Please give an overview of the past research into machine learning and artificial intelligence in medical imaging. What are we currently able to do with this research?
The two major tasks in medical imaging that appear to be naturally predestined to be solved with AI algorithms are segmentation and classification. Most of techniques used in medical imaging were conventional image processing, or more widely formulated computer vision algorithms.
Image credit: Huron Digital Pathology.
One can find many works with artificial neural networks, the backbone of deep learning. However, most works were focused on conventional computer vision which focused, and still does, on “handcrafted” features, techniques that were the results of manual design to extract useful and differentiating information from medical images.
Some progress was visible in the late 90s and early 2000s (for instance, the SIFT method in 1999, or visual dictionaries in early 2000s) but there were no breakthroughs. However, techniques like clustering and classification were in use with moderate success.
K-means (an old clustering method), support vector machines (SVM), probabilistic schemes, and decisions trees and their extended version ‘random forests’ were among successful approaches. But artificial neural networks continued to fall short of expectations not just in medical imaging, but in computer vision in general.
Shallow networks (consisting of a few layers of artificial neurons) could not solve difficult problems and deep networks (consisting of many layers of artificial neurons) could not be trained because they were too big. By the mid 2000s there was theoretical progress in this field with the first major success stories in early 2010s on large datasets like ImageNet.
Now suddenly, it was possible to recognise cats and cars in an image, perform facial recognition and automatically label images with a caption describing its content. The investigations of applications of these powerful AI methods in medical imaging has started in the past 3-4 years and is in its infancy but promising results have been reported here and there.
What applications are there for machine learning and artificial intelligence in medical imaging?
Based on recent publications, it seems that the focus of many researchers is on diagnosis, mainly cancer diagnosis, where the output of the AI software is often a “yes/no” decision for malignant/benign, respectively.
The other stream is working on segmenting (marking) specific parts of the images, again with the main attention of many works being on cancer diagnosis and analysis, but also for treatment planning and monitoring.
However, there is much more that AI can offer to medical imaging. Looking at its potentials for radiogenomics, auto-captioning of medical images, recognition of highly non-linear patterns in large datasets, and quantification and visualization of extremely complex image content, are just some examples. We are at the very beginning of an exciting path with many bifurcations.
What are the current limitations in the characterization of tissues and their attributes with artificial intelligence? What needs to be done to overcome this?
AI is a large field with a multitude of techniques based of different ideas. Deep learning is just one of them, but it is the one with the most success in recognizing image content in recent years. However, deep learning faces multiple challenges in digital pathology.
First and foremost, it requires a large number of marked (labelled) images (images in which the region of interest has been manually delineated by a pathologist) but the general workflow of digital pathology does not provide labelled images. This has led the research to work on specific cases, e.g., breast cancer, for which a small number of labelled images can be provided to demonstrate the feasibility of deep learning.
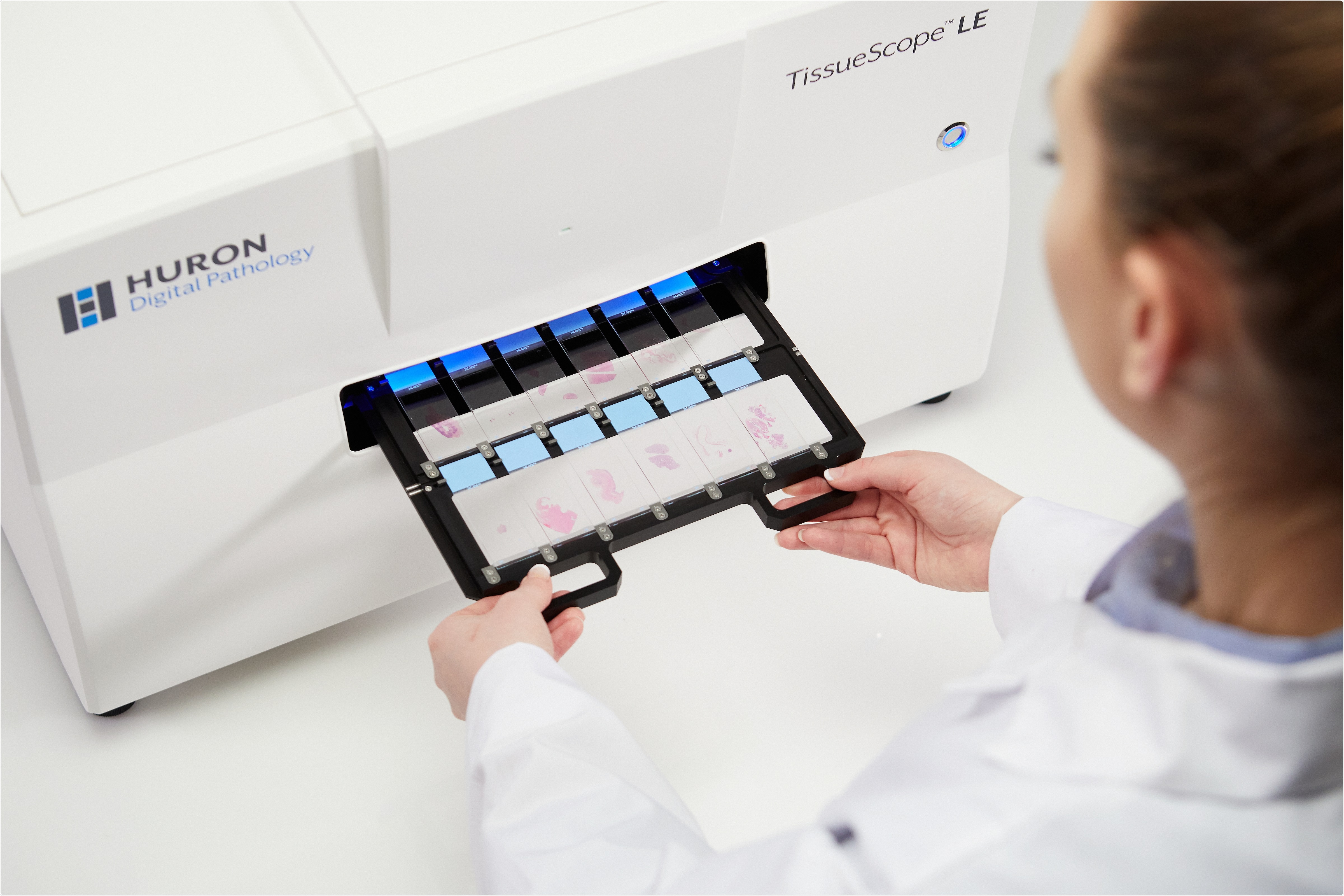

Image credit: Huron Digital Pathology.
Another major challenge for deep learning in digital pathology is the dimensionality of the problem. Pathology images are extremely large, i.e., larger than 50,000 by 50,000 pixels. Deep networks, however, can only handle small input images, i.e., not larger than 300 by 300 pixels. Down-sampling images (making them smaller) would result in a loss of information.
A further obstacle in training deep networks is that they generally perform well if they are fed with “balanced” data, that means having almost the same number of images for any category you need to recognize. Imbalanced data impedes generalization, which means the network may make grave mistakes after training.
A final problem worth mentioning is the so-called “adversarial attacks” when someone with knowledge of the system, or exploiting the presence of artefacts and noise, could eventually fool a deep network into a wrong decision, an effect that is extraordinarily important in medical imaging; we cannot allow algorithms to be fooled when we are dealing with people’s lives.
Intensive research is being conducted at many fronts to find solutions for these and other challenges. Among others, one potential solution being worked on is “transfer learning”, to learn in a different domain and transfer the knowledge into the medical domain.
Can we teach the AI with millions of labelled natural photos (e.g., cars, faces, animals, buildings) and then use the acquired knowledge on histopathology images? Other potential remedies are to inject domain knowledge into deep networks, training “generative” models that do not directly deal with classification, and combining deep solutions with conventional algorithms and handcrafted features.
How would the use of medical imaging interplay with other histopathological tests? Could they be replaced with a simple image search?
Definitely not. Image searches would be a new facilitator that will assist the pathologist and provide new insights. Presently, we may not have an accurate understanding of where the image search would fit most usefully, but we know for sure that the pathologist must remain in the center of all processing.
The tasks that we assign to the AI and computer vision will be widely specialized and customized; they naturally cannot render other existing (non-AI) technologies and other modes of tests useless. It’s all about complementing existing procedures with new insights, and not replacing them; well, at least this should be the guiding attitude.
Please give an overview of your recent research to advance this field and the techniques that you have used.
At Kimia Lab, we have been working on a multitude of techniques, from deep networks to support vector machines, from local binary patterns to Radon transform, and from deep autoencoders to dimensionality reduction.
Our research philosophy is unconditionally pathologist-centric; we are there to design AI techniques that serve the pathology community. We are convinced that this is the right way of deploying AI, namely as a smart assistant to the pathologist and not a competitor.

Image credit: Huron Digital Pathology.
We introduced a fundamental shift in our research and refrained from engaging in yes/no classification and instead are conducting many experiments to understand the polymorphic nature of tissue recognition before we attempt to design a final chain for the clinical workflow.
In addition, we have not lost our focus on non-AI computer vision for there are a lot of conventional methods that exhibited mediocre performance back in the day, but can now be rediscovered as partners to the powerful AI by relying on the faster computational platforms available.
What advantages are there to the Radon transform that you used in your research?
This is one example of our efforts to not lose sight of well-established technologies. Radon transform is an old technique and has enabled us, among others, to do computed tomography.
Projections in small and large parts of the image can provide compressed information about tissue characteristics and where significant changes occur. They can serve as inputs to AI algorithms to provide additional information in a setting where multiple technologies work together.
Radon transform is not only a mathematically sound technology but, in contrast to deep networks, is interpretable. Why a specific image is selected can be relatively easily understood when we acquire Radon projections, whereas the millions of multiplications and additions inside a network do not offer any plausible way for understanding why a specific decision has been made.
However, we need deep architectures to learn. Hence, combining the old and the new is something we are heavily investing in.
How can artificially intelligent search and categorization of medical images accelerate disease research and improve patient care?
If we abandon the classification-oriented AI (making yes/no decisions), which aims at eliminating the diagnostic role of the pathologist, then we are left with mining-oriented AI that identifies and extracts similar patterns from large archives of medical images.
Showing similar images to the pathologist when s/he is examining a new case is not something extraordinary, unless the retrieved cases are annotated with the information of evidently diagnosed patients from the past.
Then we have something that has never been done before: we are tapping into the collective wisdom of the physicians themselves to provide them with computational consultation. Consulting other pathologists for difficult cases is a common practice.
However, the image search will give us access to “computationally” consult hundreds of pathologists across the country (and the globe) through digital records. This will expedite the process, reduce error rates, save lives, release valuable pathologist time for other tasks (e.g. research and education), and finally save costs.
Where do you see the future of machine learning with regards to medical imaging?
Perhaps many of us are hoping that radiogeomics would be a revolutionary change in disease diagnosis that among others would make the biopsy superfluous, as some researchers audaciously envision.
However, for the foreseeable future, we should rather look at “consensus building”. The manifestation of medical diagnosis difficulty is clearly visible in the so-called “inter-observer variability”; doctors cannot agree on a diagnosis or measurement when given the same case.
For some cases like breast and lung cancer the disagreement can approach and even exceed 50% when the exact location of the malignancy is involved. Using AI for identifying and retrieving similar abnormalities and malignancies will open the horizon for building consensus.
If we can find several thousand cases of the past patients that can be confidently matched with the data of the current patient, then a “computational consensus” is not far away. The beauty of it is, again, that the AI will not be making any diagnostic decision but just making the existing medical wisdom accessible, the wisdom that is currently fallow under terabytes of digital dust.
As the technology advances, will there be a need to pathologists in the future?
The tasks and workload of the pathologists will certainly go through some transformation but the sensitive nature of what they do on one side, and the breadth and depth of knowledge they hold, on the other side, makes them indispensable as the ultimate recognition entities.
It is imaginable in near future that, by employing high-level visual programming languages, pathologists design and teach their own AI agents for very specific tasks. Not engineers, not computer scientists, it will be the pathologists that would have the medical knowledge to be in charge of exploiting the AI capabilities.
Where can readers find more information?
A good read on digital pathology is the book “Digital Pathology” by Pantanowitz and Parwani. A good technical read on foundations of AI is “Neural Networks And Learning Machines” by Simon Haykin. Another technical read is “Deep Learning” by Goodfellow and others.
To get an overview, “A survey on deep learning in medical image analysis” by Litjens and others is an up-to-date article. An article by myself and Dr. Pantanowitz (University of Pittsburgh Medical Center) titled “Artificial Intelligence in digital Pathology – Challenges and Opportunities” will soon appear in the Journal of Pathology Informatics.
About Prof Hamid Tizhoosh

Dr. Hamid R. Tizhoosh is a Professor in the Faculty of Engineering at University of Waterloo since 2001 where he leads the KIMIA Lab (Laboratory for Knowledge Inference in Medical Image Analysis).
Before he joined the University of Waterloo, he was a research associate at the Knowledge and Intelligence Systems Laboratory at the University of Toronto where he worked on dynamic bandwidth allocation using AI methods such as reinforcement learning.
Since 1993, His research activities encompass artificial intelligence, computer vision and medical imaging. He has developed algorithms for medical image filtering, segmentation and search. He is the author of two books, 14 book chapters, and more than 140 journal and conference papers.
He has also filed 5 patents in collaboration with WatCo (Waterloo Commercialization Office). Dr. Tizhoosh has extensive industrial experience and has worked with numerous companies. Presently, he is the AI Advisor of Huron Digital Pathology, St. Jacobs, ON, Canada.