The success or failure of the development of a biopharmaceutical product is governed by protein stability. Protein stability is considered a critical parameter during manufacturing, production, formulation, long term storage, efficacy, and delivery to patients.
Highly stable proteins can be produced more cost-effectively, may have fewer problems during the manufacturing process, and are likely to remain functional during formulation and storage without any chemical aggregation or alteration.
In the “Quality by Design” (QbD) approach to biopharmaceutical development, the characterization of stability is part of the assessment of the 'drugability' or 'developability' of a promising drug candidate and during process development and manufacturing.
In addition, stability data is used in 'fingerprinting' and higher order structure (HOS) characterization employed for manufacturing support, biosimilarity, and biocomparability. Besides these, protein HOS characterization is increasingly required in regulatory submissions for new biosimilars and biopharmaceutical drugs.
Proteins have complex nature, which underscores the importance of biophysical tools in the characterization of biopharmaceutical products. A number of biophysical tools are employed to evaluate protein stability, including, but not limited to:
- Dynamic and static light scattering (DLS and SLS)
- Circular dichroism (CD)
- Fourier transform infrared spectroscopy (FTIR)
- Size exclusion chromatography (SEC)
- Size exclusion chromatography–multi-angle light scattering (SEC-MALS)
- Analytical ultrafiltration (AUC)
- Intrinsic fluorescence (IF)
- Differential scanning fluorescence (DSF), and
- Differential scanning calorimetry (DSC)
Although these biophysical assays considerably contribute toward biopharmaceutical development, the characterization of thermal stability by DSC is very important. In a 2015 article concerning biophysical methods for monoclonal antibody higher order structure characterization, Gokarn et al. stated: "DSC remains as an unparalleled technique to assess the thermodynamic stability of proteins in a given buffer condition”[1].
This article shows the application of DSC to assess the thermal stability of protein biopharmaceuticals (mainly antibodies), and how it serves as a tool in the development of biosimilars, and as a HOS characterization tool for the comparability of biopharmaceuticals (effect of process changes, batch-to-batch comparison, and so on).
Differential scanning calorimetry (DSC)
The microcalorimetry technique, DSC is used to characterize the conformational and thermal stability of biopolymers such as proteins, lipids, nucleic acids, etc. [2-7]. This technique is capable of measuring heat capacity as a function of temperature.
This whitepaper describes DSC instruments that are used for protein characterization. These include “power compensation” instruments, with the biopolymer in solution, and placed in a matched reference cell filled with buffer and a fixed-in-place sample cell.
Next, the heat capacity (Cp) signal from a sample cell is compared against the reference cell. As the cells’ temperature is increased, the temperature difference between the sample and reference cells is constantly determined and calibrated to power units.
Furthermore, DSC is a 'forced degradation' assay – that is when the protein is subjected to increasing temperature, it starts to unfold and the protein’s Cp increases, as illustrated in Figure 1.
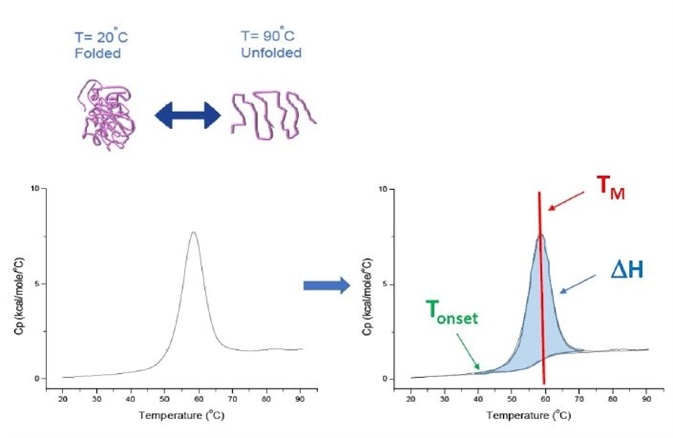
Figure 1. How DSC works. The heat capacity (Cp) changes as a protein thermally denatures. The DSC experiment starts at a temperature at which the protein is primarily folded in its native conformation. With increasing temperature, at some point the protein will begin to unfold/ denature (Tonset) and the Cp increases. At the temperature where 50% of the protein is in its native conformation, and 50% is denatured, the Cp will reach its maximum value - this is the thermal transition midpoint or TM. Above the TM the protein will be primarily denatured, and at the end of the DSC experiment all of the protein will be in its unfolded conformation. Experimental parameters for DSC include Tonset, TM, and the unfolding enthalpy (ΔH)
The heat capacity change is directly measured by DSC without the use of fluorescence or other probes or labels. For proteins that denature reversibly, the thermal transition midpoint (TM), also referred to as melting or denaturation temperature, is the temperature at which half of the protein is in its folded or native conformation, and half of it is in its unfolded or denatured conformation. The TM is regarded as the 'peak' of a DSC thermogram.
TM is a good indicator of thermal stability – when the TM is higher, the protein is more thermally stable. Antibodies are multi-domain proteins that possess more than one peak on a DSC thermogram, which means for each domain, more than one TM can be determined (Figure 2).
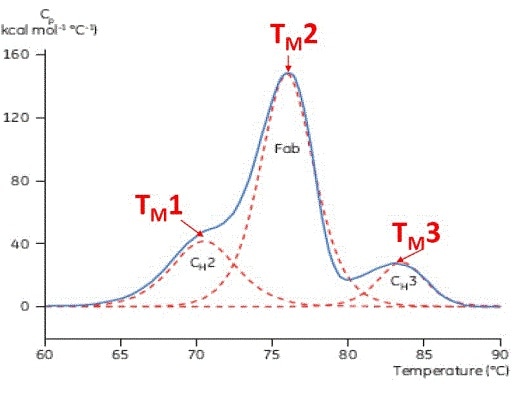
Figure 2. Representative DSC thermogram of a monoclonal antibody, with CH2, Fab, and CH3 domains identified. The dashed red lines are the deconvoluted peaks of each domain transition, with the three TMs indicated.
Protein stability, including the unfolding enthalpy (ΔH), can be characterized and rank-ordered using other parameters provided by DSC. The unfolding enthalpy (ΔH) is calculated by the area under the curve. Protein unfolding is an endothermic process, because energy input is required to break the secondary non-covalent bonds that ensure that the protein remains folded.
In addition, DSC determines ΔCp (heat capacity change of unfolding), the Tonset (start of unfolding), and T1/2 (width at 1/2 the peak height, which indicates the shape of the unfolding thermogram). Any combination of these parameters can be determined by DSC analysis.
Majority of proteins denature irreversibly and are likely to precipitate or aggregate following thermal denaturation. The DSC analysis of irreversible protein denaturation provides parameters, including TM, that are not actual thermodynamic parameters, but the rank-ordering of TM from the DSC analysis of
irreversibly denatured proteins continues to be useful for screening qualitative parameter stability.
The MicroCal VP-Capillary DSC system[8,9] offered by Malvern Panalytical is an automated DSC that is specifically designed for TM screening as well as thermodynamic characterization of biopolymers and proteins in solution.
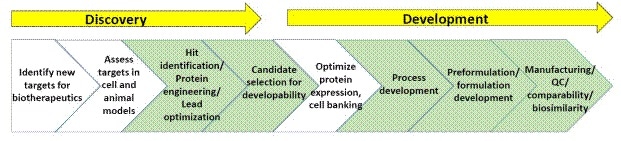
Figure 3. General schematic of processes within biopharmaceutical discovery and development
A schematic of biopharmaceutical discovery and development processes is shown in Figure 3. The sections highlighted in green are the points where biophysical characterization, including stability assays, is generally used (A list of suggested reading on biopharmaceutical discovery and development is given at the end of this article).
When developing a preferred biopharmaceutical, researchers first look for highly stable biomolecules during candidate selection, and if required, they may have to introduce more stability through protein engineering.
During the purification process, the protein is taken away from conditions where it is active, correctly folded, and stable, and as a result, additives, buffers, purification, and storage conditions have to be used so that the protein remains as stable as possible during this process.
It is important to ensure that formulated subcutaneous (SC) protein drugs remain stable and unaffected at extremely high protein concentrations (sometimes about 100 mg/mL) inside their container closure, (such as prefilled syringe, or vial) for a number of years.
When protein molecules are subjected to stresses like pressure, chemicals, heat, high concentration, mixing, and pH changes - any and all which can take place during the production and formulation of biopharmaceuticals - the protein conformation favors the unfolded or denatured protein.
Additionally, proteins in solution are also sensitive to modifications like oxidation and deamidation that also lead to inactive, denatured proteins. In a protein biopharmaceutical, modifications, including denaturation, may lead to aggregate formation that is either nonfunctional or has reduced efficacy as a drug.
Most importantly, aggregation of proteins could cause a fatal immunogenic response in patients. Therefore, to achieve a more cost-effective production, and an effective, successful, safe drug product, a stable protein may be used as the basis for a biopharmaceutical.
The most basic component of the protein structure and polypeptide chain is a protein's primary (1°) structure, or the sequence of amino acids. Beyond this, the three-dimensional structure of protein, also known as higher order structure (HOS), should be understood and characterized.
Three levels of protein HOS exist:
- Secondary (2°), referred to the local folding patterns of a protein’s primary structure, including random coils, turns, α-helix, and β-sheet
- Tertiary (3°), referred to a protein’s final 3D structure, originating from a range of secondary structural elements; and
- Quaternary (4°), which describes structures involving the interaction of two or more different or identical polypeptide chains
DSC refers to the measure of the conformational changes and stability in quaternary and tertiary structure that take place during thermal denaturaton of a protein and provides an indication of how extrinsic and intrinsic factors influence a protein's stability.
As a predictor of long-term stability, DSC is regarded as the best quantitative assay for thermal stability employed in the characterization of biopharmaceutical proteins [1,10-14]. TM from DSC is a parameter that is often employed to rank-order stability during the selection of candidates (developability), process development, and formulation screening, with more stable proteins with a higher TM.
ΔCp, T1/2, Enthalpy (ΔH), and Tonset from DSC are also employed in rank-ordering stability, quantitative analysis of protein unfolding, validation of DSC data, and higher order structure 'fingerprinting'[10-14].
If the protein is highly similar or the same (Figure 4), DSC analysis of a protein in defined solution conditions will be quantitative and reproducible. This means, there will be a reproducible pattern in DSC thermograms, and parameters including ΔH, TM, and Tonset, will be well within an acceptable range [12-14].
If the DSC fit parameters change and the compared thermograms are different, it indicates that events were taking place such as protein misfolding, aggregation, degradation, post-translational modification, change in solvent, or other higher order structure changes that impact the conformational stability.
DSC provides quantitative and reproducible data, making it an important assay in HOS and comparability analysis for product evaluation during development (including site-to-site and batch-to-batch comparability), biosimilarity, and the comparison of protein variants as well as modified products (including structural changes owing to oxidation and glycosylation).
Regulatory support documents also use DSC data as a HOS characterization for submissions of a new drug and biosimilar. In a survey, researchers in the biopharmaceutical sector ranked DSC as a "very useful" to "extremely useful" HOS biophysical tool for formulation development, candidate selection, product characterization, process development, comparability, and biosimilarity[15].
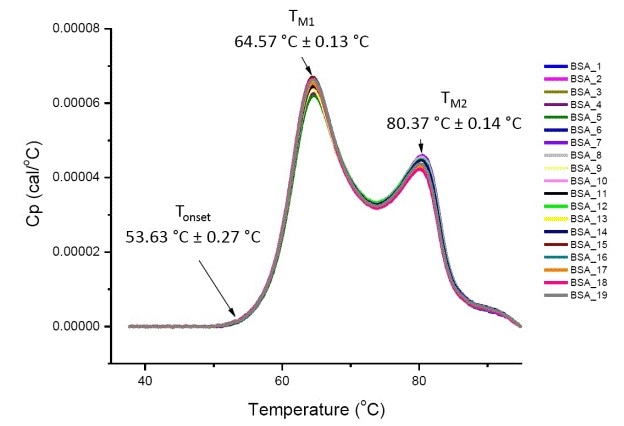
Figure 4. Nineteen DSC thermograms of bovine serum albumin (Sigma A1933, chromatographically purified) in PBS. DSC data shown after scan rate normalization, buffer-buffer subtraction, and integration baseline subtraction. Mean and standard deviations of Tonset, TM1, and TM2 are shown.
Antibodies are an example of multi-domain proteins where DSC thermograms exhibit more than a single unfolding transition (Figure 2). DSC not only characterizes and quantitates the different domains, but also establishes individual TMs for all transitions.
The thermogram peak(s) represent TM values that can be easily determined from the DSC data, without any need for complicated data analysis. DSF, CD, and IF are alternative biophysical assays that can determine TM, but these may only detect the 'most dominant' TM in the case of multi-domain proteins, or the first TM that occurs at the lowest temperature. Complex data fitting may be required to extract more than one TM from fluorescent and spectroscopic data and that TM may not be reproducible.
However, compared to other TM screening assays, DSC needs more protein sample for each scan and can also cause lower-throughput. If limited sample is available, an initial TM rank-ordering with IF or DSF can be performed and then a number of samples can be selected to confirm TM using DSC.
The TM results must be confirmed with DSC, and spectroscopy or fluorescence should not be simply relied upon to calculate TM for stability assessment. In fluorescence-based assays, it is very common to see artifacts that obstruct with the output, and because of these artifacts the TM results may seem to shift to a higher or lower value.
There are certain proteins and buffer conditions that cannot be used with fluorescence, which can make it very difficult to calculate TM differences. Spectroscopy and fluorescence do not have the ability to determine thermodynamic parameters, including the calorimetric enthalpy, whereas DSC can offer all this data as a thermal stability 'data suite'.
The biopharmaceutical industry considers DSC the “gold standard” thermal stability assay because the method:
- Is a direct measurement of protein unfolding, and hence, there is no need for tag, probe, or label. This means that there are no possible detection artifacts which are often found in spectroscopic and fluorescence assays.
- Determines heat changes related to protein unfolding.
- Determines native proteins in solution, and can be employed with almost all additives and buffers that are commonly used in development and purification of biopharmaceuticals. Some of these additives and buffers cannot be used with spectroscopy or fluorescence.
- Possess high precision temperature control and has a working temperature range of up to 130 °C, thus detecting most high TM transitions. There are other TM screening assays, but they can heat samples up to just 100 °C or lower.
- Can be easily used for experimental set-up.
- Is a 'forced degradation' assay, which means proteins no longer have to be stored in buffer before analysis. DLS and SEC-HPLC need incubating samples in buffer at increased temperature to detect conformational changes.
- Has integrated data analysis software and simple data output.
- Is information-rich, offering conformational stability, thermodynamic data, and TM determination
- Can be employed to overcome separate unfolding transitions and define simple single-domain proteins, as well as multi-domain proteins and protein complexes.
- Comes with high-throughput automation (MicroCal VP-Capillary DSC system) for fast screening of thermal stability
- Can be employed as a primary assay for the characterization of biotherapeutic thermal stability, and can even be used with other biophysical screening tools which are complementary or orthogonal, and/or to confirm other data
Biopharmaceutical comparability
Biopharmaceutical drugs, including monoclonal antibodies, are complex proteins that are expressed by bacterial or mammalian cells. As each protein is unique, elaborate studies, characterization, and optimization are required to convert it into a drug.
The preferred properties of the protein drug are characterized, including dosage, purity, and potency, all as part of discovery and early development. To investigate these characteristics, biological, biochemical, and biophysical assays have been developed and optimized across product development.
For each process, detailed HOS (including stability) and protein characterization is performed to define the protein's critical process parameters and critical quality attributes, required to regulate and validate the processes across the entire life span of products.
In the “Quality by Design” (QbD) approach for biopharmaceutical development, stability characterization and critical quality attributes are part of drug assessment during process development and manufacturing support[16,17].
During the process of protein drugs, the proteins are exposed to the following different conditions:
- Repeated freeze/thaw cycles, high pressures and agitation/mixing
- Formulation additives (excipients)
- Different types of solutions, including varying salts, buffer, and pH
- Range of protein concentrations
- Various material surfaces the protein comes in contact with (chromatography media, diafiltration/ultrafiltration membranes, chromatography media and so on)
- Exposure to proteases, oxidants, product variants, cell growth media and the likes
Any or all the above conditions can affect the interactions and forces that maintains a protein in its native folded conformation, leading to inactive or denatured protein. There may also be chemical denaturation caused by oxidation, changes in glycosylation or similar post-translational modifications, or deamidation
Usually, denatured proteins form aggregates, which presents a major challenge during the development of biopharmaceuticals. Aggregates can be termed as associated states of the protein monomer, and the formation of aggregates can be irreversible or reversible, ranging from a protein dimer to huge particles that can be seen with the naked eye.
At the most, protein aggregation lowers the efficacy and potency of drugs, leading to increased production cost. A major issue about particles and protein aggregates is their ability to promote immunogenic responses, which in severe cases may lead to patient death.
Due to this problem, protein aggregation has been extensively investigated within the biopharmaceutical sector. This includes detection of particles and aggregation, characterization and quantification during formulation and production, and a search for solutions to remove or reduce protein aggregation in biopharmaceuticals.
It must be shown that the manufactured protein drug is similar in structure, size distribution, stability, and functional and biochemical assays to:
- The same protein manufactured at various sites
- The same protein produced in earlier lots as well as the reference protein lot
- The same protein produced using a modified downstream or upstream process
- The same protein in a different formulation
- Scale-up of downstream or upstream process
Functional, HOS, biophysical, and biochemical assays for comparability or biocomparability are specifically designed to show that the protein product thus manufactured is 'highly similar' in the selected critical quality attributes, in comparison to a reference protein. The HOS of protein drug candidates should be carefully examined at all stages of development.
During process development and post drug approval, there may be some changes to the manufacturing process of a biopharmaceutical. The reasons for these changes include:
- A change in the manufacturing site
- Increase in scale
- Improvement of the manufacturing process (such as reduced cost and product yield)
- Regulatory and compliance changes
- Improvement of product stability
After making considerable changes to the manufacturing process, a comparability exercise is performed to assess the effect of the change(s) on the critical quality attributes, efficacy, and safety of the biopharmaceutical product.
The comparability demonstration does not essentially mean that there are identical critical quality attributes of the post-change and pre-change product, but rather that they are extremely similar. Moreover, product quality should not be negatively affected by manufacturing changes.
In protein therapeutics, comparability is considered a major problem and is handled by a number of international regulatory agencies [18,19,20]. As such, there is no single analytical technique that can be employed for comparability assessment of protein drugs. Comparability and HOS studies include, but are not limited to:
- DSC
- DLS
- CD
- SEC
- AUC
- Raman spectroscopy
- Fluorescence
- Microscopy
- Resonant mass measurement (Archimedes from Malvern Panalytical)
- Nanoparticle Tracking Analysis (NanoSight from Malvern Panalytical)
Bioassays, surface plasmon resonance (SPR), and isothermal titration calorimetry (ITC) are also used for monitoring the efficacy and bioactivity of proteins. Each biophysical method provides specific data, and when this information is evaluated collectively they provide structural data for HOS comparability and characterization.
With the help of the HOS 'map', consistency in protein structure is ensured on scale-up during lot-to-lot comparability, process development cycle, when production is altered, and when processes transfer to new manufacturing locations. Biosimilars can also be evaluated with the data obtained from HOS characterization.
During a manufacturing process (or change), testing and establishing comparability is not the same as the testing that was carried out during the development of biosimilar drug products. In essence, biosimilar development is a much more complicated process. Normally, a biosimilar is not produced by the same company that formulated the innovator (also known as reference or parental product).
Therefore, during the development of biosimilars, 'reverse engineering' has to establish new downstream and upstream processes and the biosimilar manufacturer is required to match the quality of product (biosimilarity) within the limited range of the innovator’s commercial product, demanding more biochemical and biophysical proof to show biosimilarity than what is usually required to show comparability during manufacturing.
HOS comparability of biopharmaceuticals –case studies using DSC
DSC provides data on a protein’s thermal stability under various solvent conditions, and DSC results obtained are reproducible (Figure 4). As a result, DSC is frequently considered in comparability studies to demonstrate that the process does not change the product stability, and that the products manufactured from different manufacturing sites and lots are very similar.
Jiang and Nahri
A number of biophysical methods for comparability during process development and manufacturing were reviewed by Jiang and Nahri[21]. In the experiment, DSC was used as one of the biophysical characterization tools in comparability of a monoclonal antibody drug product expressed by two different cell lines (cell lines 1 and 2), which also underwent a change in the manufacturing process (processes 2pA and 2pB).
Three protein products were obtained that were evaluated by FTIR (to find out the secondary structure), intrinsic fluorescence and near-UV CD (to compare the tertiary structure), DSC (to compare solubility and thermal stability), DLS (to compare the size distribution and hydrodynamic properties of proteins), and fluorescence and ANS binding (to compare surface hydrophobicity).
These biophysical assays provided results that indicated similarity in the overall secondary and tertiary structures of the three samples. The DSC results showed that there was increased thermal stability of the protein from cell line 2pB when compared to both the cell line 1 sample and the 2pA sample. According to DSC, the 2pA and cell line 1 samples also seemed to be more heterogeneous.
Usually, two or three transitions are observed for antibodies (refer Figure 2 for an example of a 'typical' antibody DSC thermogram). According to the DSC analysis of these samples, there were a higher number of overlapped thermal transitions, which indicated heterogeneity of the samples.
The DSC results indicated that the stability and homogeneity of the target monoclonal antibody were improved by the process change. Based on the functional, biochemical, and biophysical assay comparability data described in the article, process 2pB and cell line 2 were considered to yield the more stable protein[21].
In another second example from Jiang and Nahri[21], DSC was used to observe the comparability of a protein product developed at different manufacturing sites. Protein Y contained two monomeric polypeptide chains, disulfide-linked via the molecule’s Fc region.
During the development of Protein Y, it was manufactured at different sites. In order to confirm that the protein present in these different lots was similar in terms of native conformation, size distribution, thermal stability, and secondary and tertiary structures, four different Protein Y samples, representing reference the protein standard and three samples produced at different sites were examined by DSC, AUC, FTIR, fluorescence, near-UV CD, and far-UV CD.
The DSC scans of three samples from different manufacturing sites and the reference standard demonstrated two thermal transitions for individual sample, and within the experimental variability, all four DSC profiles were identical.
This showed that the protein samples did not have any different thermal stability and that all the samples were folded into the native conformation. The collective biophysical characterization results, including those provided by DSC, showed that all four samples of Protein Y samples were similar, uniform, and possessed the appropriate secondary and tertiary structure[21].
If the DSC thermograms had been non-identical, this may have indicated that chemical denaturation had transpired in the protein associated with the formulation or process, or a change in post-translational modification.
Arthur, et al.
Arthur, et al.[22] showed the sensitivity of DSC in detecting the HOS changes in stability caused by oxidation, which is a common chemical degradation pathway. When a biopharmaceutical product is exposed to oxidation during the purification process, or storage in the final formulation, it will not only affect efficacy but may also lead to the formation of protein aggregation.
In this analysis, protein products from three structural groups were examined at different oxidation levels. All proteins displayed a linear decrease in TM as a function of methionine oxidation. Variations in the rate of change in TM and Variations in domain TM stability, within and across structural classes[22] were also observed.
On the other hand, fluorescence spectroscopy and near-UV CD were much less sensitive to conformational changes induced by oxidation. Compared to these spectroscopic techniques, DSC was found to be a more suitable structural characterization technique for observing protein oxidation [22].
For a single protein (IgG2B) in the study, DSC detected the changes in TM preceding a loss in relative potency, indicating that DSC is a prime indicator of decreased antigen binding. Obvious changes in oxidized methionine by mass spectrometry (MS) took place at oxidation levels below those with a detectable functional or conformational effect.
Using TM shifts from DSC in tandem with MS and potency techniques, the association between changes in HOS and conformational stability, a primary structural modification, and functional impact can be consistently characterized.
Morar-Mitrica et al.
In yet another study, Morar-Mitrica et al.[12] discussed a number of case studies that incorporated DSC in HOS and comparability analyses of biopharmaceuticals, including extended DSC characterization of glycosylated monoclonal antibodies with varying levels of glycosylation (a standard post-translational modification), the characterization of monoclonal antibody oxidation by visible TM shifts (similar to those described in [22]), the use of ΔH, TM, Tonset, and T1/2 for comparability studies of biopharmaceuticals, and reproducible DSC thermogram profiles.
Shahrokl et al.
Shahrokl et al.[23] described the utility of DSC in comparability studies as part of the biological license application, citing DSC’s ease of use as well as the simplicity of its data output. The authors showed how DSC was employed for comparability of a 51 kDa glycoprotein before and after making a change in its production process.
The process change was mainly adopted to boost product yield and remove animal-derived materials arising in the cell culture medium. With the help of DSC, the authors assessed three lots of the glycoprotein that were developed by means of each manufacturing process, and they observed that the superimposable DSC thermograms demonstrated a transition at 60.7 °C +/- 0.1 °C.
These DSC results demonstrate that the change in the manufacturing process did not affect the glycoprotein’s conformational stability[23].
Lubiniecki et al.
Lubiniecki et al.[24] assessed the effect of manufacturing changes on the function and structure of antibodies throughout the product development. The authors conducted three comparability studies for two types of IgG1 monoclonal antibody candidates - antibody A and antibody B.
DSC was incorporated in two of the comparability studies as one of the biophysical and biochemical analytical tools. One study observed the process scale-up and transfer to the production site, along with the transition from a lyophilized to a liquid dosage form.
The DSC thermograms for the liquid and lyophilized formulations for antibody A and antibody B demonstrated good correlation[24]. Used in combination with SEC, CD, MS, and other assays, DSC demonstrated that the manufacturing change did not affect the antibody structure and function.
In the third comparability study, the liquid formulation of antibodies in prefilled vials or syringes was evaluated [24]. Results obtained from DSC, SEC, CD, and other assays indicated similar biological activity, molecular structure, and degradation profiles, except there was a small yet considerable increase in the levels of subvisible particles in prefilled vials or syringes[24].
Using DSC for characterization of biosimilars (biosimilarity)
A biosimilar product, also known as a “subsequent entry biologic” or “follow-on biologic,” is a biopharmaceutical product that has obtained a regulatory agency approval based upon its high similarity with an earlier-approved product, also called a reference product or an innovator or parental product.
Biosimilar products have been available worldwide for the several years [25-29]. During October 2016, 4 and 19 biosimilars have been approved by the US FDA and the EMEA, respectively.
Developing a biosimilar is a completely different process compared to the development of generic small molecule drugs. A small molecule drug is a chemical and its synthesis is a controlled process. On the other hand, biopharmaceutical products are normally proteins and therefore, a certain degree of protein variation can be observed in different lots of the same product because of the inherent variability of the manufacturing process and the biological expression system.
These variations include differences in post-translational modification and HOS. Manufacturing biosimilars is a complex process compared to the manufacture of generics due to the complex structures, high molecular weights, and natural variations of proteins. Therefore, a biosimilar product cannot be an exact replicate of the reference product, and can only be characterized as 'highly similar' or 'similar’ to the reference product.
The development of biosimilar products involves analytical, functional and physicochemical comparisons of the biosimilar and the reference protein. These assays are supported by comparative clinical and non-clinical data to establish equivalent safety and efficacy.
Before approval, it is essential to verify that there is no clinically meaningful variation in the biosimilar product in terms of effectiveness and safety compared to the reference product. Only slight variations in clinically-inactive components are acceptable in biosimilar products. For the same approved indications, the biosimilar product is expected to function like the reference product.
Regulatory agencies assess biosimilar products based on their degree of similarity with their reference products. Since biopharmaceuticals are complex in nature, it is not possible for two different manufacturers to synthesize two identical biopharmaceutical products even if they use processes, host expression systems, and equivalent technologies.
Therefore, biosimilar product manufacturers have to rely on the results of comparability studies and analysis of HOS as described above [30,31]. The desire for more analytical data, and the need for shorter timelines for biosimilar development requires the demonstration of comparability to the reference product at every stage, especially during production.
Biosimilar products need to be 'reverse-engineered' in order to achieve the high similarity with the reference product. The analytical characterization of a biosimilar product involves primary and higher order structure (secondary, tertiary, and quaternary) evaluation, biological activity, and investigation of process and product impurities.
DSC is widely used as a HOS biophysical assay to demonstrate that a biosimilar product has a highly similar DSC profile 'fingerprint', as well as similar thermodynamic parameters such as Tonset and TM in comparison with the reference molecule.
Sinha-Datta et al.[37] compared two therapeutic monoclonal antibodies (mAb1-i and mAb2-i) with their biosimilars (mAb1-B and mAb2-B1, B2, B3) using DSC (MicroCal VP-Capillary DSC) and surface plasmon resonance (SPR) as the analytical instruments for thermal stability, affinity and kinetics.
The SPR results revealed that the biosimilars showed high similarity to their reference samples in terms of biofunction. The researchers further confirmed the biosimilarity between the biosimilar and their parental products using DSC, which revealed good structural similarity between the biosimilar and parental antibodies, with the primary TM at 84.1°C for mAb1 and 72.8°C for mAb2.
Conclusion
Information discussed in this article shows the value and significance of using DSC as a biophysical stability and HOS assay for biopharmaceutical comparability and biosimilarity characterization. During production, biopharmaceutical manufacturers can take meaningful decisions on protein comparability and stability with DSC results and other biochemical and biophysical assays, ensuring the biosimilarity of each batch of protein to the reference lot.
They can also ensure that any manufacturing or process changes do not have an impact on the conformational protein stability. DSC is also incorporated as a HOS assay for the development of biosimilar products to show that the biosimilar product has high similarity to the innovator.
Suggested reading
- Analytical Techniques for Biopharmaceutical Development, R. Rodriguez-Diaz, T. Wehr, S. Tuck (eds.), Taylor & Francis, New York, NY, USA (2005).
- Biophysical Characterization of Proteins in Developing Biopharmaceuticals, D.J. Houde, S.A. Berkowitz (eds.), Elsevier, Amsterdam, Netherlands (2015).
- Biophysical Methods for Biotherapeutics: Discovery and Development Applications, T.K. Das (ed.) John Wiley & Sons, Hoboken, NJ, USA (2014).
- Biophysics for Therapeutic Protein Development, L.O. Nahri (ed.), Springer, New York, NY, USA (2013).
- State-of-the-Art and Emerging Technologies for Therapeutic Monoclonal Antibody Characterization Volume 1. Monoclonal Antibody Therapeutics: Structure, Function, and Regulatory Space, J.E. Schiel, D.D. Davis, O.V. Borisov (eds.), ACS Symposium Series Vol 1176 (2014). doi: 10.1021/bk-2014-1176.
- State-of-the-Art and Emerging Technologies for Therapeutic Monoclonal Antibody Characterization Volume 2. Biopharmaceutical Characterization: The NISTmAb Case Study, J.E. Schiel, D.D. Davis, O.V. Borisov (eds.), ACS Symposium Series Vol 1201 (2015). doi: 10.1021/bk-2015-1201.
- State-of-the-Art and Emerging Technologies for Therapeutic Monoclonal Antibody Characterization Volume 3. Defining the Next Generation of Analytical and Biophysical Techniques, J.E. Schiel, D.D. Davis, O.V. Borisov (eds.), ACS Symposium Series Vol 1202 (2015) DOI: 10.1021/bk-2015-1202.
References
- Gokarn, Y., Agarwal, S., Arthur, K., et al., Biophysical Techniques for Characterizing the Higher Order Structure and Interactions of Monoclonal Antibodies, in: State-of-the-Art and Emerging Technologies for Therapeutic Monoclonal Antibody Characterization Volume 2. Biopharmaceutical Characterization: The NISTmAb Case Study. J.E. Schiel, D.D. Davis, O.V. Borisov (eds.), ACS Symposium Series Vol 1201, American Chemical Society, Washington DC, USA, pages 285-327 (2015).
- Cooper, A., Nutley, M. A., and Wadood, A., Differential Scanning Calorimetry, in: Protein-Ligand Interactions: Hydrodynamics and Calorimetry, A Practical Approach, S.E. Harding, B.Z. Choudry (eds). Oxford University Press, Oxford, UK, p. 287-318 (2001).
- Malvern Panalytical Whitepaper “Differential Scanning Calorimetry (DSC) Theory and Practice” https://www.malvernpanalytical.com/en.
- Bruylants, G., Wouters, J., and Michaux, C., Current Med. Chem. 12, 2011-2020 (2005) doi: 10.2174/0929867054546564.
- Jelesarov, I., and Bosshard. H.R., J. Mol. Recognit. 12, 3-18 (1999) doi: 10.1002/(SICI)1099 1352(199901/02)12:1<3:AID-JMR441>3.0.CO; 2-6.
- Choi, M.H., and Prenner, E.J., J. Pharm. Bioallied Sci. 3, 39-59 (2011) doi:10.4103/0975-7406.76463.
- Johnson, C.M., Arch. Biochem. Biophys. 531, 100-109 (2013) doi: 10.1016/j.abb.2012.09.008.
- Plotnikov, V., Rochalski, A., Brandts, M., Brandts, J.F., Williston, S., Frasca, V., and Lin, L.N., Assay Drug Devel. Technol. 1, 83-90 (2004) doi:10.1089/154065802761001338.
- www.malvern.com
- Demarest, S.J., and Frasca. V., Differential Scanning Calorimetry in the Biopharmaceutical Sciences, in: Biophysical Characterization of Proteins in Developing Biopharmaceuticals, D.J. Houde, S.A. Berkowitz (eds.), Elsevier, Amsterdam, Netherlands, p. 287-306 (2015).
- Remmele, R.L., Microcalorimetric Approaches to Biopharmaceutical Development, in: Analytical Techniques for Biopharmaceutical Development, R. Rodriguez-Diaz, T. Wehr, S. Tuck (eds.), Taylor & Francis, New York, NY, USA, p. 327-381 (2005).
- Morar-Mitrica, S., Nesta, D., and Crofts, G., BioPharm Asia 2, 46-55 (2013).
- Kirkitadze, M., Hu, J., Tang, M., and Carpick, B., Pharm. Bioprocess. 2, 491-498 (2014) doi: 10.4155/PBP.14.27.
- Wen, J., Arthur, K., Chemmalil, L., Muzammil, S., Gabrielson, J., and Jiang, Y., J. Pharm. Sci. 101,955-964 (2012) doi: 10.1002/jps.22820.
- Gabrielson, J.P., and Weiss, W.F., J. Pharm. Sci. 104, 1240-1245 (2015) doi: 10.1002/jps24393.
- Cooney, B., Jones, S.D., and Levine, H., BioProcess Int. 14(6), 28-35 (2016). https://bioprocessintl.com/
- Cooney, B., Jones, S.D., and Levine, H., BioProcess Int. 14(8), 24-33 (2016). https://bioprocessintl.com/
- https://www.who.int/
- European Medicines Agency: Comparability of biotechnology-derived medicinal products after a change in the manufacturing process - non-clinical and clinical issues. EMEA/CHMP/ BMWP/101695/2006.https://www.ema.europa.eu/en/homepage
- http://www.fda.gov/downloads/Drugs/GuidanceComplianceRegulatoryInformation/Guidances/UCM496611.pdf
- Jiang, Y., and Nahri, L.O., J. Am. Pharm. Rev. 9, 34-43 (2006).
- Arthur, K.K., Dinh, N., and Gabrielson, J. P., J. Pharm. Sci. 104, 1544-1554 (2015) doi: 10.1002/jps.24313.
- Shahrokh, Z., Salamat-Miller, N., and Thomas, J.J., Biophysical Analyses Suitable for Chemistry, Manufacturing, and Control Sections of the Biologic License Application (BLA), in: Biophysical Methods for Biotherapeutics: Discovery and Development Applications, T.K. Das (ed.) John Wiley & Sons, Hoboken NJ USA pages 317-353 (2014).
- Lubiniecki, A., Volkin, D.B., Federici, M, et al., Biologicals 39, 9-22 (2011). doi: 10.1016/j.biologicals.2010.08.004.
- http://www.fda.gov/Drugs/DevelopmentApprovalProcess/HowDrugsareDevelopedandApproved/ApprovalApplications/TherapeuticBiologicApplications/Biosimilars/
- European Medicines Agency: Similar biological medicinal products containing biotechnologyderived proteins as active substance: non-clinical and clinical issues. EMEA/CHMP/BMWP/42832/2005 Rev. 1 .https://www.ema.europa.eu/en/homepage
- https://www.who.int/
- Bas, T.G., and Oliu Castillo, C., Biomed Res Int. 2016, 5910403 (2016). doi: 10.1155/2016/5910403.
- Tsuruta, L.R., Lopes dos Santos, M., and Moro, A.M., Biotechnol Prog. 31, 1139-1149 (2015). doi: 10.1002/btpr.2066.
- Kálmán-Szekeres, Z., Olajos, M., and Ganzler, K., J. Pharm. Biomed. Anal. 69, 185-195 (2012). doi: 10.1016/j.jpba.2012.04.037.
- Berkowitz, S.A., Engen, J.R., Mazzeo, J.R., and Jones, G.B., Nat. Rev. Drug Discov. 11, 527-540 (2012). doi: 10.1038/nrd3746.
- https://www.fda.gov/
- Jung, S.K., Lee, K. H., Jeon, J. W., et al., MAbs 6, 1163-1177 (2014). doi: 10.4161/mabs.32221.
- Liu, J., Eris, T., Li, C., Cao, S., and Kuhns, S., BioDrugs 30, 321-338 (2016). doi:10.1007/s40259-016-0184-3.
- Sinha-Datta, U., Khan, S., and Wadgaonkar, D., Biosimilars 5, 83-91 (2015) doi:10.2147/BS.S85537 https://www.dovepress.com/label-free-interaction-analysis-asa-tool-to-demonstrate-biosimilarity-peer-reviewed-article-BS
About Malvern Panalytical

Malvern Panalytical provides the materials and biophysical characterization technology and expertise that enable scientists and engineers to understand and control the properties of dispersed systems.
These systems range from proteins and polymers in solution, particle and nanoparticle suspensions and emulsions, through to sprays and aerosols, industrial bulk powders and high concentration slurries.
Used at all stages of research, development and manufacturing, Malvern Panalytical’s materials characterization instruments provide critical information that helps accelerate research and product development, enhance and maintain product quality and optimize process efficiency.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.Net which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.