Several months into the COVID-19 pandemic caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), we still do not know much about the early evolutionary history and order of mutational events of the virus during this pandemic.
The lack of a closely related outgroup sequence, widespread sequencing errors, and a limited number of phylogenetically informative genetic variants in the genomes are all barriers that have made inference and rooting of the SARS-CoV-2 phylogeny complicated. Thus the traditional approach of studying viral spread and evolution in which a reliable genome phylogeny is first identified and then visible differences among sequences are mapped has not helped in identifying early mutational events in SARS-CoV-2 evolution. Viral lineage definition and cataloging are also complicated because of phylogenetic noise.
An understanding of the evolution of SARS-CoV-2 is critical to find out how, when, and why COVID-19 emerged and spread across the world and to develop more effective therapeutic approaches. Global sequencing of 1000s of genomes revealed several common genetic variants that are critical to discovering the evolutionary history of the virus and tracking its spread across continents over time. Despite that, our understanding of the fundamental details in the evolution and spread of SARS-CoV-2 remains unclear.
Using mutation order approach to map SARS-CoV-2 mutational history
Researchers from Temple University, Philadelphia, PA, recently presented the cryptic mutational history, phylogeny, and dynamics of the SARS-CoV-2 virus in a preprint paper published on the bioRxiv* server, after analyzing 1000s of high-quality genomes.
To overcome phylogenetic noise and other issues, the team used a mutation order approach that is independent of phylogeny inference as an intermediate step in the mapping of the SARS-CoV-2 genomes’ mutational history. The mutation order approach is ideal for analyzing SARS-CoV-2 genomes because the clonal evolution of the virus shows no evidence for early-stage recombination. That preserves the collinearity of genomic variants. This makes it possible to use shared co-occurrence of genomic variants and the frequencies of individuals to reliably determine the mutational history of the virus despite sequencing errors and other artifacts.
The team analyzed 29,681 SARS-CoV-2 genomes (29KG dataset), each with nearly 28,000 bases, sampled from 97 countries and 40 regions worldwide. They investigated 49 single nucleotide variants (SNVs) occurring with more than 1% variant frequency (vf > 1%).
The reconstructed mutational progression is concordant with the coronavirus sampling timing and it predicts the progenitor genome that was spreading globally months after the emergence of COVID-19. Over time, mutations in the virus gave rise to 7 significant lineages some of which originated in Europe and North America after the emergence of the ancestral lineages in China.
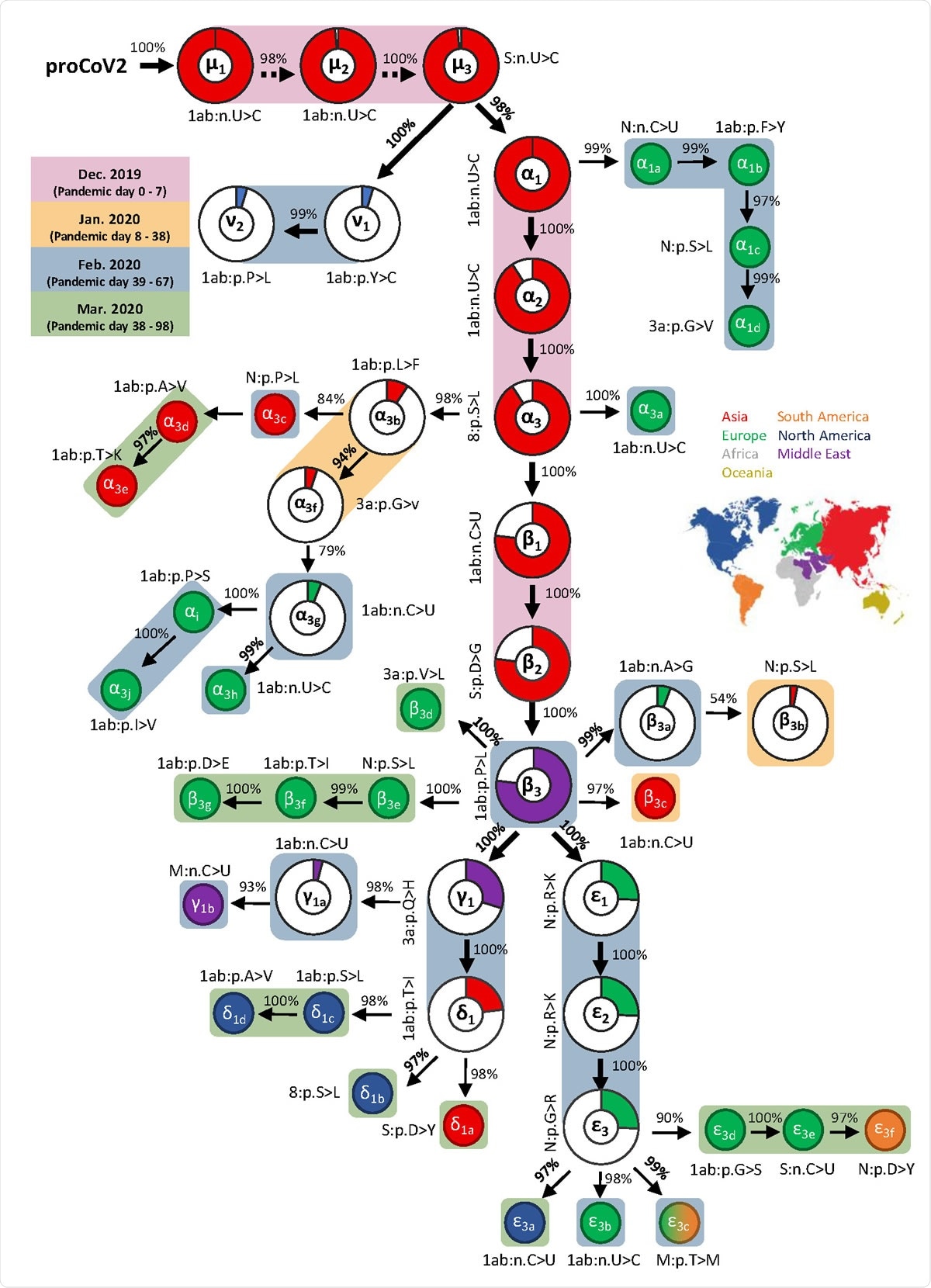
