.jpg)
Illustration of SARS-CoV-2. Image Credit: Orpheus FX / Shutterstock
The ongoing worldwide pandemic of coronavirus disease (COVID-19), caused by the novel SARS-CoV-2, revealed a pressing need for new preventative and treatment approaches. The primary immunogenic target for such endeavors at the moment is viral S-protein due to its pivotal role in viral life-cycle and its interaction with the human ACE2 receptor.
The S-protein can be divided into two regions or domains: an N-terminal S1 domain and a C-terminal S2 fusion domain. In short, after the S1 domain binds to the host receptor, it induces the S2 domain to change its formation during the cell-virus fusion process. Furthermore, receptor binding employs the Receptor Binding Domain (RBD) in S1, which is followed by the proteolytic separation of the spike by host protease enzymes.
Such conformational plasticity is highly specific for enveloped-virus fusion-protein structure since conserved viral fusion elements have to be protected from host immune responses – retaining at the same time sufficiently steep free-energy gradient to enable fusion with the host cell.
Exposed viral elements are generally well adapted to be permissive and amenable to mutations via genetic drifting and host immune adaptation. Nonetheless, the aforementioned conformational plasticity can pose certain difficulties in the context of vaccine and drug design.
The lesson that we have learned in the elusive quest to produce a widely protective HIV vaccine has shown the significance of understanding and precisely controlling fusion protein dynamics. SARS-CoV-2 is, in all probability, no exception in this regard.
Consequently, researchers from Duke Human Vaccine Institute and Duke University in Durham, as well as from Genome Integrity and Structural Biology Laboratory at Research Triangle Park in the US, aimed to understand S-protein mobility/flexibility to inform structure-based immunogen design.
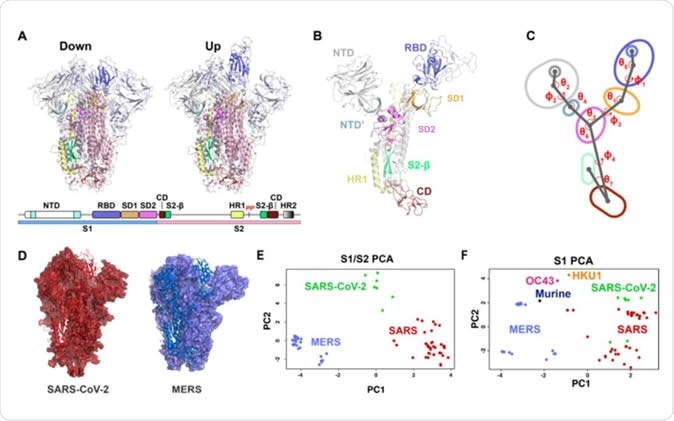
Vector-based analysis of the β-CoV S-protein demonstrates remarkable variability in S-protein conformation within ‘up’ and ‘down’ states between CoV strains. A) Cartoon representations of the ‘down’ (upper left) and ‘up’ (upper right) state structures colored according to the specified domains (lower). B) A single protomer of the β-CoV S-protein with labeled domains. C) A simplified diagram of the β-CoV S-protein depicting the centroids and vectors connecting them with the determine angles (θ) and dihedrals (ɸ) labeled. D) The SARS-2 (left; red) and MERS (right; blue) structures each with a single protomer depicted in cartoon representation and the remaining two in a surface representation. E) Principal components analysis of the SARS, SARS-2, and MERS protomers including measures between S1 and S2 domains. F) Principal components analysis of the SARS, SARS-2, MERS, HKU1, and Murine CoV protomers including measures only between S1 domains.
Designing and visualizing mutations
A structure-based vector analysis of available betacoronavirus S-protein structures was implemented as an overall approach. For the purposes of this study, the researchers have designed mutations that modify the conformational distribution of the domains in the S-protein.
The effect of their designs was visualized with the use of a structural determination pipeline depending initially on single particle analysis by a negative stain electron microscopy, enabling in turn low-cost and rapid assessment of the spike ectodomains at low resolution. This was followed by cryogenic electron microscopy for high-resolution information on mutation-induced changes.
Engineered S-protein conformations
"Our results reveal a heterogeneous conformational landscape of the SARS-CoV-2 spike that is highly susceptible to modification by the introduction of mutations at sites of contact between the S1 and S2 domains", say study authors.
Despite a general similarity in domain organization, different strains of betacoronaviruses exhibit distinct S-protein configurations. Even scarce mutations may result in dramatic shifts in S-protein structure due to a relatively small contact area between S1 and S2 subunits observed in this paper.
The conformational plasticity of the SARS-CoV-2 S-protein is shown to be greater than comparable HIV-1 Env protein. Based on this analysis, two soluble ectodomain constructs were developed by researchers, where highly immunogenic and mobile RBD is locked in either the characteristic 'down' position or is induced into the hitherto unobserved 'up' configuration. This is an entirely new discovery that stemmed from this study.
In a nutshell, the researchers presented data on modified SARS-CoV-2 ectodomain constructs in thus far unobserved stabilized conformations, with possible direct implications in applicable vaccine design via engineered coronavirus spike proteins.
Implications for vaccine development
"From the perspective of immunogen development, the constructs developed here present an opportunity to examine the ability of differentially stabilized S-protein particles to induce two different, yet important antibody responses," explain study authors.
Certain complicating factors – most notably the potential for vaccine enhancement or immune backfiring which can actually worsen the disease – may still favor the utilization of truncated, single-domain constructs that have fewer potentially weakly or non-neutralizing epitopes. However, the findings of this study are certainly in favor of an innovative approach.
"The designs presented here will allow for a detailed characterization of not only vaccine immunogenicity but also antigenicity, paving the way for next-generation vaccines for the novel SARS-CoV-2 and the eventual development of a broadly neutralizing betacoronavirus vaccine", highlight study authors in this paper available on bioRxiv preprint server.
The rational design approach presented here lays out the method for precise control of the RBD orientation distribution, allowing in turn exploratory efforts to delineate the role of conformational dynamics from the perspective of drug and vaccine development.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources