As individuals and as populations, our risks of getting diseases are determined partly genetically and partly by our environment. The microbiome is an important part of that environment that mediates between the outside world and the inside world of our bodies.
Our guts are filled with bugs. There's up to a kilogram of bugs inside you, with a collection of about 10 million genes. The human body has about 10 trillion cells, and the microbiome is about 100 trillion organisms living inside you at any one time.
The “exposome” is a concept that involves thinking about all the things from the environment that get into your body. It relates to your diet and to the microbes in your gut, which process the diet and make molecules of their own - microbial metabolites. Thus the “exposome” is the measured sum of the environmental exposures.
We also need to take into account the drugs we take and the chemicals we're exposed to in the environment. Together, these exposures, plus your genes, determine your disease risk as an individual.
The exposome is all the different exposures that impact your body's function. Your genome, the genes that you have, are really just instructions.
Genes are just a set of instructions for building an organism in a certain way, and how that organism is built is determined by not only the program, the genes, and the genome that runs it but also the substrates available to it through the diet and microbial signaling from the gut.
Ultimately, our risk of getting a disease is highly dependent on all of those interactions. The other thing that's complicated about this is that the interactions vary throughout life, so exposures to diet variation, drugs, etc., that occur in babies have very different effects to the same source of exposures that occur in adult life, so there's a sort of conditional interaction between your genes and your environment that progresses throughout life, changing the risks for disease at any stage of life and the risks for getting diseases in the future.
In particular, probably the most important exposures, the most important interactions, are those in the first few years of life, where your body is programmed for a lot of the later biology that it's going to experience.
It's interesting to note that the microbiome takes about 3 years to develop because babies are born without microbes, and the microbes develop very, very quickly. In fact, there are trillions of bacteria within a few days of the baby being born, and it takes about 3 years before you get an ecology in the child that is similar to that of an adult, so anything that upsets that development of the ecology in early life can be quite influential for risk factors later on.

How important is the microbiome in human health, and why are disorders of the gut microbiome associated with multiple non-infectious diseases?
It's only recently we've started to understand how complex the microbiome is and how important it is to us. A lot of the bacteria that live inside us are not readily culturable, so microbiologists have only known a little bit about the biology of the microbes historically because you can't dig many of the bugs out and actually make them grow. The reason for that is that these microbes live in an environment that has other microbes of different species, and they all work together and need each other.
Modern genomics and the technologies originally developed for the Human Genome Project can also be used for microbiome sequencing. About 10 or maybe 15 years ago, people started using modern gene-sequencing technology, so we can go into detail about what is in this microbial community to find out who is there.
Now we know that there are probably 2 to 3,000 species in the human gut. The number will vary between people, but those 2 to 3,000 aren't the same species in different human individuals, so there might be bugs in you that are not in me and vice versa.
If you think about that in terms of genetic diversity, humans, you and I, say, are 99.99% similar genetically. Our microbiomes are probably less than 5% similar genetically. We have amazing microbiological genetic diversity within us, and the microbes perform many biological functions for humans, which means that functional variability is enormous between individuals as well.
To give you an idea of what I mean by function, there are biological pathways that control our body, and many of them are under the host's genetic control. For example, the enzymatic sequence for breaking down sugar and glucose into smaller bits that can be used for energy metabolism is under human genetic control.
There are lots of other pieces of biochemical machinery in the body that are not actually under human genetic control. In fact, quite a lot of cholesterol metabolism, for instance. People talk about cholesterol a lot, the good cholesterol and bad cholesterol, but in fact, if you look at the cholesterol pathway, the conversions from all the different metabolites, the different sequential chemical conversions, and cholesterol metabolism lead to steroid metabolism, etc., these are highly related.
Microbe-signaling molecules partly determine something like 90% of all the conversions in those pathways. If you like, some fundamental parts of our machinery are controlled by microbial activity. Microbes make drug-like compounds; they switch on and off our pathways, and it's only been recently understood just how deep that connectivity is. This is the basis of what biologists call symbiosis.
Symbiosis involves two or more organisms that are physiologically connected and perform functions for each other, ultimately resulting in a situation where one cannot do without the other; extreme examples of symbiosis are organisms that cannot survive without each other.
