Secondary lymphoid organs initiate adaptive immunity, and this is influenced by milieu generated by the innate immune system’s initial activation.
Longitudinal studies on humoral COVID-19 immunity and studies in convalescent subjects revealed that humoral immunity is generally short-lived, while the majority of SARS-CoV-2 antibodies demonstrate limited somatic hypermutation (Brouwer et al., 2020; Long et al., 2020; Robbiani et al., 2020).
Properly comprehending how the adaptive immune system is modulated in severe COVID-19 infections requires interrogation of secondary lymphoid organs during the acute phase of infection, where these responses are generated. However, many previous studies were primarily focused on peripheral blood samples.
SARS-CoV-2 infection leads to a wide range of clinical manifestations ranging from asymptomatic to rapidly fatal, but the causes of this heterogeneity are currently unknown. Seriously ill patients can suffer from a life-threatening acute respiratory distress syndrome, and even when cared for in an ICU or equivalent setting, some patients may still suffer from severe lung damage (Zhu et al., 2020; Zhou et al., 2020).
The virus tends to be located in airways and lungs during the early stages of infection, but this changes as the disease progresses (Schaefer et al., 2020). Damage-associated molecular patterns (DAMPs) are released by infected pneumocytes, and these are prone to combining with viral pathogen-associated molecular patterns (PAMPs), activating innate immunity (Vardhana and Wolchok, 2020).
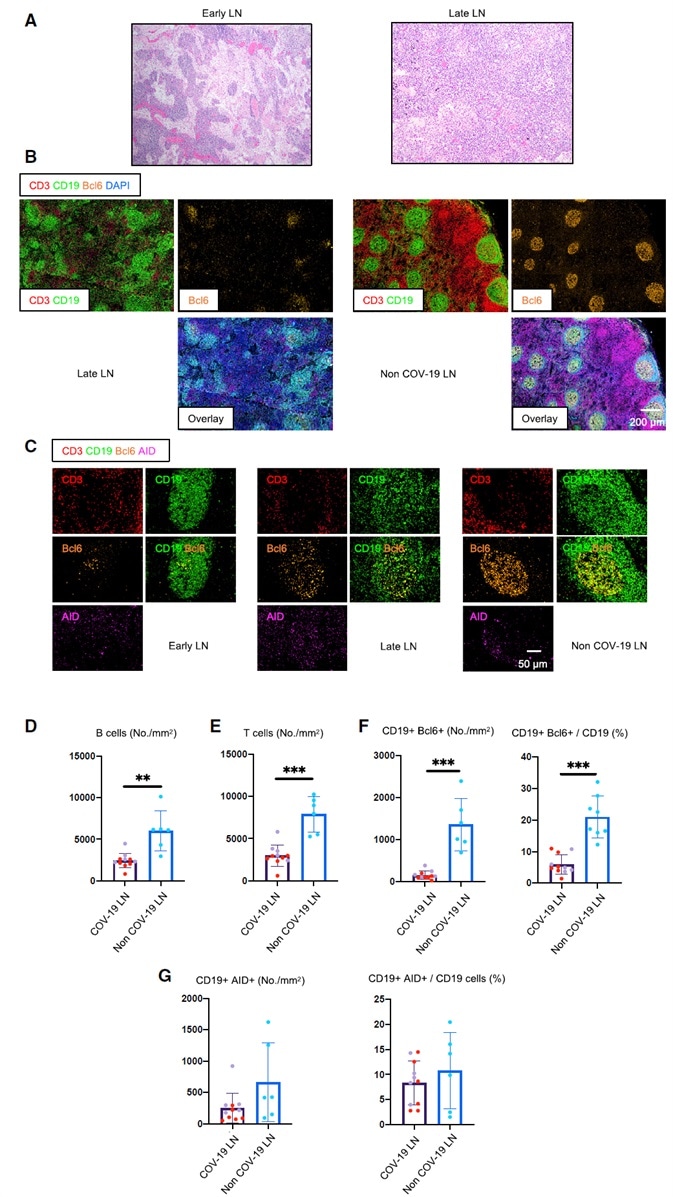
Figure 1. Early Loss of Germinal Centers and Bcl-6-Expressing B Cells in COVID-19 Thoracic Lymph Nodes.
(A) Hematoxylin-eosin staining of lymph nodes from early (left) and late (right) COVID-19 patients.
(B) Low-power images of CD3 (red), CD19 (green), Bcl-6 (orange), and DAPI (blue) staining in a lymph node from a late COVID-19 patient (left) and a non-COVID-19 thoracic lymph node (right).
(C) Representative multi-color immunofluorescence images of CD3 (red), CD19 (green), Bcl-6 (orange), and AID (purple) staining in lymph nodes from early (left) and late (middle) COVID-19 patients and a non-COVID-19 lymph node (right).
(D and E) Absolute numbers of CD19+ B cells (D) and CD3+ T cells (E) in lymph nodes from COVID-19 patients (purple, n = 11) and non-COVID-19 patients (blue, n = 6). COVID-19 samples include early (purple, n = 5) and late (red, n = 6) COVID-19 patients.
(F and G) Absolute numbers and relative proportion of Bcl6+ B cells (F) and AID+ B cells (G) in the pool of CD19+ B cells in lymph nodes from COVID-19 patients (purple, n = 11) and non-COVID-19 patients (blue, n = 6).
COV-19, COVID-19; LN, lymph node. Mann-Whitney U test was used to calculate p value. Error bars represent mean ± SEM. **p < 0.01; ***p < 0.001.
Image Credit: TissueGnostics
Cytokine milieu generated throughout this process would generally influence the induction of lymphocyte activation by antigen conveyed directly in the lymph, or via dendritic cells to draining lymph nodes. It is also likely that viremia results in the initiation of immune responses in the spleen.
There are many similarities between the features of severe COVID-19 and severe acute respiratory syndrome (SARS). SARS-CoV-2 infection (Guan et al., 2020) has been found to include progressive lymphopenia, and the degree of lymphopenia has been correlated with increases in circulating interleukin-6 (IL-6) and IL-8 (Zhang et al., 2020).
Lymphopenia has also been observed in SARS, at the peak of active disease. This was characterized by acute respiratory distress and cytokine storm (Perlman and Dandekar, 2005). Autopsy studies of SARS patients have revealed lymphoid organ atrophy, including spleen, lymph nodes, Peyer’s patches, and the loss of germinal centers (Gu et al., 2005).
Autopsy studies in COVID-19 patients have also found lymphocyte depletion in spleen and lymph nodes (Lax et al., 2020), as well as splenic white pulp atrophy (Xu et al., 2020; Buja et al., 2020).
Many viral and non-viral infections lead to cytokine storm, as well as lymphopenia and acute respiratory distress (Tisoncik et al., 2012). Furthermore, splenic white pulp atrophy has previously been histopathologically demonstrated in Marburg disease, Ebola (Martines et al., 2015; Rippey et. al., 1984) and H5N1 influenza (Gao et al., 2010; Lu et al., 2008).
When considered together, these factors suggest that numerous different viral and infectious triggers may contribute to a comparable constellation of immunological phenomena, potentially driving pathology.
In individuals with COVID-19, the durability and magnitude of antibody responses are increased in those with more severe disease (Ju et al., 2020; Amanat et al., 2020). However, these are often of low magnitude (Robbiani et al., 2020) and seem to lack durability (Long et al., 2020). There is a possibility that this is similar to Middle East respiratory syndrome (MERS) and SARS, where humoral responses were generally found to be not durable, except in a particular subset of individuals (Mo et al., 2006; Zumla et al., 2015).
There have been documented cases of impaired infection-induced protective immunity where patients with less severe respiratory tract infections (Galanti et al., 2019) suffered from repeated infections with the human coronaviruses CoV 229E, OC43, NL63, and HKU1.
It is possible that reinfection is a result of viral strain subtypes, but the cause of the widespread lack of durable humoral immune responses to coronaviruses remains unknown. Better understanding of changes in humoral immune system components - particularly in secondary lymphoid organs - offers an opportunity to learn why natural infections with coronaviruses tend not to lead to durable immunity.
A granular analysis of B and T lymphocytes in draining lymph nodes and spleens of SARS or MERS patients was never reported, leaving the underlying basis for the lymphopenia and the general lack of durability of antibody responses in those diseases unresolved.
COVID-19 disease primarily affects the lungs, so the Massachusetts Consortium on Pathogen Readiness Specimen Working Group has carried out an investigation into thoracic lymph nodes, analyzing lymphoid architecture and lymphocyte populations by utilizing multispectral imaging, multi-color immunofluorescence, and cell-cell interaction analyses from the initial onset of the disease in individuals with a range of disease outcomes.
Because viremia has been noted in COVID-19 (Zheng et al., 2020; Lescure et al., 2020), the group also interrogated spleens the acute- and late-disease settings, complementing these studies with an examination of peripheral blood samples in a different cohort, allowing convalescence to be studied as well.
The group’s results identified a notable lack of lymph node and splenic germinal centers and Bcl-6-expressing B cells. The group also detected defective Bcl-6+ T follicular helper cell generation and differentiation, as well as dysregulated SARS-CoV-2-specific humoral immunity in the early stages of COVID-19 disease.
These results have helped develop a mechanistic explanation for the limited durability of humoral immunity, as well as helping explain the less robust somatic hypermutation observed in this disease in the wake of natural infection.
Results
A human-tissue-imaging platform was used with quantitative high-resolution automated slide-scanning microscopy, employing both confocal and regular approaches along with multispectral imaging. The platform was used to interrogate human lymphoid and non-lymphoid organs at the single-cell level, with these particular approaches being essential in preserving architecture over broad swaths of tissue.
In severely ill COVID-19 patients who succumbed in under 8 days following admission (this group is designated ‘‘early’’ - less than 10 days from the start of respiratory symptoms, Table S1), thoracic lymph nodes were found to display a lack of germinal centers. These were also lacking in patients who succumbed later (this group is designated as “late” - 15–36 days after being admitted, Figures 1A and 1B). Controls took the form of thoracic lymph nodes from age-matched individuals who had succumbed from non-COVID-19 causes (Table S2).
Quantitation showed significant early loss of both B and T cells. Absolute numbers declined to approximately one-third of their non-COVID-19 controls, persisting in late disease (Figures 1D and 1E). Distinct B and T cell zones could always be clearly observed, however.
It is possible that human control lymph nodes include germinal centers due to ongoing adaptive immunity that has been initiated by commensal antigens. One unexpected occurrence in this study was the absence of germinal centers in thoracic lymph nodes of acutely ill COVID-19 patients, whose lungs were already known to contain very high viral loads (Schaefer et al., 2020).
Bcl-6-expressing germinal center B cells were also found to be reduced in COVID-19, but there was found to a preservation of AID-ex- pressing B cells. These were diffusely distributed when compared to controls, however.
