The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) continues to wreak its havoc the world over, seemingly remaining one step ahead of scientists. With the rollout of SARS-CoV-2 vaccines, the focus shifted to making vaccines available in the vast quantities required for the global immunization effort.
However, the emergence of multiple antibody-resistant mutants has cast doubts on the success of this strategy. Especially concerning are the three variants of concern (VOCs), namely, B.1.1.7 (20I/501Y.V1), B.1.351 (20H/501Y.V2), and P1 (20J/501Y.V3).
A new research paper posted to the medRxiv* preprint server describes the changes in genomic sequence observed during the course of infection in a patient on the drug tacrolimus, along with steroids, both potent immunosuppressants, and who also received convalescent plasma treatment. These mutations were observed to occur within three weeks from infection.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Immune escape mutations in VOCs
All three VOCs carry the N501Y mutation in the viral spike protein, the viral component that mediates host cell attachment and entry. They show a high binding affinity to the host cell receptor, the angiotensin-converting enzyme 2 (ACE2), to which their increased transmissibility is attributed.
These VOCs have multiple grouped mutations affecting the N-terminal domain (NTD) and the receptor-binding domain (RBD) of the spike protein. Some of them show resistance to convalescent plasma containing polyclonal antibodies to the virus and commercially available antibody cocktails, as well as to the antibodies elicited by COVID-19 vaccines.
These VOCs are also reported to cause reinfection.
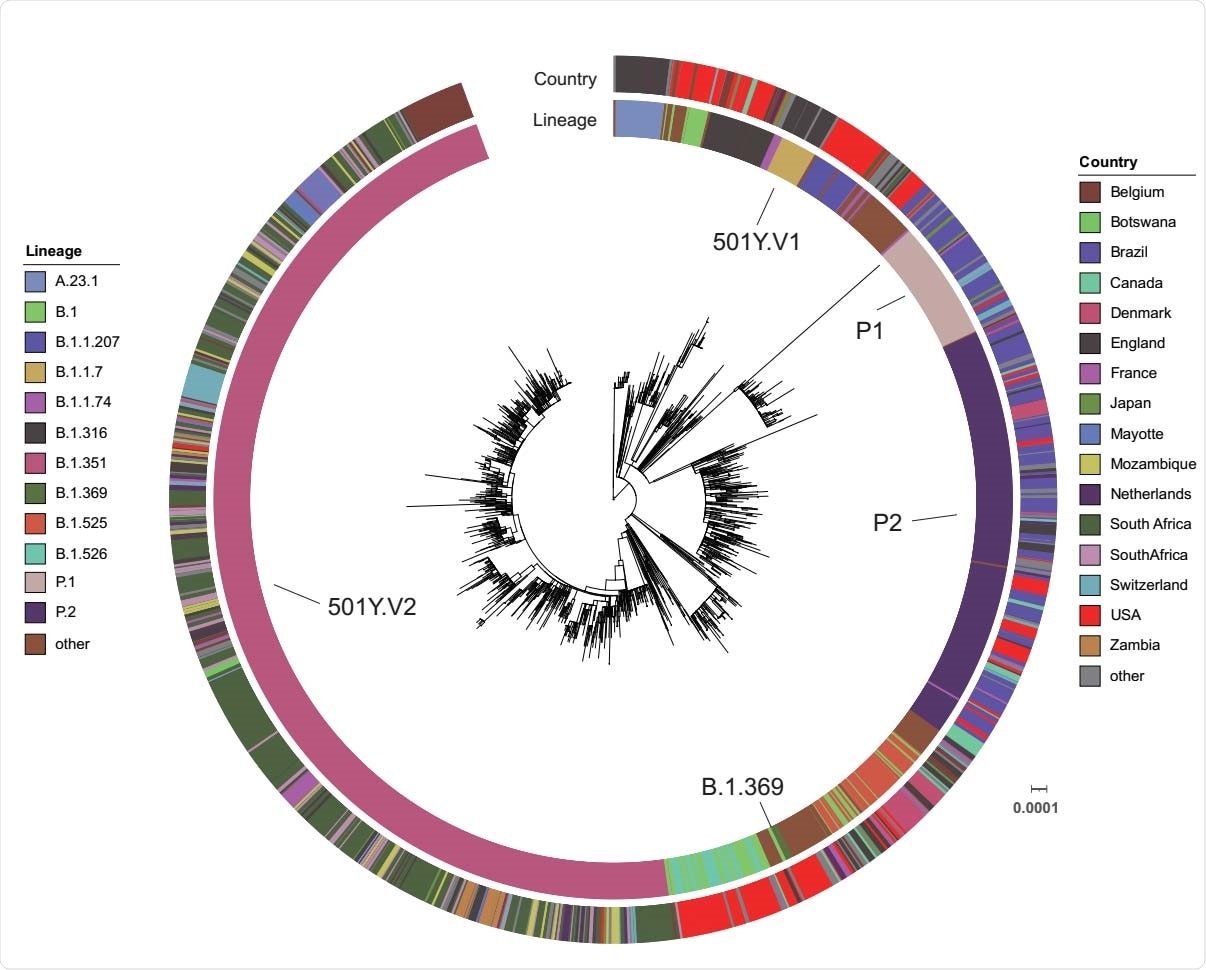
Global distribution of S protein E484K mutation. A maximum-likelihood phylogenetic tree with the 4 patient sequences (B.1.369) and 1,983 selected SARS-CoV-2 genomes from the GISAID database (date as 2/12/2021) is annotated using iTOL (https://itol.embl.de/). The scale represents 0.0001 nucleotide substitutions per site. The SARS-CoV-2 Pangolin lineage and country are illustrated as two outer rings in different colors.
