Researchers Meghan May, Bahman Rostama and Ryan F. Relich from the Department of Biomedical Sciences, College of Medicine, University of New England, Biddeford, Department of Pathology and Laboratory Medicine, Indiana and University School of Medicine, Indianapolis, IN USA, worked on the evolution of the pathogenic strains of the betacoronaviruses including the SARS CoV-2 that has caused the COVID-19 pandemic.
Their study titled, “Selectomic and Evolvability Analyses of the Highly Pathogenic Betacoronaviruses SARS-CoV-2, SARS-CoV, and MERS-CoV,” has been published prior to peer review on the open-access preprint server Biorxiv*.
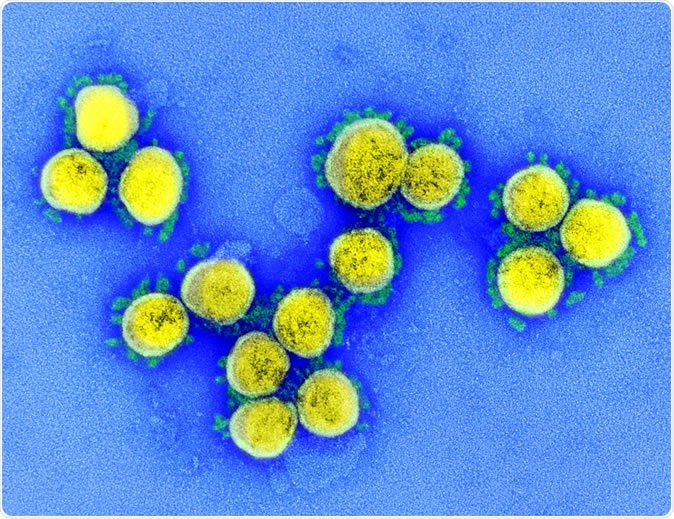
SARS-CoV-2 - Transmission electron micrograph of SARS-CoV-2 virus particles, isolated from a patient. Image captured and color-enhanced at the NIAID Integrated Research Facility (IRF) in Fort Detrick, Maryland. Credit: NIAID

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
What was this study about?
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the novel virus that causes COVID-19 disease and has resulted in worldwide pandemic affecting nearly all the.
The research team wrote, “The novel coronavirus SARS-CoV-2 is the third Betacoronavirus to emerge in the past 20 twenty years.”
This virus emerged in December 2019 with the first reported cases seen in Wuhan, Hubei province of China. This virus is said to have genomic similarities with the previous outbreak caused by SARS CoV that causes severe acute respiratory syndrome in 2003. Similarly, the MERS CoV or the Middle East Respiratory syndrome also caused an outbreak.
The team wrote that, to date, there had been seven coronaviruses that have been identified by scientists. These lead to respiratory tract infections among humans. Four of these include the “human coronaviruses (HCoVs) HCoV-229, 49 HCoV-OC43, HCoV-NL63, and HCoV-HKU1,” they wrote. These are part of several other respiratory viruses that circulate across the world during the winter months. They lead to mild upper or lower respiratory tract infections among populations. These HCoVs, however, are also capable of causing severe infections, and these are particularly severe among vulnerable populations such as those with a compromised immune system or elderly.
The SARS CoV-1 contains a single sense RNA that is positive sense and has a length ranging between 27 and 32 kbp (kilobase pairs). This RNA codes for four proteins that form the structure of the virus called the “spike, S; 93 envelope, E; membrane, M, and nucleoprotein, N,” and nonstructural proteins (NSPs) of the virus. It has an open reading frame (ORF) termed ORF1ab, wrote the researchers.
.jpg)
Novel Coronavirus SARS-CoV-2: This scanning electron microscope image shows SARS-CoV-2 (round gold objects) emerging from the surface of cells cultured in the lab. SARS-CoV-2, also known as 2019-nCoV, is the virus that causes COVID-19. The virus shown was isolated from a patient in the U.S. Credit: NIAID-RML
They added that SARS lead to chills, fever, headache, muscle pain, and generalized malaise. In some, it can lead to severe pneumonia and often become life-threatening. The case fatality rate or the risk of dying due to the infection is around 10 percent. The case fatality rate, however, increases with increasing age.
This study looked at the “genomic plasticity and evolutionary dynamics” of this class of betacoronaviruses in order to help develop diagnostic tests for molecular confirmation of the infection or develop vaccines to prevent the infection.
What was done in the study?
The team determined the selectome for the SARS-CoV-2 virus by assessing diversification (two types – pervasive and episodic) of the amino acid coding positions and their selection for the subsequent strains. They also looked at the evolution of the pathogenic betacoronaviruses to assess the movement of this virus from animals to humans (zoonotic spread or transmission). Selectomes for SARS-CoV and MERS-CoV and calculations of evolution into SARS-CoV-2 was also analyzed. Using these methods, they looked at the amino acid locations where there have been “pervasive diversifying selection or episodic diversifying selection.”
What was found?
The study showed that there was a significant site where evolution took place. This was at the “ORF1a polyprotein, the spike protein, and the membrane protein,” of the novel coronavirus or the SARS-CoV-2. They wrote that the virus is “highly plastic” (at the HCoV-OC43) as well as “highly conserved” (at the HCoV-229E). They wrote that this technique could help identify the wildlife versions of this virus that could potentially mutate into virulent forms. They wrote, “This underscores the need to conduct continuous viromic surveillance of wildlife reservoirs, as identification of coronaviruses with significantly evolvable traits are likely at higher probability for successful zoonotic spillover events.”
They noted that there was pervasive diversifying selection in ORF1a polyprotein and ORF1b protein and the S protein of MERS-CoV and SARS CoV which signified its evolvability. All three viruses showed pervasive diversifying selection. Episodic diversifying selection was seen in the same proteins of the SARS-CoV-2, they wrote.
Conclusions
The authors wrote that that “host adaptation of SARS-CoV-2 to humans is likely to lead to viral diversification parallel to that seen for SARS-CoV and MERS-CoV. This study also provides insights into the need for viral surveillance to prevent spillage of the viruses from wildlife into humans.
The authors concluded, “These findings provide a comprehensive view of zoonotic, highly pathogenic betacoronavirus evolutionary dynamics that can be directly applied to diagnostic assay and vaccine design for SARS-CoV-2.”

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
Journal reference:
- Preliminary scientific report.
Selectomic and Evolvability Analyses of the Highly Pathogenic Betacoronaviruses SARS-CoV-2, SARS-CoV, and MERS-CoV Meghan May, Bahman Rostama, Ryan F. Relich bioRxiv 2020.05.05.078956; doi: https://doi.org/10.1101/2020.05.05.078956