important.
What are the main challenges when preparing whole-mount serial sections of breast specimens that are diagnostic-quality while preserving 3-D conformation?
The first challenge in the case of breast tissue, is that the tissue can be quite flaccid when the connections from the breast have been removed. Once the various fibers and Cooper's ligaments have been removed, the tissue tends to be flaccid, and, that flopping or slumping of the tissue can distort the relationships between the tumor and the margins. The first challenge really was addressed by using the gel and we found that to be quite successful.
Another challenge is simply the time involved. We have to do this quickly enough for there to be no degradation of the tissue during our processing. As I mentioned, we've been quite successful with that, just by modifying the standard techniques used in pathology to speed things up. In general, the times that we've managed to achieve have been considered acceptable.
The third challenge is the fact that you do need to use a special kind of microtome for cutting the tissue. I think, with practice, the technologists become quite good at this, so it's just a matter of becoming experienced in making these very nice thin 4-micron thick serial sections. A good technologist can just do this all day and get very good results.
The final challenge is the cost associated with having the larger slides and more glass and specialized equipment. We feel that if this technique becomes more widely used, then there will be some advantages associated with scale and the higher cost will no longer be a factor.
Why do you use the TissueScope from Huron Digital Pathology in your research?
We're interested in looking at whole mounts from the breast and, in the case of mastectomies or even larger lumpectomies, this is the only system we're aware of that can digitize such large slides. It's really about the capacity for covering this large area and doing quite a good job with these large slides.

Image credit: Huron Digital Pathology.
How flexible is the TissueScope in terms of the size of the slide?
The system has carriers for a wide range of different size slides. While we really take advantage of the system because of its unique ability to do these larger, 5 x 7 inch slides, there's certainly no problem with doing smaller slides. On the slide carriers, one can actually put multiple slides in and scan them in the same run, so the flexibility is very good.
What impact do you think whole-mount scanning of breast tissue will have on clinical outcomes moving forwards?
Since we really believe that we're seeing a more accurate picture with respect to margins and focality, which are prognostic factors, we think this is the right way to be assessing the results of breast surgery and using this information to guide the next phase of therapy.
It's hard to know the impact, which will depend on whether the technique is adopted by multiple centers, but certainly our pathologists are very enthusiastic about this because they feel that they can get a better assessment of the samples. They can also do it more quickly and I think they walk away from the exam feeling more confident that they've appreciated what's going on in a particular case.
Adoption is really key and that will probably depend to some extent on more people being aware of the technique, the results of some of our publications and other people trying to replicate them. I think it's very exciting.
There are only a few places in the world where whole mount is being done right now, although the people who are doing it are very enthusiastic and I think we have a challenge in terms of making our results more widely known. Hopefully this will lead towards a greater adoption of the approach.
What are your future research plans with regards to whole-mount scanning?
I’m glad that you asked because I'm a researcher and not a clinical pathologist. We felt that the whole mount technique would provide some benefits to the pathologist in terms of day-to-day clinical work, but we're very interested in cancer research and understanding the implications of the heterogeneity of various biomarkers in the breast and the implications with respect to therapy.
Currently, we're looking both at the histological and immunohistochemical biomarkers in different areas of a lesion; examining the heterogeneity of those biomarker signals and, in some cases, trying to correlate those with genomic information and perhaps even to use both the genomic and histopathologic information in combination. That's the direction of our current work.
I think the challenge in breast cancer and prostate cancer is overtreatment or undertreatment of the disease. If we can get a better assessment of how aggressive treatment needs to be, I think that would help a lot in terms of providing more appropriate precision therapy.
Where can readers find more information?
- Clarke GM, Eidt S, Sun L, Mawdsley GE, Zubovits JT, Yaffe MJ. Whole-specimen histopathology: A method to produce whole-mount breast serial sections for 3-D digital histopathology imaging. Histopathology 2007 Jan; 50(2):232-42.
-
Clarke GM, Holloway CMB, Zubovits JT, Nofech-Mozes S, Liu K, Murray M, Wang D, Yaffe MJ. Whole-mount pathology of breast lumpectomy specimens improves detection of tumor margins and focality. Histopathology 2016. Article first published online: 19 JAN 2016
About Dr Martin Yaffe
Dr. Martin Yaffe received training in Physics and Medical Physics at The University of Manitoba and a PhD in Medical Biophysics from The University of Toronto where he is now Professor of Medical Biophysics and a Senior Scientist at Sunnybrook Research Institute. He is also Director of the Smarter Imaging Research Program at The Ontario Institute for Cancer Research. He holds the Tory Family Chair in Cancer Research.
Dr. Yaffe has been actively involved in research on breast cancer detection and breast cancer imaging for over 35 years and his group pioneered the development and clinical evaluation of digital mammography and quantitative measurement of breast density, an important risk factor for breast cancer. Over the past decade he has turned much of his attention to the application of imaging methods in pathology and has developed methods for whole-mount digital histopathology and multiplex imaging of cancer biomarkers. He created the state-of-the art Biomarker Imaging Research Laboratory (BIRL) to further this work.
Dr. Yaffe is a Fellow of The American Association of Physicists in Medicine and an Honorary Fellow of The Society of Breast Imaging. In February, 2016 Dr. Yaffe was inducted as a Member of The Order of Canada.