As the coronavirus disease 2019 (COVID-19) pandemic continues, scientists are racing to develop vaccines that immunize against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative pathogen of COVID-19, as well as antiviral drugs that help mitigate the impact of its infection. However, the process of detecting the presence of the virus within infected cells is tedious, often involving secondary antibody reactions, as is the identification of successful neutralization.
Researchers from the Texas Biomedical Research Institute, USA and the University of Alabama at Birmingham, USA, released a new study on the preprint server bioRxiv* in November 2020 reports the development of a fluorescent reporter gene-bearing recombinant form of SARS-CoV-2, which is capable of replication. This can bypass the cumbersome traditional methods of virus detection, providing a valid surrogate for viral infection.
_-_1200.jpg)
This transmission electron microscope image shows SARS-CoV-2—also known as 2019-nCoV, the virus that causes COVID-19—isolated from a patient in the U.S. Virus particles are shown emerging from the surface of cells cultured in the lab. The spikes on the outer edge of the virus particles give coronaviruses their name, crown-like. Image captured and colorized at NIAID's Rocky Mountain Laboratories (RML) in Hamilton, Montana. Image Credit: NIAID / Flickr

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
The use of reverse genetics has led to the generation of recombinant viruses. This allows the detailed study of viral biology. Using reporter genes inserted into these viruses allows the easy and rapid detection of the virus once it enters a host cell, as well as the activity of preventive or therapeutic drugs in viral infection. In fact, such efficient detection methods are ideal for high-throughput screening (HTS) of extensive libraries of biological molecules that possess antiviral activity.
Earlier studies have shown that recombinant SARS-CoV-2 (rSARS-CoV-2) viruses can be made to express fluorescent or bioluminescent genes such as mNeonGreen and GFP, or Nluc genes, respectively. These made use of laborious viral assembly, transcription and transfection steps, however.
Reporter-expressing rSARS-CoV-2
The current study uses fluorescent Venus/mCherry, or bioluminescent Nluc reporter genes, using a novel reverse genetics technique that is bacterial artificial chromosome (BAC)-based. The researchers substituted the 7a ORF with one of these reporter genes.
Characteristics of reporter rSARS-CoV-2
In Vero E6 cells, the rSARS-CoV-2 viruses containing the reporter genes are able to grow easily and produce the same type of cytopathic plaques as the wild-type virus rSARS-CoV-2/WT. The cells infected with rSARS-CoV-2 expressing these reporter genes were detected using green or red filters for Venus and mCherry genes, respectively.
These cells showed fluorescence and bioluminescence at 24 hours after infection, with the protein expression increasing until 96 hours. At this point, the cytopathic effect (CPE) led to reduced fluorescence. While the CPE was seen in cells infected with the wild-type virus, the fluorescence was not.
The red fluorescence of mCherry is advantageous over green in that it is not a common phenomenon, unlike the green fluorescence seen with a variety of genetically modified cell lines and organisms. Also, green fluorescence is absorbed by hemoglobin, while tissues show autofluorescence.
The researchers found a clear match between the expression of the reporter gene and viral replication, making it possible to quickly and easily identify infected cells. The expression of the reporter genes did not affect viral health or replication, and the virus remained genetically stable.
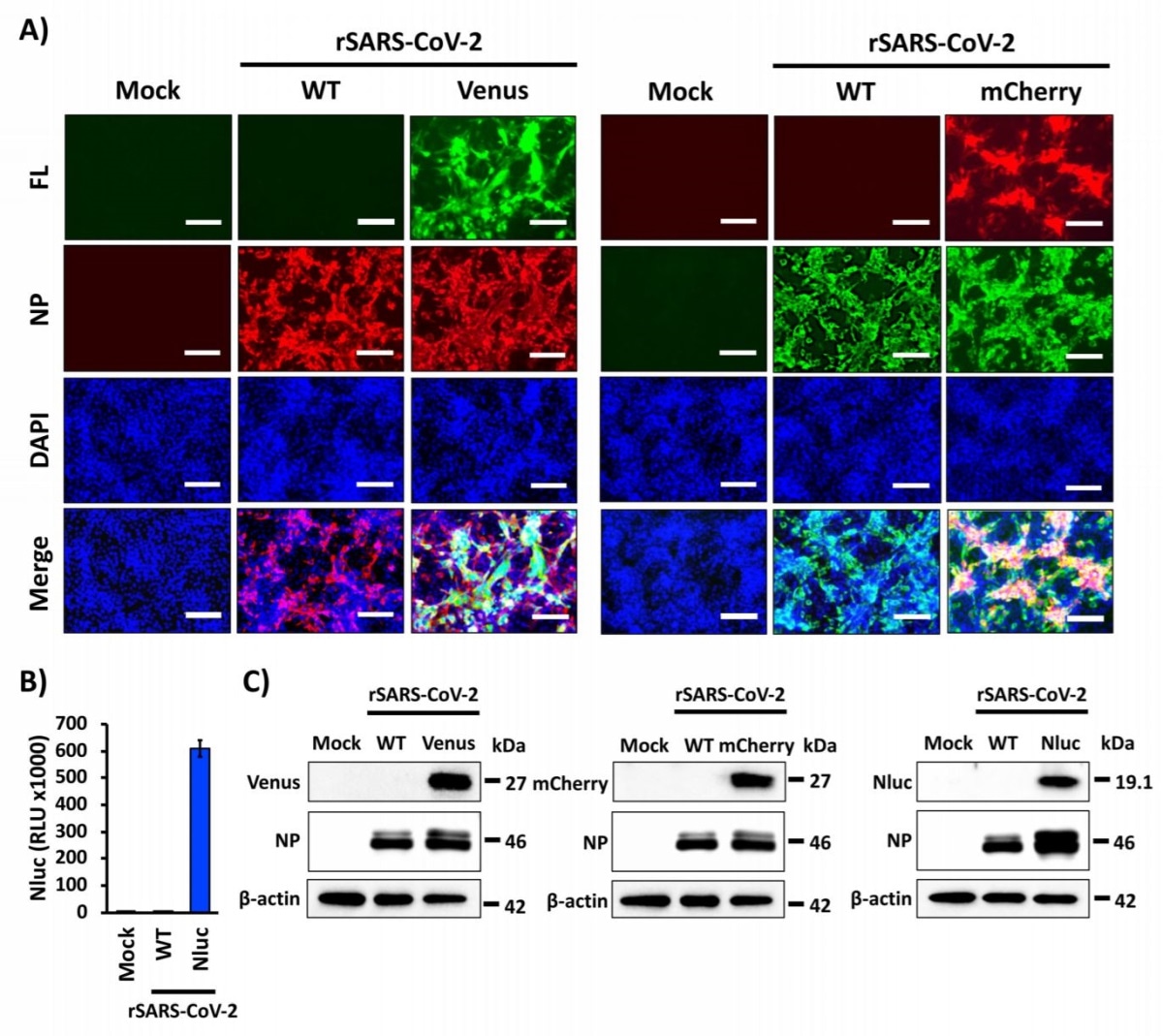