Humans can survive, just about, without their microbes, but they don't do very well at all. In fact, all animals with a gut rely on their gut microbes performing many biochemical functions, as a result of which they exhibit "genome reduction."
Genome reduction is actually a reduction in the number of genes because you've got someone else to do the job for you. This is quite an interesting idea because it means that humans are actually incomplete as an organism. They need microbes to complete them and to control their pathways properly.
To give you an example, if you'd make gnotobiotic (germ-free) animals, e.g., by C-sectioning rats and keeping them in a germ-free environment so they never get exposed to microbes, the normal surface of the gut, the so-called villus surface of the gut, in those rats never, ever develops.
The gut has got lots and lots of very complex surface area, increasing invaginations and villi, little projections. The development of all of that anatomy is dependent on microbial signaling, so it's an extraordinary thing to say that your genome does not even encode your gut structure.
It actually requires microbial genomes to work. That's just one rather splendid example of genome reduction, but the important thing is the diversity of the microbes.
If you can imagine that microbes are partly controlling your biology, your microbes are different from mine. Our biology is different because our microbes are different. Therefore, the way that we interact with the exposome, the way that we interact with diseases, and the way that we respond to drugs are dependent on variations that ultimately advance variations in our microbes.
We've only really started to understand just how deep that connection is in the last 5 to 10 years. If you can bring it back to humans and things like understanding how drugs work in the body, microbes can detoxify drugs to make them less toxic or more toxic, so there are some splendid examples.
Gut microbes metabolize amphetamines. Digoxin, the heart drug, is metabolized by microbes, so your different microbial composition changes how drugs work in you as well as the likelihood of diseases occurring in you.
When you consider this from the population level, the population's disease risks change partly due to people's behavior, what they eat, and how they exercise, but also because of their microbes. At the individual level, microbes also change drug therapy.
When it comes to personalized healthcare, just having the human genomic part of it is not good enough because it doesn't explain much of the physiological variation between individuals, which we need to understand and optimize therapies for individuals.
The microbiome is going to be important in the future because it will help us understand disease risk and the way that the environment and the human body interact through the microbiome. Still, in personalized healthcare, we're going to have to understand the microbiome to get that fine-tuning of therapeutic management that we're going to need in the future.

How does this complexity impact pharmacogenomics?
Pharmacogenomics has had few successes, given the effort and money involved. You can certainly stratify people into different genetic classes of disease. Breast cancer is a good example, where there are about 10 different genetic sub-varieties that you can detect, and some are more or less responsive to certain drugs.
If you know about somebody's genetic background, you can start predicting whether a particular drug is likely to be good for that person. There have been some successes in this area.
The point is that even within a specific genetic subclassification, there's still quite a variety of patient variations and responses to therapy. Some people respond better than others, and that extra variation has to do with things like physiological variation, microbial variation, and lifestyle variation.
If you put the genetic information together with the microbial and sort of the metabolic phenotypes you can measure, you start to cover all of the bases. You end up having a much deeper understanding of human biology that you need for personalized healthcare.
What new technologies and systems have been developed to help us understand the complexity of human diseases?
Genomics has been tremendously important in helping us understand the basis of disease, but the fact of the matter is that most people in the world die of non-genetic causes; they die of environmental causes, starvation, and they die of infectious diseases.
We need technologies that go beyond genes, and so people have developed technologies looking at proteins, which are the machines made on genetic instructions to run the body. There are technologies for looking at metabolism, which is the small-molecule currency of the body, the energy balance, the breakdown of food, the excretion, and the building and excretion of metabolic products that we don't need.
Each of those different areas requires different technologies, so each level of what we call "biomolecular organization," the genetic, the protein, and the metabolic, requires a different type of technology to explore the variations in human biology.
Genes give you the potential of what might happen in relation to an environmental stimulus or stressor. The proteins are the machines that execute the variation in the genes, and the metabolites are the products that tell you what's actually happening in the body. Because the patterns reflect fundamental activities of the chemistry within the body, it often tells you why there's a problem as well.
Some of the most important technologies developed recently involve metabolic analyses, which allow you to analyze thousands of metabolites quickly and precisely, giving you deep insight into human biology.

In what ways is our understanding still limited?
What we are challenged by is the number of levels of biological interaction and understanding what it means for the human. We think of this like the microbes playing a tune, a chemical tune. Microbes make molecules that switch on and off receptors in our bodies and control pathways.
Another challenge is understanding all of these different microbes, all talking to each other and talking to us, and which ones are controlling the overall signaling process. We simply don't understand that yet. When we do understand that, we will eventually be able to start making interventions for it.
Imagine that a particular metabolic pathway in the human body is controlled by a molecule made by a microbe that can switch on and off, maybe a pathway in the body. If you switch it off, you may change a disease-risk factor, a cancer, or something like that. We think such things exist.
If you could understand which microbe was making which molecule, you could work out a way of switching off that microbial metabolite production, thus changing the activities in the human body and potentially changing the disease risk.
That's the big prize in the future. It's a whole new way of thinking about therapy for the human body, which involves controlling microbial activity that exists within us as a way of controlling our bodies and maybe even intervening in disease processes or preventing disease processes from occurring at all.
Although our understanding is still highly limited because of the complexity, the long-term value of understanding is going to be critical because it will change the way that we look at medicine and the way that we develop therapies in the future.
What are the main challenges in personalized medicine and public health to date?
Understanding real human biological complexity in a way that you can do something about it. We talk a lot about “clinical actionability,” which is the ability for you to take certain pieces of scientific knowledge about a system or a person and to be able to act on it so that the doctor who looks at this piece of information, says, "I know what to do next."
At the moment, we have many technologies that describe human biology in ever more detail, but most of that detail is not very helpful to the doctor. He doesn't know how to use it to make a decision. The biggest challenge to personalized healthcare is translating the knowledge of biological complexity into an actionable pathway that a doctor can use.
The same thing applies to public health, where we are interested in understanding the basic causes of disease, whether they're genetic or environmental, and changes in disease patterns, and again it's about describing human biology in great detail, but at the population level rather than at the individual level.
We could also say that public health research exists to inform future healthcare policy. The challenge here is getting the biological knowledge and then expressing it in a way that it can be useful toward healthcare policy and also practically for advising people as to how to run their lives.
How can we overcome challenges such as antibiotic resistance, obesity, and physical inactivity?
Education is one part of it. The reason we've got antibiotic resistance is because we've misused antibiotics. The massive overuse of antibiotics in agriculture for increasing growth in cattle, etc., has led to an environmental pool of antimicrobial resistance.
People didn't understand the long-term consequences of antimicrobial use in agriculture, which has resulted in this environmental problem of microbial resistance.
Doctors overprescribe antibiotics, and people often don't take a full course because they want to save some for later in case they get sick again. Still, of course, the prescription is to destroy all of the bugs, the hostile bugs, and any that are left will remain because they're resistant to that drug.
We've ended up in a really, very serious situation with antimicrobials, and within the next decade, there probably will be microbes that emerge that are resistant to all known antibiotics. These could cause huge problems for the healthcare system.
We're already beginning to see it: 1 in 7 of hospital-acquired infections in the UK are non-treatable, and every person who gets an infectious complication in a hospital tends to increase their stay in hospital by 30%.
If you work that through, that's financially enormous. What we've got happening now is increased numbers of resistant bugs out there with new classes of drug resistance emerging. So this problem is going to get worse and worse, such that unless something is done about it in the next 20 or 30 years, we will go back to 19th-century healthcare, where the majority of the population died of infectious diseases and not old age.
This is really quite an existential issue for modern society. In fact, there are some interesting angles one could take on it, including the microbiome, which I'm going to come back to, but the same sort of thing applies to obesity.
Why are people obese? Well, they eat too much, and they don't exercise enough. It's really standard, plain pharmacodynamics, as a matter of fact. Why do they eat too much and not exercise enough? Because they're not properly educated to know that this will have huge impacts on their health in the long term. There tend to be associations between low socioeconomic status and obesity, and indeed, low educational level and obesity.
It's not to say that poor people are stupid, not at all. It's because what tends to happen is that people are informed about healthcare according to their socioeconomic level. Also, their ability to change their diet is determined by their socioeconomic level as well.
Any study on obesity will show that socioeconomic status, even salary, is correlated with obesity and other related issues. Again, we've got to think about solutions that work not just for rich people but also for poorer people.
Interestingly, of course, the obesity epidemic that we're seeing now is partly related to changes in the microbiome, which brings us back to antibiotic use again. Microbes control a significant amount of calorific transfer from the diet to the body.
Quite a lot of calories go into rebuilding the microbiome. There are two parts to it. Maybe 10% of your protein intake is actually related to building the microbes that live inside you because they turn over, and you have to replenish them.
Another part of the protein in your diet goes to rebuilding the gut wall because your gut wall turns over every few days as well, so that's sort of on the negative side, if you like, of the calorific balance that microbes take up calories because they need to build their cells.
On the other hand, they also perform digestive and fermentation functions in the body and in the gut, which makes more calories available. So, by changing which microbes are there, you change the balance of power between negative and positive uses of calories, potentially changing an individual's weight gain on a given diet. Understanding the microbiome is also important to understanding obesity in really quite complex ways.
Physical inactivity is a product of our modern lifestyle: We've got cars, we have fancy TVs, and we like sitting around in front of them. Many people have sedentary jobs, where they sit in front of computers all day. That is a consequence of modern life, and people need to be educated as much as possible about healthy lifestyles and dietary choices.
Can you please give an overview of the MRC-NIHR National Phenome Centre? What are the center's main aims regarding metabolic phenotyping?
It came from the Olympic drug-testing center that was set up in 2012 to look at substances of abuse in athletes. It was a big analytical facility that looked at several hundred different substances that athletes might take to performance-enhance themselves.
In 2011 and early 2012, the Prime Minister talked a lot about the Olympic legacy and justified why we spent billions of pounds on the Olympics and what we would get out of it. People normally talk about swimming pools, sports stadiums, and travel infrastructure.
At that time, Professor Elliott, Head of Epidemiology at Imperial, and I wrote to Dame Sally Davies, Chief Medical Officer for England, and said, "Well, we've got a great idea for Olympic legacy.
Supposing at the end of the Olympic Games, we take the Olympic drug-testing facility and repurpose it for a national facility for metabolic analysis for populations." We were in a very fortunate, very timely situation in making a suggestion that suited the politics of the MRC, the NIHR, and the government.
Then we were asked to write the grant proposal, which we did, 75 pages, some of it for justification by the peer review. The MRC and NIHR awarded us 10 million pounds, and we also raised about 11 million pounds from industry to move the Olympic drug-testing facility into Imperial and create a national facility for large-scale phenotyping, which is what the National Phenome Centre.
It's the first of its kind in the world, and it's the only laboratory in the world that can do phenotyping at that scale. There are a lot of metabolic laboratories around the world, but there are none of them that can do hundreds of thousands of samples a year because the Olympic drug-testing facility was built for forensic-level analysis because you're basically doing drug testing on people. If you get that wrong, you're all over the newspapers!
We've built a very high-quality control laboratory and a high-throughput laboratory that allows us to screen large epidemiological cohorts and populations for metabolic features associated with disease risks, as I mentioned earlier about obesity.
The questions are: What are the metabolic features of obesity that tell you about the gene-environment interactions that precipitated the obesity? How does diet impact your metabolism at the population level? Are there markers you can detect that predict cancer in individuals based on metabolic profiles? Are there markers you can get that predict stroke or cardiovascular disease?
The National Phenome Centre is a fantastic laboratory, and it's built on an industrial scale. We're very lucky to have it here at Imperial. I'm currently involved in creating an international network of phenome centers because now that we've built ours, lots of other countries want one as well. There are going to be ones in America, Japan, and China. There's already one in Singapore linked to Imperial, and one in Birmingham has just opened too.
In the next few years, there will be national phenome centers that all use the same technology that we do and harmonize our data so that we can all share and exchange information about our population's disease patterns. By sharing our technology and data, we'll be able to make the first global atlas of metabolic disease around the world, which is only made possible by doing large-scale phenotyping studies.
The other part of that is the same technology can be used to do patient profiling as well, so we can use it for personalized healthcare. We have a Clinical Phenome Centre in Imperial as well, which is funded by the National Institute of Health Research, NIHR, and here we're not just looking at populations.
It will be individual responses to drug therapy as well, through the effects on metabolites, so that we can record so that we know whether somebody is getting metabolically better or worse when they're under drug therapy.