Mutational history graph of SARS-CoV-2. Thick arrows mark the pathway of widespread variants (frequency, vf ≥ 5%), and thin arrows show paths leading to other common mutations (5% > vf > 1%). The size of the pie in pie-charts is proportional to variant frequency in the 29KG dataset, with pie-charts shown for variants with vf > 3% and pie color based on the region of the world where that mutation was first observed. A circle is used for all other variants, with the filled color corresponding to the earliest sampling region. The co-occurrence index of each mutation and its predecessor mutation is shown next to the arrow connecting them. Base changes (n.) are shown for synonymous mutations, and amino acid changes (p.) are shown for non-synonymous mutations along with the gene/protein names. A rounded rectangular background indicates the earliest month in which a mutation was first found.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Spatiotemporal patterns continue to evolve as the pandemic progresses
The first mutations were sampled in Asia (China) and have the highest frequency in the 29KG dataset. Over 95% of the variants in this mutation showed a very high co-occurrence index (COI), which means in the graph, each variant was present in the genomic background of the preceding variant. The average COI for variants exceeds 96.9%, which indicates a strong signal for reliable mutation history inference.
Mutational barcoding proves that the genome signatures of North American coronaviruses are different from that of the coronaviruses prevalent in Europe and Asia that converged over time. These spatiotemporal patterns keep evolving as the pandemic progresses and can be tracked live online.
According to the team, the mutation order approach that helped them find the key mutational events, the evolution timeline, evolutionary lineages, and the spatial distribution of variants are applicable for pathogen analysis during the initial stages of outbreak. This approach is also scalable for bigger datasets since the need for more phylogenetically informative variants does not increase with an increase in the number of samples. In fact, bigger datasets provide more accurate estimates of variant frequencies and thus enable more reliable detection of lower frequency variants.
By applying this method to an extensive SARS-CoV-2 genome collection, the team was able to reconstruct the progenitor viral genome and identify mutant lineages, which enabled smooth tracking of distinct SARS-CoV-2 lineages over time and space. This helps us to have a better understanding of the past, current, and future evolution of SARS-CoV-2 and COVID-19.
An initial implementation of the SARS-CoV-2 phylogeny and global spatiotemporal patterns developed using GISAID data is available at http://igem.temple.edu/COVID-19.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
An evolutionary portrait of the progenitor SARS-CoV-2 and its dominant offshoots in COVID-19 pandemic Sudhir Kumar, Qiqing Tao, Steven Weaver, Maxwell Sanderford, Marcos A. Caraballo-Ortiz, Sudip Sharma, Sergei L. K. Pond, Sayaka Miura bioRxiv 2020.09.24.311845; doi: https://www.biorxiv.org/content/10.1101/2020.09.24.311845v1
- Peer reviewed and published scientific report.
Kumar, Sudhir, Qiqing Tao, Steven Weaver, Maxwell Sanderford, Marcos A Caraballo-Ortiz, Sudip Sharma, Sergei L K Pond, and Sayaka Miura. 2021. “An Evolutionary Portrait of the Progenitor SARS-CoV-2 and Its Dominant Offshoots in COVID-19 Pandemic.” Edited by Meredith Yeager. Molecular Biology and Evolution 38 (8): 3046–59. https://doi.org/10.1093/molbev/msab118. https://academic.oup.com/mbe/article/38/8/3046/6257226.