Characterization of reporter-expressing rSARS-CoV-2. A) Fluorescent expression: Vero E6 cells were mock infected or infected (MOI 0.01) with WT and Venus- or mCherry-expressing rSARS-CoV-2. At 48 h post-infection, cells were fixed and permeabilized, visualized for Venus (left) or mCherry (right) expression, and immunostained with a SARS-CoV NP MAb (1C7). DAPI was used for nuclear staining. Merged images for Venus (left) or mCherry (right), viral NP, and DAPI are illustrated. Representative images (20X magnification) are shown. Scale bar, 100 µm. B) Nluc expression: Vero E6 cells were mock-infected or infected (MOI 0.01) with WT and Nluc-expressing rSARS-CoV-2. At 48 h post-infection, Nluc expression in tissue culture supernatants was analyzed using a Synergy LX microplate reader (BioTek). C) Western blot: Vero E6 cells were mock-infected or infected (MOI 0.01) with WT and Venus (left), mCherry (center) or Nluc (right) expressing rSARS-CoV-2. At 48 h post634 infection, viral NP and reporter gene protein expression levels were analyzed using specific antibodies. An antibody against beta-actin was used as internal control. The size of molecular markers is shown in the right in each of the Western blots.
Reporter-based microneutralization assay
The study also reports a microneutralization assay to identify neutralizing antibodies or antiviral drugs with an accuracy equivalent to that of conventional microneutralization assays using rSARS-CoV-2/WT. Using the human monoclonal antibody 1212C2, which is a powerful neutralizing antibody (NAb) against SARS-CoV-2 (in vivo and in vitro), they found that its half-maximal neutralization titer NT50 against the recombinant reporter-expressing viruses were similar to the values with the wild-type recombinant virus.
The researchers also tested the inhibition of the reporter virus strains with remdesivir using a microneutralization assay. They found that the half-maximal effective concentration EC50 against all three recombinant strains was similar to that against rSARS-CoV-2/WT. This makes it easy to identify antiviral compounds based on their fluorescence or bioluminescence, in a simple assay that does not require monoclonal antibodies for virus detection.
What are the implications?
The researchers say, “Rapid and sensitive screening assays to identify compounds with antiviral activity or to assess efficacy of vaccine candidates for the therapeutic and prophylactic treatment of SARS-CoV-2 infections, respectively, are urgently needed. In this study, we demonstrate that reporter-expressing rSARS-CoV-2 546 represent an excellent option for the rapid identification and characterization of both antivirals and NAbs for the therapeutic and/or prophylactic treatment of SARS-CoV-2 infections.”
Reporter genes in viruses enable both basic viral research and studies on protein translation following viral infection. These findings show the feasibility of using this reporter gene-expressing rSARS-CoV-2 to study viral biology; to detect therapeutic activity by microneutralization assays; for in vivo studies of therapeutics; and to screen large arrays of biologicals for potential antiviral activity. Additionally, this technique could be used to produce vaccines against COVID-19.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Chiem, K. et al. (2020). Generation and Characterization of recombinant SARS-CoV-2 expressing reporter genes. bioRxiv preprint. doi: https://doi.org/10.1101/2020.11.16.386003. https://www.biorxiv.org/content/10.1101/2020.11.16.386003v1
- Peer reviewed and published scientific report.
Chiem, Kevin, Desarey Morales Vasquez, Jun-Gyu Park, Roy Neal Platt, Tim Anderson, Mark R. Walter, James J. Kobie, Chengjin Ye, and Luis Martinez-Sobrido. 2021. “Generation and Characterization of Recombinant SARS-CoV-2 Expressing Reporter Genes.” Edited by Colin R. Parrish. Journal of Virology 95 (7). https://doi.org/10.1128/jvi.02209-20. https://journals.asm.org/doi/10.1128/JVI.02209-20.