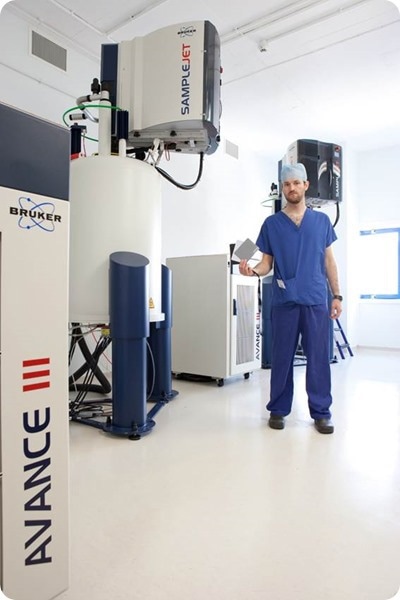
How important are technologies such as NMR in research at the MRC-NIHR National Phenome Centre?
For metabolic phenotyping, there are only 2 important analytical technologies: nuclear magnetic resonance spectroscopy and mass spectrometry. Both of these tools, these technologies can measure hundreds or thousands of metabolites in any sample in the same analytical run.
They work in quite different ways. Mass spectrometry works on the molecular weight of molecules, their masses, how big they are, how they break up, and how they fragment.
NMR looks at fundamental magnetic connectivity properties between atomic nuclei, which are characteristic of individual molecules. They're complementary to each other. We screen every sample they get in the National Phenome Centre and our Clinical Phenome Centre by NMR spectroscopy in the first instance.
NMR gives us a broad profile of maybe 1,000 different metabolites present in the samples. We get data that’s actually like a fingerprint, which is very useful. In things like blood plasma, we can measure all of the plasma lipoproteins quantitatively from NMR spectroscopy.
NMR is the definitive technology for measuring lipoproteins, and the HDL versus LDL balance tells you a lot about your heart disease risks, coronary artery disease, etc. Intrinsically, NMR carries useful biological data.
There are lots of different sub-types of mass spectrometry. We use several different sub-types in the mass spectrometer according to the types of molecules we're interested in. Now, with mass spectrometry, you have to do a bit more work.
You have to not only set the sample right using some chromatographic process, but the mass spectrometer measures molecules individually and says what they are. You can also measure them more sensitively with mass spec. You can measure more of them by mass spec than you can with NMR – maybe up to 25,000 metabolites.
What do you think the future holds for personalized medicine and surgery around the world?
Surgery is about the most personalized healthcare you can get because it's one surgeon cutting you up, normally, so it's a one-on-one, literally physical interaction. Of course, we're trying to revolutionize surgery by use of technology, like the iKnife, which measures the chemistry of the smoke that's created when a surgeon cuts and that can be analyzed using a mass spectrometer so that almost instantaneously, the surgeon knows what he's cutting through based on the chemical composition of the smoke.
There is a readout that basically does a mathematical analysis of the smoke chemistry in relation to a database where we have already annotated, if you like, the smoke chemistry. That's fantastic for the personalization of surgery because it gives the surgeon more knowledge about the local biology of that patient to help in real time with the surgical procedure.
Of course, with personalized medicine more generally, when you look at maybe kinds of chemotherapy or other medical interventions rather than surgical ones, then metabolic phenotyping technologies give you much more detail about the individual variation in the biology of those patients.
We created a concept a few years ago called "Pharmacometabonomics," which is the metabolic equivalent of pharmacogenomics, where we use a pre-interventional metabolic signature and a mathematical model of it to say, "Well, you'll be this sort of person metabolically. Therefore, we know from experience that you're in this particular category, and then we can potentially find which drugs will be good for you or bad for you.” We've already demonstrated that with anticancer drugs, you can predict the efficacy and toxicity of the drugs in humans based on plasma metabolic profiles.
At the moment, we're in this phase now trying to get into clinical trials for all these things, where we've got all these scientific observations, and we're now trying to create patient-journey clinical trials, so we really evaluate quantitatively how much improvement in healthcare we get by the deployment of these technologies.
Of course, we have to cost them in relation to care pathways as well, so in order to get improved personalized healthcare and implement it properly in the healthcare systems, we actually have to improve people's health and shorten their journeys.
We have to save money in a healthcare system that's very strapped for cash, so there are 2 ways to save money in the healthcare system, really. Very basically, it's 1, spend less time in the hospital when you're there, and 2, go there less often.
