In the current COVID-19 pandemic, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has led to millions of infections worldwide, with over 1.5 million deaths.
The virus has also undergone several mutations, of which one, the D614G, has attracted much attention due to its unknown replication or transmissibility advantage on the virus. As a result, the strains carrying this mutation have become dominant wherever they have been introduced, in a very short period of time.
The entry of the virus into the host cell leads to the translation of its viral genomic RNA (gRNA) and further proteolytic cleavage. This is followed by the synthesis of subgenomic viral RNAs encoding the spike protein and other major structural proteins. These structural proteins drive virion production and are synthesized within the endoplasmic reticulum (ER) and transported to the ER-Golgi intermediate compartment (ERGIC), another subcellular compartment where proteins are processed, inside vesicles.
The mechanism of viral exit from infected cells has been thought to be a biosynthetic secretory pathway, but some have suggested another lysosomal pathway.
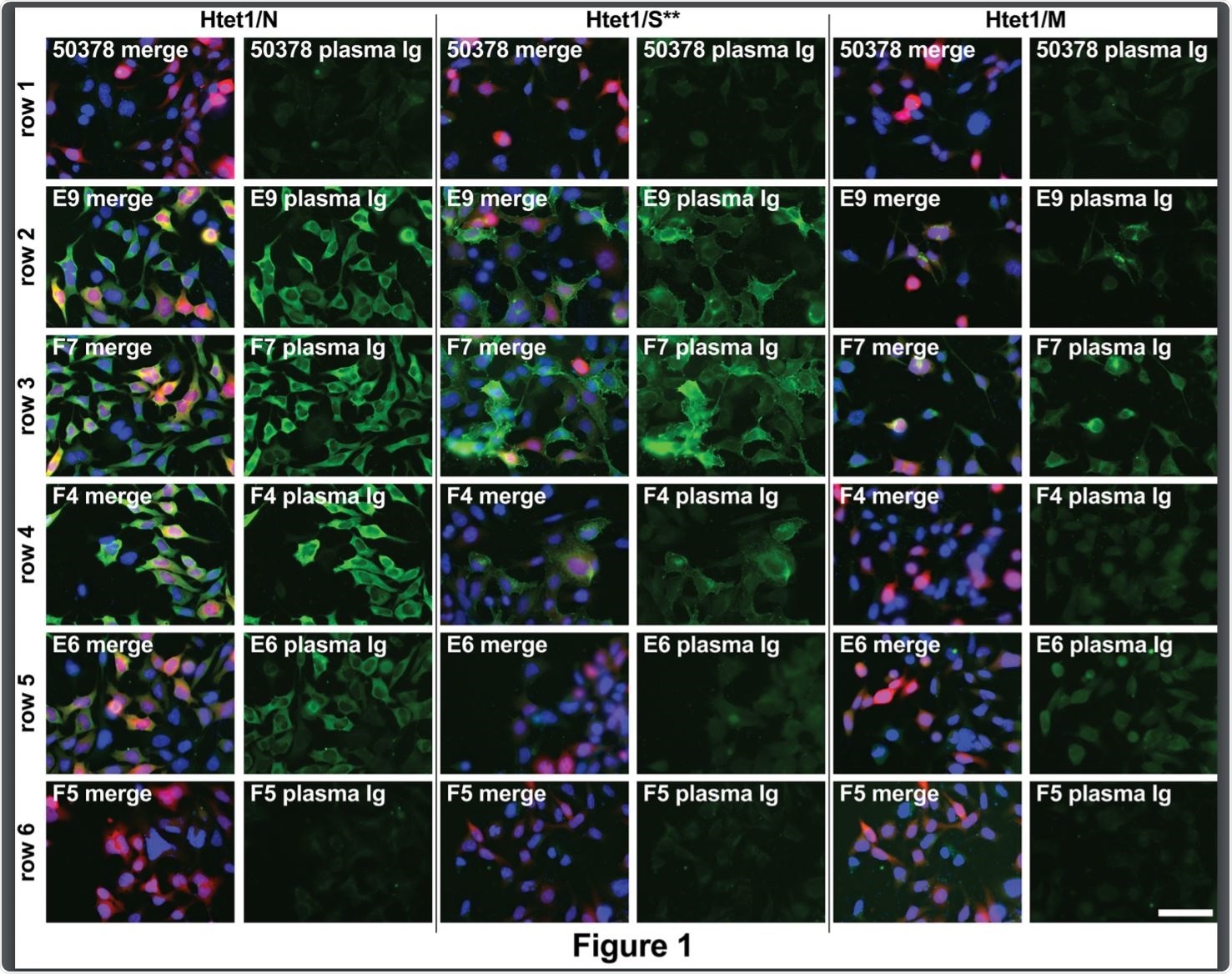
Microscopy-based serology. Image Credit: https://www.biorxiv.org/content/10.1101/2020.12.08.417022v1.full.pdf

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Effects of D614G Mutation
The emergence of the novel coronavirus was followed soon after by the entry of a strain containing the D614G mutation, at the N-terminal side of the polybasic cleavage site. This site is essential because it engages the host protease that cleaves the spike protein into its S1 and S2 subunits. This cleavage is essential for cell entry via the host cell receptors, the angiotensin-converting enzyme 2 (ACE2) molecule, and the neuropilin 1 molecule. This mutation does not appear to affect receptor binding affinity or the level of spike protein associated with the virions.
