Manu Vanaerschot and colleagues say the World Health Organization (WHO) recommends that in some areas where community spread of SARS-CoV-2 is high, RT-PCR screening of just one discriminatory target is considered sufficient.
However, the current study identified a nucleotide change in an N gene primer sequence that disrupted annealing and amplification, thereby reducing diagnostic sensitivity.
The researchers say the findings strongly support the routine use of at least two targets when testing for SARS-CoV-2 using RT-PCR, even in areas where transmission rates are high.
A pre-print version of the paper is available on the server bioRxiv*, while the article undergoes peer review.
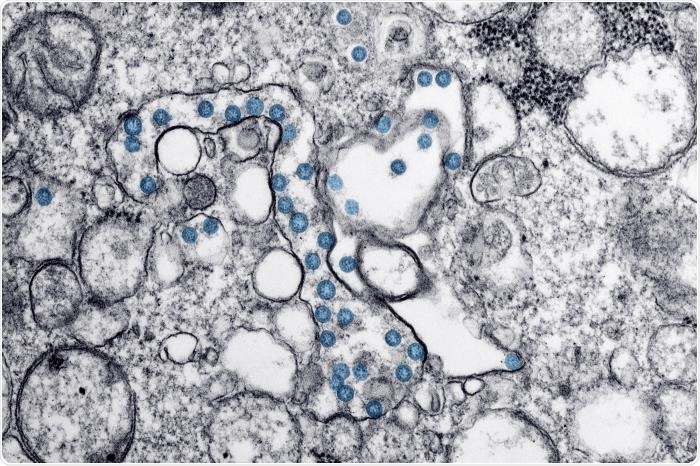
Transmission electron microscopic image of an isolate from the first U.S. case of COVID-19, formerly known as 2019-nCoV. The spherical viral particles, colorized blue, contain cross-sections through the viral genome, seen as black dots. Image Credit: CDC/ Hannah A Bullock; Azaibi Tamin

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Standard testing generally targets more than one viral gene
Routine testing for SARS-CoV-2 involves using RT-PCR to test for the viral genome in respiratory samples, and this is usually carried out using primer pairs that target more than one viral gene.
However, the WHO recommends that “in areas where the COVID-19 virus is widely spread, a simpler algorithm might be adopted in which, for example, screening by RT-PCR of a single discriminatory target is considered sufficient.”
Of the FDA-approved Emergency Use Authorizations that currently exist, 36 only screen for one target in RT-PCR assays.
The COVID-19 diagnostic laboratory at the Chan Zuckerberg Biohub and the University of California San Francisco (CLIAHUB) has been receiving samples for no-cost testing from various counties in California since April 7th, 2020.
The protocol involves assays that target the N gene and the E gene in SAR-VoV-2, and other groups commonly use these assays.
What did the study involve?
In July, the researchers identified clustering of 35 samples from Madera County that showed poor assay performance for the N gene, but not the E gene, and the team has now set out to identify potential reasons why.
The researchers assessed the concordance of the cycle threshold (Ct) values for the two assays in 3,957 SARS-CoV-2-positive tests that were conducted between May 27th and August 7th. The Ct value is a relative measure of the concentration of a target during PCR.
Of 3,629 samples that were positive for the E gene and N gene, the difference in Ct value between the N and E gene (ΔCt[N-E]) was 0.40.
The team defined two potential mutant sample sets.
The first set (set A), which comprised 42 samples collected from Madera County between June 30th and August 4th, included samples that had an ΔCt(N-E) value of 2.96 or more and an E gene Ct value 30 or less.
The team reports that in sample set A, the ΔCt(N-E) was an average of 5.35, representing an approximate 41-fold impairment of N gene amplification.
The second sample set (Set B), which comprised 19 samples collected from Madera County over the same time period, included samples that had an E gene Ct value of more than 30 and an ΔCt(N-E) of more than 2.96 or an E gene Ct value of more than 30 and no N gene Ct value detected.
Sequencing identified a mutation
Sequencing of the detected N gene fragment in samples from set A, identified 34 samples that had a single mutation in the forward primer binding site in the N gene, which corresponds to G29140T in the SARS-CoV-2 genome.
The other eight samples had a wild-type N gene fragment, indicating that the higher average ΔCt(N-E) of 5.99 that was observed in these samples was an artifact.
Of 17 randomly-selected control samples from Madera county that had an ΔCt(N-E) value of less than 2.96, none contained the G29140T mutation.
Sequencing of the detected gene fragment in samples from set B identified twelve samples that had the G29140T mutation.
Importantly, RT-PCR targeting the N gene did not detect SARS-COV-2 in five of ten 10 samples, says the team.
“Fortunately, these cases could still be recognized as infected by use of the E gene assay,” write the researchers.
To assess the effect the G29140T mutation had on PCR targeting of the N gene, the researchers synthesized a new N gene forward primer that had complete complementarity to the mutated sequence and compared the performance of this primer to the original primer among 16 mutant and 14 wild- type samples.
Among the mutant samples, the ΔCt(N-E) fell from 5.44 with the original primer to 0.19 with the mutated primer. Among wild-type samples, the ΔCt(N-E) increased from 0.46 with the original primer to 7.34 with the mutated primer.
“These data validate that the G29140T mutation is causative for the observed aberrant N gene Ct values,” says the team.
At least two targets should be used
Vanaerschot and colleagues say the findings demonstrate that even in areas where community spread of SARS-CoV-2 is high, mutations can arise that impair recognition of RT-PCR primers and decrease diagnostic sensitivity. This could lead to under-diagnosis if laboratories only used one target for SARS-CoV-2 detection.
“Our findings strongly support the routine use of at least two targets for SARS-CoV-2 detection by RT-PCR,” concludes the team.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Vanaerschot M, et al. Identification of a polymorphism in the N gene of SARS-CoV-2 that adversely impacts detection by a widely-used RT-PCR assay. bioRxiv 2020. doi: https://doi.org/10.1101/2020.08.25.265074
- Peer reviewed and published scientific report.
Vanaerschot, Manu, Sabrina A. Mann, James T. Webber, Jack Kamm, Sidney M. Bell, John Bell, Si Noon Hong, et al. 2020. “Identification of a Polymorphism in the N Gene of SARS-CoV-2 That Adversely Impacts Detection by Reverse Transcription-PCR.” Edited by Angela M. Caliendo. Journal of Clinical Microbiology 59 (1). https://doi.org/10.1128/jcm.02369-20. https://journals.asm.org/doi/10.1128/JCM.02369-20.