At the moment, it takes 3, or 4 or even 5 visits to a hospital for problems we've sorted out. What we'd like it to be is 1 or 2. 1 or 2 visits to the hospital, and then you're saving an enormous amount of time in the administration, etc.
Then the other thing is, if your hospital journey takes 5 days, can we improve treatment? Hence, it only takes 3 days, and it does not only improve it by shortening the journey point but actually has you better in a shorter period, better for the patient, and you get that by doing personalized healthcare.
The more you understand the patient's problems, the more clinically actionable they are. You can intervene more quickly and effectively, which will help the patient recover quicker, hopefully, improve the chances of their survival if it's a dangerous disease, or cancer, or something like that, and at the same time save money because it's shortening the journey.
If you can get all the components aligned, then the feature of personalized healthcare will be fantastic, but it's only going to work for certain disease areas. It won't work for everything, and one of the things that we're trying to do at the moment is to understand what is going to work for and what it's not going to work for.
Certainly, cancer is an obvious area because cancer is biologically, genetically, and physiologically quite diverse, so the technologies that we've been developing, which are about measuring diversity and variation in patients, are going to be very useful; for instance, we think in personalized cancer therapies in the future.
We've got a lot of work still to do. We see light at the end of the tunnel, and what we've got to do, hopefully, is emerge from the tunnel before global warming or multiple antibiotic resistance gets us because those are looming up.
What do you think the future holds for antibiotic resistance?
Weaponizing the microbiome may be the way around antimicrobial resistance. If we really understood our microbes well enough, we could program and engineer them to destroy pathogens. That would be a much more effective solution than trying to invent new antibiotics every 15 years and trying to do that forever, but we have a long, long way to go!
Where can readers find more information?
http://www.imperial.ac.uk/phenome-centre
- Holmes, E., Kinross, J., Gibson, G. R., Burcelin, R., Jia, W., Pettersson, S., and Nicholson, J. K. (2012) Therapeutic modulation of microbiota-host metabolic interactions, Sci Transl Med 4, 137rv136.
- Nicholson, J. K., Holmes, E., Kinross, J., Burcelin, R., Gibson, G., Jia, W., and Pettersson, S. (2012) Host-gut microbiota metabolic interactions, Science 336, 1262-1267.
- Nicholson, J. K., Holmes, E., Kinross, J. M., Darzi, A. W., Takats, Z., and Lindon, J. C. (2012) Metabolic phenotyping in clinical and surgical environments, Nature 491, 384-392.
- Want, E. J., Masson, P., Michopoulos, F., Wilson, I. D., Theodoridis, G., Plumb, R. S., Shockcor, J., Loftus, N., Holmes, E., and Nicholson, J. K. (2013) Global metabolic profiling of animal and human tissues via UPLC-MS, Nat Protoc 8, 17-32.
- Balog, J., Sasi-Szabo, L., Kinross, J., Lewis, M. R., Muirhead, L. J., Veselkov, K., Mirnezami, R., Dezso, B., Damjanovich, L., Darzi, A., Nicholson, J. K., and Takats, Z. (2013) Intraoperative tissue identification using rapid evaporative ionization mass spectrometry, Sci Transl Med 5, 194ra193.
- Zheng, X., Zhao, A., Xie, G., Chi, Y., Zhao, L., Li, H., Wang, C., Bao, Y., Jia, W., Luther, M., Su, M., Nicholson, J. K., and Jia, W. (2013) Melamine-induced renal toxicity is mediated by the gut microbiota, Sci Transl Med 5, 172ra122.
- Merrifield, C. A., Lewis, M. C., Claus, S. P., Pearce, J. T., Cloarec, O., Duncker, S., Heinzmann, S. S., Dumas, M. E., Kochhar, S., Rezzi, S., Mercenier, A., Nicholson, J. K., Bailey, M., and Holmes, E. (2013) Weaning diet induces sustained metabolic phenotype shift in the pig and influences host response to Bifidobacterium lactis NCC2818, Gut 62, 842-851.
- Elliott, P., Posma, J. M., Chan, Q., Garcia-Perez, I., Wijeyesekera, A., Bictash, M., Ebbels, T. M., Ueshima, H., Zhao, L., van Horn, L., Daviglus, M., Stamler, J., Holmes, E., and Nicholson, J. K. (2015) Urinary metabolic signatures of human adiposity, Sci Transl Med 7, 285ra262.