Case study
The current study focused on a man in his early fifties who presented with symptoms of COVID-19 to a Northern Jersey hospital. He was a kidney transplant patient with a history of graft rejection and recurrent collapsing focal segmental glomerulosclerosis.
The patient was on three immunosuppressive medications mycophenolic acid, tacrolimus, and prednisone. His symptoms were confirmed to be due to COVID-19 by a reverse transcriptase-polymerase chain reaction (RT PCR) test. In view of his medical history and current symptoms of respiratory difficulty, he was admitted to the intensive care unit.
Mycophenolate was discontinued, but tacrolimus and prednisone were continued. Other therapies included antibiotics, convalescent plasma with a high titer of neutralizing antibodies, and the interleukin (IL)-6 inhibitor tocilizumab.
The patient continued to show worsening respiratory symptoms, and intubation became unavoidable on day 2. The patient continued to deteriorate and required extracorporeal oxygenation (ECMO) and anti-thrombotic therapy on the fifth day.
After a complicated clinical course, including septic shock, the patient was eventually taken off ECMO and mechanical ventilation. After almost 50 days, he was shifted to the step-down unit, and after 64 days, he was discharged to a care facility where he was on the ventilator only at night.
However, he died on day 94, supposedly due to poor oxygenation secondary to respiratory failure, resulting from COVID-19 pneumonia.
What did the patient's viral samples show?
The researchers found that the nasopharyngeal swabs from day 0, 34 and 41, and tracheal fluid from day 7, 21, and 27 were positive for PCR testing in this patient. They then carried out genomic sequencing of the first swab and all three tracheal samples.
The first swab and the first tracheal aspirate showed identical genomic sequences, with 14 mutations relative to the wild-type Wuhan-Hu1 virus. This put it in GISAID clade GH, or lineage B.1.369.
The second tracheal fluid sample showed the virus had acquired five spike mutations in the NTD or RBD. Of these, the Q493R and the NTD 243-244LA deletion were present in 70% of reads. The latter is in the so-called recurrent deletion region (RDR).
The ORF1a A138T mutation, and the spike 141-144LGVY deletion, were found in about 30% each, while the E484 172 K and Q493K substitutions had a frequency of ~20% and ~10%, respectively.
The third tracheal sample on day 27 showed predominantly ORF1a:A138T, and the two spike mutations – 141-144LGVY deletion and E484K substitution – at over 95% frequency.
Like the NTD 243-244LA deletion, the 141-144LGVY deletion is also in the RDR. Both these deletions are associated with resistance to neutralizing monoclonal antibodies targeting the NTD.
Origin of VOC mutations
The emergence of the VOCs may be due to the virus's opportunity to replicate endlessly within immunosuppressed patients. For instance, the New England Journal of Medicine (NEJM) recently reported the detection of antibody-resistant mutants that allow immune evasion.
The RDR deletions are found in chronic SARS-CoV-2 infections in patients with weakened immunity and thus could be a mechanism to speed up the occurrence of adaptive mutations that overcome viral proofreading activity.
The Q493K/R and E484K substitutions in the RBD are also linked to resistance to monoclonal antibodies or convalescent plasma. Resistance to neutralizing antibodies is a potentially huge challenge to vaccine-mediated control of the pandemic.
The E484K is well known for its presence in B.1.351 and B.1.1.28 variants, first detected in South Africa and Brazil and fast spreading over the region. However, it is also present in more than 17,000 variants found in more than 60 countries.
Interestingly, the paired 141-144LGVY deletion-E484K substitution became predominant in the last tracheal aspirate in the current study, indicating that it replaced the other haplotypes by compensating for lower fitness or because it resists antibody-mediated neutralization more effectively.
What are the implications?
The study findings add to the body of evidence that an immunocompromised host is likely to harbor chronic infection and develop severe or critical illness in COVID-19. Both humoral and cellular responses are probably crucial to the adaptive immune response against the virus.
Innate antiviral strategies include interferons type I and III activations, with CD8+ effector T cell activation for killing infected cells, enhanced by CD4+ T cells, which also increase B cell responses. Memory T cells preserve the memory of the virus for long-term immunity once the virus is cleared from the host.
However, the presence of chronic infection mutes this response, at least in part, and may eventually cause T cell exhaustion by the persistent stimulation of these cells by antigen presentation. This is often marked by lymphopenia.
The current patient was on an immunosuppressive regime directed against T cell function and replication and probably the proliferative burst of effector T cells. The continued use of tacrolimus and prednisone during the infection may have contributed significantly to the poor cellular immune response.
The administration of convalescent plasma in such a situation could have led to the formation of a "breeding ground," which resulted in the independent emergence of an immune escape variant.
This would account for the origin of VOCs in similar situations, which could spread in the community or via the hospital, requiring more intensive precautions and surveillance to contain such transmission.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources