The steady growth of the biotechnology industry has increased the prominence of the study of protein aggregation. Monitoring and reducing the aggregation of proteins is an important aspect of protein therapeutics because aggregation is the potential cause of immunogenicity and reduced activity. Osmolytes are commonly used as excipients for controlling aggregation. There are a number of excipients to bring about stability by preferential exclusion. Carbohydrates and polyols are a special subclass of excipients. Addition of polyols is specifically done to improve the stability and tonicity of protein formulations.
Besides prevention of aggregation, some of the other effects that the addition of osmolytes polyols has on protein formulations are retention of activity, improved protein refolding, suppression of chemical degradation, and increased thermal stability of proteins. A higher degree of thermodynamic stability can be brought about by increasing the number of hydroxyl groups on the polyols carbon chain. This results in an entropic effect when the polyol is eliminated from the protein surface.
The influence of polyol on water-peptide interactions is much greater than on non-polar group interactions that occur as a result of protein denaturation. A methodical study involving multiple proteins for determining the influence of various stereoisomers of a polyol on the thermal stability of a protein has not yet been performed. This article focuses on gaining a comprehensive understanding on the influence of the polyol size and hydroxyl group orientation on the degree of thermal stability of seven monoclonal antibodies of various isotopes. Differential scanning calorimetry (DSC) was the chosen method for monitoring thermal stability. The unfolding of proteins caused by heat denaturation was monitored by DSC so that unfolding of the domains of an antibody can be analyzed, thermal reversibility can be monitored, and formulations for excipients can be screened.
Experimental Procedures
Materials
This experiment used seven monoclonal antibodies of different isotopes: two were λIgG1 (L1A and L1B); two were κIgG1 (K1A and K1B); two were κIgG2 (K2A and K2B); and one was κIgG4 (K4). The excipients used were xylitol, mannitol, erythritol, ribitol, sorbitol, glycerol and histidine. A high throughput DSC fitted with an autosampler was used for thermal denaturation.
Sample Preparation
RODI water was used for the dialysis of each antibody, followed by preparation of buffer banks and stock solutions for xylitol, mannitol, ribitol, erythritol, sorbitol, glycerol and histidine. Antibody solutions of 1.25mg/mL in 10mM histidine of pH 6.0 were prepared using a liquid handling robot. The concentration of the titrated polyol solution was in the range of 0-350mM.
Thermal Denaturation
All the samples and their buffer controls were scanned within a temperature range of 30-95°C at 60°C/h for the thermal denaturation process. The thermal midpoints (Tm) were determined for each transition through deconvolution and baseline subtraction of the thermograms using the manufacturer-supplied software. Three or four transitions were fitted to each antibody based on the degree of resolution of the unfolding of the Fab and Fc domains from each other. One of the Fab domains and CH2 domain remain unresolved in the three unfolding transitions of L1B shown in Figure 1.
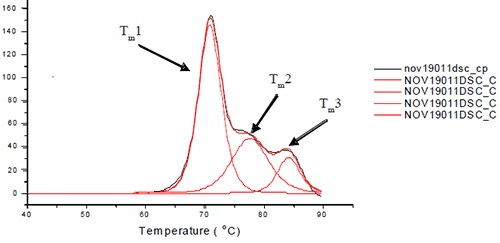
Figure 1. Representative thermogram for L1B. For L1B, Tm1 is a combination of the unfolding of the CH2 and one of the Fc domains. Tm2 is the unfolding of the CH3 domain, and Tm3 is the unfolding of the other Fc domain. Note that the order in which the domains unfold may change for different antibody isotypes.
Results and Discussion
Analyzing the thermograms for each antibody added with or without the polyol showed that the Tm value of the molecule formulated with the polyol was higher. Plotting Tm of each protein against polyol concentration revealed that the thermal stability increased linearly with polyol concentration (Figure 2).
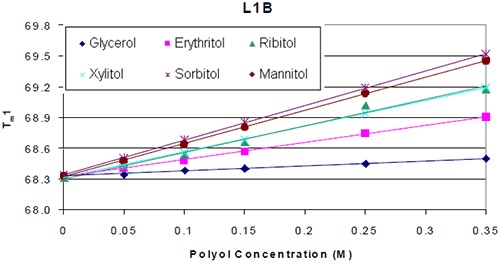
Figure 2. Thermal stability of L1B with respect to polyol concentration
As shown in Figure 2, formulations containing mannitol and sorbitol exhibited the most obvious increase in thermal stability. Ribitol comes next in the order of thermal stability increase, followed by xylitol, erythritol, and finally glycerol. The influence of using different stereoisomers on thermal stability was trivial. This was the case for the seven different antibodies analyzed. Figure 3A shows the results of the analysis of Tm increase per mole of polyol with the corresponding protein domain. Tm increase in correspondence with the polyol concentration was observed for CH3 domain and two FAB domains of K4 as well. However, the influence of the various stereoisomers of the polyol on the thermal stability was less significant. Similar results were observed for all the other six antibodies.
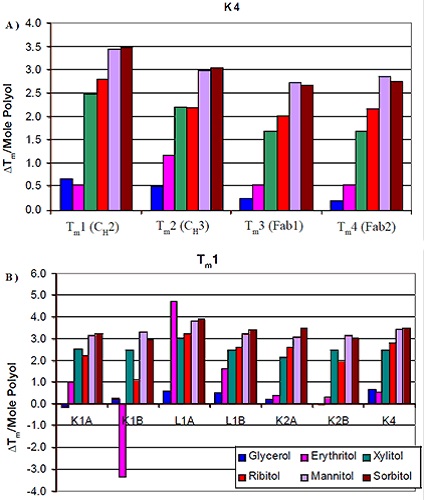
Figure 3. Change in Tm per mole of polyol as a function of domain (A) and isotype (B).
It is evident from the data that there was not much difference in the thermal stability induced by different stereoisomers on a specific domain of an antibody. Moreover, the domains more or less remained the same with respect to increased polyol concentration and Tm. Therefore, the thermal stability increase was also analyzed with regard to isotype (Figure 3B). The comparison of the first Tm for the antibodies showed a comparable increase in Tm for all of them with regard to mannitol and sorbitol concentrations, but slight variations were seen for the κIgG1 antibodies between ribitol and ylitol. The thermal stability induced by xylitol was more pronounced than ribitol for these antibodies. Also, there was variation in the increase of thermal stability corresponding to erythritol concentration.
The increase in thermal stability due to the addition of erythritol was comparatively lesser than the addition of ribitol and xylitol, but greater than the addition of glycerol. Apart from the increase in thermal stability, erythritol also induced some unique interactions. The thermal stability of K1B antibody was decreased by erythritol. Conversely, the thermal stability of the L1A antibody was increased by erythritol to a level higher than the thermal stability increase induced by either sorbitol or mannitol. The results suggested that the stability inducing mechanism of erythritol is different from other larger polyols. A single plot of the variations of the Tm value per mole of polyol against the initial Tm value of the 0mM polyol control is created by combining the data acquired for all the seven antibodies (Figure 4A).
The degree of thermal stability change caused by the addition of polyol was found be more at lower initial transition temperatures than at higher initial transition temperatures. Like earlier results, the thermal stability induced by various stereoisomers did not vary significantly. The next step was normalizing the data for the number of hydroxyl groups present in the polyols. After normalization, the variations in the thermal stability increase imparted by the addition of mannitol/sorbitol began to reduce in comparison with xylitol/ribitol (Figure 4B).
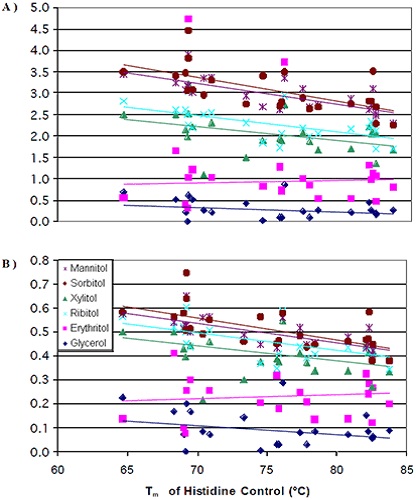
Figure 4. Tm of the histidine control versus change in Tm per mole of polyol (A) and per mole of hydroxyl groups (B)
However, the reason behind the thermal stability induced by erythritol and glycerol being very less compared to other polyols, and the variation of thermal stability induced by polyol based on the initial transition temperatures remained unclear. It has been demonstrated that the addition of polyol osmolytes makes the denatured state less stable through preferential exclusion. Based on the experiments conducted by Liu and Bolen, the amino acid side chains tend to interact more with osmolytes than water. The preferential interaction of the peptide backbone with water rather than osmolytes decreases the stability of the denatured state. It can be assumed that the interaction with water could be similar to all polyols due to their similar structure. However, their size is the decisive factor behind the variations in the thermal stability increase induced by them.