The S2 subunit engages the host cell membrane to fuse with the virus, allowing the viral particle to enter the cell. This is where this mutation causes a shift of the spike protein trafficking towards the lysosomes, but away from the biosynthetic secretory pathway. This alteration is observed both in the location of the spike protein within the infected cell and outside of it.
Thus, the researchers postulate that a single mutation related to lysosomes could account for the diverse effects of this mutation on the production of the spike protein within the infected cell, the more rapid entry of the virus into the host cell, and greater transmissibility.
The researchers prepared a cell line expressing SARS-CoV-2 spike protein in order to explore this theory. They found that the spike protein was found not only at the plasma membrane and within the Golgi apparatus, but also within large intracellular components outside the Golgi apparatus.
Their research has been published in the preprint server bioRxiv*.
In other words, the different spike variants in human infected cells are seen to belong to antigenically different groups, within different components of the cell, and eliciting different antibodies in different patients, in response to the variation in conformation.
The same plasma samples were therefore tested against a cell line expressing the D614G spike mutation. This single nucleotide change induced a remarkable alteration in the results. All four plasma types showed spikes to be localized within the non-Golgi large intracellular compartments, forming large clusters.
This could mean that this mutation increases the trafficking of the spike to these locations, or that the new conformation caused by this mutation makes it more readily detectable by antibodies in the plasma.
Spike Expression Inhibits Lysosome Function
All vaccine candidates in the forefront today are based on the spike protein. For this reason, the researchers explored these spike-associated locations. They found that they were not part of the ER, ERGIC, Golgi apparatus, endosomes, or plasma membrane. They carried markers for lysosomal membrane markers, Lamp1 and Lamp2, and lysosome-associated proteins Lamp3 (CD63) and mTOR.
Lysosomes are the end-point for the trafficking of materials that enter the cell by fluid-phase endocytosis. However, the expression of the spike protein disrupts normal lysosome function, as shown by the failure to take up test materials during the current experiment.
Earlier, in mouse hepatitis virus (MHV) infection, host cell secretory pathways were seen to change, with the KDELR protein shifting from the ER/Golgi organelles to the lysosome. In the current experiment, the researchers saw a similar shift in KDELR in some spike-expressing cells.
Lysosome Clusters Induced by Spike Expression
This was due to lysosome clustering and not fusion, as shown by vacuolin-1-induced swelling of the lysosomes. These treated lysosomes still showed both the mutated spike protein and Lamp2, showing both to be present in the same membrane, but the number of lysosomes remained constant, indicating that fusion did not occur.
Lysosomal Trafficking Independent of Microtubules and Endocytosis
Moreover, spike trafficking to the lysosomes with the appearance of spike-induced lysosomal clustering continued in the presence of molecules that inhibit endocytosis, indicating that this pathway is not active in the current case.
Secondly, it also persisted after the addition of microtubule inhibitors which affect the trafficking of certain proteins by endocytosis, and also disrupt lysosome function via the suppression of microtubule-dependent lysosome movement. Thus, it is not a function of microtubule transport.
Next, the researchers added bafilomycin A1, a powerful inhibitor of the enzyme V-ATPase. This prevents the acidification of several intracellular organelles, including lysosomes, as well as disrupting the maturation process of lysosomal degradative enzymes. Under these conditions, spike trafficking to the lysosomes failed, with the protein being localized to Golgi bodies and the cell membrane.
Conclusions
The D614G mutation has a slight effect on proteolytic cleavage at the S1/S2 boundary, but its major effect is to shift spike sorting towards the lysosome rather than to other intracellular compartments.
Overall, this is the first observable effect of the mutation and may be responsible for the increased viral fitness with this change. The possibility is that lysosomal sorting promotes the part of viral biogenesis that occurs in the lysosome. This may also be responsible for the rapid viral entry seen following this mutation.
The researchers postulate that the interactions between the spike protein and cell surface proteins speeds up viral entry, as well as enhancing the lysosomal trafficking of newly produced spike proteins. This could be via the interactions of the latter with other newly synthesized spike protein molecules in the ER, Golgi bodies, or ERGIC. In fact, it is known that this virus depends on many proteins that interact with lysosomes for the successful establishment of infection.
The ability to infect the host cell is easily prevented by suppressing trafficking between the plasma membrane and lysosomes.
These results are consistent with a lysosomal pathway of coronavirus biogenesis and raise the possibility that a common mechanism may underly the D614G mutation’s effects on spike protein trafficking in infected cells and the accelerated entry of SARS-CoV-2 into uninfected cells.”

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources