Leopoldo N. Segal and colleagues suggest lung microbes could be predictive of severe COVID infection and antibody resistance.
"These data highlight the importance of SARS-CoV-2 abundance in the lower airways as a predictor for mortality, and the significant contribution of the host cell transcriptome, which reflects the lower airway cell response to infection," wrote the researchers.
The findings could help identify patients most at-risk for poor clinical outcomes and provide alternative treatments early on.
The study "Microbial signatures in the lower airways of mechanically ventilated COVID19 patients associated with poor clinical outcome" is available as a preprint on the medRxiv* server, while the article undergoes peer review.
Classifying microbial signatures
The collected lower airway samples from patients on ventilation from COVID-19 infection during the first wave in New York City. They collected airway samples from 142 patients with COVID-19 infection.
Mycoplasma salivarium in the airways is linked to poorer clinical outcomes
Using metagenomics, the researchers linked the microbes living in the lung microbiome with patients' clinical outcomes.
Results showed that having high amounts of Mycoplasma salivarium was associated with a higher viral load of SARS-CoV-2. In addition, a limited immunoglobulin response in the lower airways correlated with increased mortality risk.
"The data presented here through the use of direct quantitative methods (RT-PCR) and a semiquantitative untargeted approach (metatranscriptome sequencing) support the hypothesis that the SARS-CoV-2 viral load in the lower airways plays a critical role in the clinical progression of critically ill COVID-19 patients," wrote the researchers.
No evidence of poorer clinical outcomes from coinfection with respiratory pathogens
While most patients were given broad-spectrum antibiotics and antifungals, there was no evidence of worsening effects from coinfection with bacterial, viral, and fungi respiratory pathogens.
To look at the risk of mortality from COVID-19 infection, the team analyzed lab cultures from 589 patients hospitalized for respiratory failure from severe COVID-19 infection.
Results showed that patients with poor clinical outcomes did not succumb to other respiratory infections. There was also no link between positive microbial cultures and mortality in severe COVID-19 infections.
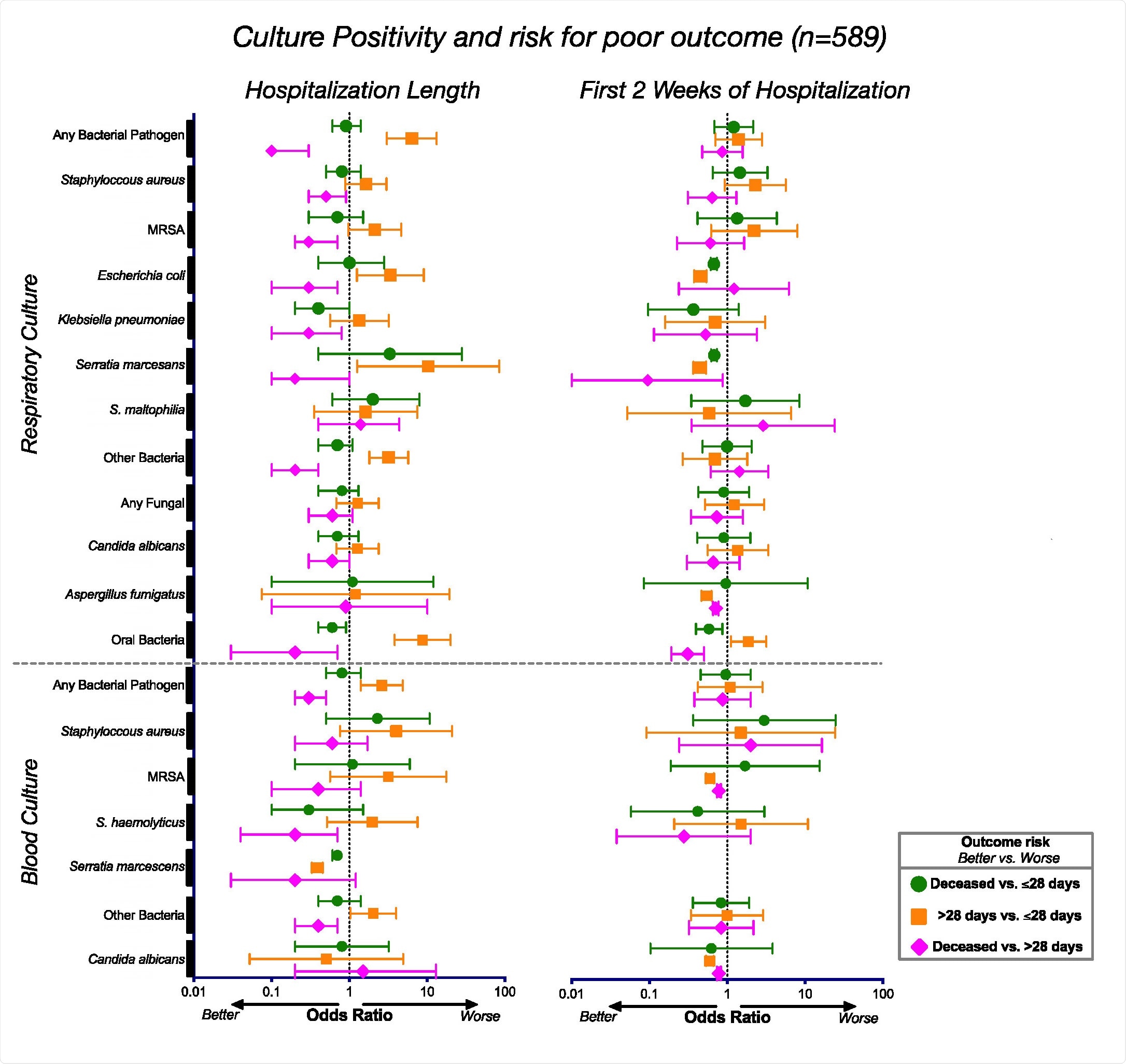
Associations between culture positivity and clinical outcome. Odds ratios and corresponding 95% confidence intervals for rates of culture positivity for the whole cohort (n=589) during the length of their hospitalization (left) and during the first 2 weeks of hospitalization (right).
Bacterial changes observed in patients ventilated for more than 28 days
Looking beyond microbes, the lower airways showed a high presence of SARS-CoV-2, which was associated with death. A tiny sample of patients had influenza A or B viruses, suggesting it unlikely that the flu occurred simultaneously with coronavirus infection.
When observing bacteria in the lower airways, metatranscriptome data found phages actively present. The researchers suggest this could be evidence that alternations in the bacterial microbiome could be happening in patients with severe COVID-19 infection.
Changes were observed in Staphylococcus phages, and Mycoplasma salivarium was actively present in patients needing ventilation for more than 28 and patients who died compared to patients who were ventilated for less than 28 days.
Microbial impact on the immune response
Patients with poor clinical outcomes expressed pathways that activated genes related to degradation, transport, as well as expressing antimicrobial resistance genes and signaling.
The researchers write:
"These differences may indicate important functional differences leading to a different metabolic environment in the lower airways that could impact host immune responses. It could also be representative of differences in microbial pressure in patients with higher viral loads and different inflammatory environments."
There was also an upregulation in the Sirtuin and Ferroptosis signaling pathways in the most severely ill COVID-19 cases. This coincided with inactivated immune response features, including phagocytes, neutrophils, granulocytes, and leukocytes. A downregulation of immunoglobulin expression levels and mitochondrial dysfunction were also observed.
Based on the data, the team suggests the lungs of critically ill patients that require ventilation from COVID-19 infection express an imbalanced state rather than elevated inflammation. Doing so appears to be predictive of worsening prognoses.
Further analysis found survival-associated differences in interferon responses. Activation of type I interferon was a predictive factor for increased mortality.
"While further longitudinal data will be needed to clarify the role of interferon signaling on the disease, the data presented here suggest that combining microbial and host signatures could help understand the increased risk for mortality in critically ill COVID-19 patients."
*Important Notice
medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.