It is hypothesized that the interaction with the peptide backbone by the smaller polylols that exhibit less steric hindrance was higher, thus inducing a more modest thermal stability increase. The exclusion of smaller polyols from the peptide backbone was to a lesser degree. Protein folding relies heavily on the hydrogen bonding and the hydrophobic effect. Proteins having a less compact structure with weaker hydrogen and hydrophobic bonding interactions tend to show a lower Tms. The protein structure becomes compact when the polyol is added, thereby reducing the exposure of the peptide backbone to the polylol and compressing the protein structure further. Proteins can be compressed further if they have lower initial Tms, thus experiencing a greater thermal stability increase upon polyol addition.
Conclusion
The experimental data obtained are in line with the findings of Oakenfull, Smith and Back, and Morikawa and Gekko. As per their findings, Tm increases linearly with the concentration of alcohol. The presence of more alcohols per molecule in larger polyols makes them to influence the protein stability. Testing seven antibodies under similar conditions has revealed that stabilization mechanism is not protein specific. A notable observation is that increase in Tm induced by polyol is more for proteins having lower initial Tm when compared to proteins with a higher initial Tm.
Consequently, the structure of antibodies with a higher initial Tms is more rigid and compressed and has stronger hydrogen and hydrophobic bonds. Therefore, the addition of polyols does not induce any favorable benefits in such proteins. On the contrary, more flexible structure with weaker intramolecular bonds is observed in proteins with lower initial Tms. Therefore, there is more room for the peptide backbone to compress and lesser likelihood of polyol interaction when polyol is added to such proteins. As a result, their hydrogen and hydrophobic bonds become stronger than that of proteins with higher initial Tms.
Acknowledgements
Produced from content authored by David Sek; Donna Luisi; Nicholas Warne from Malvern Panalytical
References
- Rosenberg, A. AAPS Journal (2006) 8(3), E501-E507.
- Runkel, L. Meier, W. Pepinsky, R., et al., (1998) Pharm. Res. 15, 641-649.
- Townsend, M., and DeLuca, P., (1990) ¬J. Pharm. Sci. 79, 1083- 1086.
- Wang, W. (1999) Inter. J. Pharm., 185, 129-188.
- Gekko, K. and Timasheff, S. (1981) Biochemistry 20, 4667-4676.
- Gekko, K. and Timasheff, S. (1981) Biochemistry 20, 4677-4686.
- Mishra, R., Seckler, R., and Bhat, R. (2005) J. Biol. Chem. 280(16), 15553-15560.
- Back, J., Oakenfull, D., and Smith, M. (1979) Biochemistry 18(23), 5191-5196.
- Arakawa, T. and Timasheff, S. (1982) Biochemistry 21, 6536-6544.
- Singh, S. and Singh, J. (2003) AAPS PharmaSciTech (2003) 4(3), 1-9.
- Gekko, K. and Morikawa T. (1981) J. Biochem 90, 39-50.
- Gekko, K. and Morikawa T. (1981) J. Biochem 90, 51-60.
- Tischenko, V., Zav’yalov, V., Medgyesi, G., Potekhin, S. Privaloav, P., (1982) Eur. J. Biochem. 126, 517-521.
- Remmele, R., Saraswaty, D., Bhat, D., Phan, D., and Gombotz, W., (1999) Biochemistry, 38, 5241-5247.
- Sanberg, A. Leckner, J. Shi, Y. et. al (2002) Biochemistry 41, 1060-1069.
- Lepock, J. Ritchie, K., Kolios, M. (1992) Biochemistry, 31, 12706-12712.
- Remmele, R., Nightlinger, N., Srinivasan, S. and Gombotz, W. (1998) Pharm. Res. 15(2), 200-208.
- Chan, B., Randall, C., and Lee, Y., (1993) Pharm. Res. 20, 1478-1483.
- Liu, Y. and Bolen, D. (1995) Biochemistry, 34, 12884-12891.
About Malvern Panalytical

Malvern Panalytical provides the materials and biophysical characterization technology and expertise that enable scientists and engineers to understand and control the properties of dispersed systems.
These systems range from proteins and polymers in solution, particle and nanoparticle suspensions and emulsions, through to sprays and aerosols, industrial bulk powders and high concentration slurries.
Used at all stages of research, development and manufacturing, Malvern Panalytical’s materials characterization instruments provide critical information that helps accelerate research and product development, enhance and maintain product quality and optimize process efficiency.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.Net which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.