- O'Keefe, S. J., Li, J. V., Lahti, L., Ou, J., Carbonero, F., Mohammed, K., Posma, J. M., Kinross, J., Wahl, E., Ruder, E., Vipperla, K., Naidoo, V., Mtshali, L., Tims, S., Puylaert, P. G., DeLany, J., Krasinskas, A., Benefiel, A. C., Kaseb, H. O., Newton, K., Nicholson, J. K., de Vos, W. M., Gaskins, H. R., and Zoetendal, E. G. (2015) Fat, fibre and cancer risk in African Americans and rural Africans, Nat Commun 6, 6342.
- Phetcharaburanin, J., Lees, H., Marchesi, J. R., Nicholson, J. K., Holmes, E., Seyfried, F., and Li, J. V. (2016) Systemic Characterization of an Obese Phenotype in the Zucker Rat Model Defining Metabolic Axes of Energy Metabolism and Host-Microbial Interactions, J Proteome Res.
About Professor Jeremy K. Nicholson
- Professor of Biological Chemistry
- Head of the Department of Surgery, Cancer and Interventional Medicine
- Director of the MRC-NIHR National Phenome Centre
- Director of the Centre for Gut and Digestive Health (Institute of Global Health Innovation)
- Faculty of Medicine, Imperial College London
Professor Nicholson obtained his BSc from Liverpool University (1977) and his PhD from London University (1980) in Biochemistry, working on the application of analytical electron microscopy and the applications of energy dispersive X-ray microanalysis in molecular toxicology and inorganic biochemistry. After several academic appointments at London University (School of Pharmacy and Birkbeck College, London, 1981-1991), he was appointed Professor of Biological Chemistry (1992).
In 1998, he moved to Imperial College London as Professor and Head of Biological Chemistry and subsequently Head of the Department of Biomolecular Medicine (2006) and Head of the Department of Surgery, Cancer and Interventional Medicine in 2009, where he ran a series of research programs in stratified medicine, molecular phenotyping and molecular systems biology.
In 2012, Nicholson became the Director of the world’s first National Phenome Centre specializing in large-scale molecular phenotyping, and he also directs the Imperial Biomedical Research Centre Stratified Medicine program and Clinical Phenome Centre. Nicholson is the author of over 700 peer-reviewed scientific papers and many other articles/patents on the development and application of novel spectroscopic and chemometric approaches to the investigation of metabolic systems failure, metabolome-wide association studies, and pharmaocometabonomics. Nicholson is a Fellow of the Royal Society of Chemistry, The Royal College of Pathologists, The British Toxicological Society, The Royal Society of Biology and is a consultant to several pharmaceutical/healthcare companies.
He is a founder and director of Metabometrix (incorporated 2001), an Imperial College spin-off company specializing in molecular phenotyping, clinical diagnostics, and toxicological screening. Nicholson’s research has been recognized by several awards including The Royal Society of Chemistry (RSC) Silver (1992) and Gold (1997) Medals for Analytical Chemistry; the Chromatographic Society Jubilee Silver Medal (1994); the Pfizer Prize for Chemical and Medicinal Technology (2002); the RSC medal for Chemical Biology (2003); the RSC Interdisciplinary Prize (2008) the RSC Theophilus Redwood Lectureship (2008); the Pfizer Global Research Prize for Chemistry (2006); the NIH Stars in Cancer and Nutrition Distinguished Lecturer (2010), the Semelweiss-Budapest Prize for Biomedicine (2010), The Warren Lecturer, Vanderbilt University (2015).
He is a Thomson-Reuters ISI Highly cited researcher (2014 and 2015, Pharmacology and Toxicology, WoS H index = 108). Professor Nicholson was elected as a Fellow of the UK Academy of Medical Sciences in 2010, elected Lifetime Honorary Member of the US Society of Toxicology in 2013, and Honorary Lifetime Member of the International Metabolomics society in 2013.
He holds honorary professorships at 12 Universities (including The Mayo Clinic, USA, University of New South Wales, Chinese Academy of Sciences, Wuhan and Dalian, Tsinghua University, Beijing and Shanghai Jiao Tong University, Nanyang Technological University Singapore. In 2014, Nicholson was Elected as an Albert Einstein Professor of the Chinese Academy of Sciences.