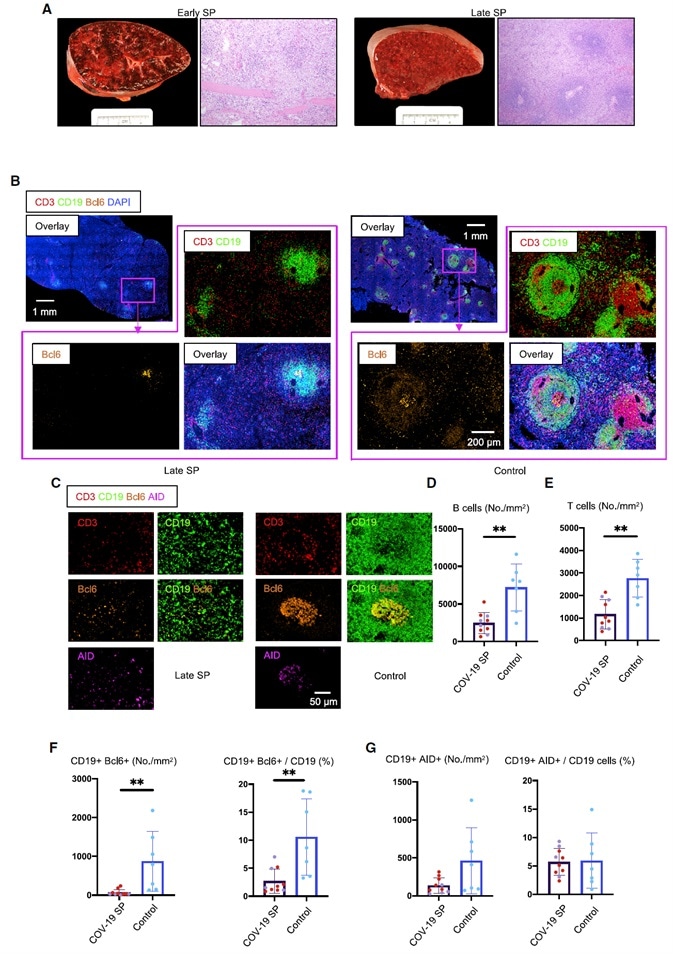
Figure 2. White Pulp Attrition, Early Loss of Germinal Centers, and Bcl-6-Expressing B Cells in COVID-19 Spleens.
(A) Cross-sectional view of whole spleen and hematoxylin-eosin staining from early (left) and late (right) COVID-19 patients.
(B) Low-power images of CD3 (red), CD19 (green), Bcl-6 (orange), and DAPI (blue) staining in a spleen from a late COVID-19 patient (left) and a control (right).
(C) Representative multi-color immunofluorescence image of CD3 (red), CD19 (green), Bcl-6 (orange), and AID (purple) staining in spleens from a late COVID-19 patient (left) and a control (right).
(D and E) Absolute numbers of CD19+ B cells (D) and CD3+ T cells (E) in spleens from early (purple, n = 4) and late (red, n = 6) COVID-19 patients and controls (blue, n = 7).
(F and G) Absolute numbers and relative proportion of Bcl-6+ B cells (F) and AID+ B cells (G) in spleens from early (purple, n = 4) and late (red, n = 6) COVID-19 patients and controls (blue, n = 7).
SP, spleen. Multiple comparisons are controlled for by Kruskal-Wallis test. Error bars represent mean ± SEM. *p < 0.05.
Image Credit: TissueGnostics
Click here to read the full paper!
Acknowledgments
Produced from materials originally authored by Naoki Kaneko, Hsiao-Hsuan Kuo, Julie Boucau, Robert F. Padera, Jr. and Shiv Pillai from the Massachusetts Consortium on Pathogen Readiness Specimen Working Group.
About TissueGnostics
TissueGnostics (TG) is an Austrian company focusing on integrated solutions for high content and/or high throughput scanning and analysis of biomedical, veterinary, natural sciences, and technical microscopy samples.
TG has been founded by scientists from the Vienna University Hospital (AKH) in 2003. It is now a globally active company with subsidiaries in the EU, the USA, and China, and customers in 30 countries.
TissueGnostics portfolio
TG scanning systems are currently based on versatile automated microscopy systems with or without image analysis capabilities. We strive to provide cutting-edge technology solutions, such as multispectral imaging and context-based image analysis as well as established features like Z-Stacking and Extended Focus. This is combined with a strong emphasis on automation, ease of use of all solutions, and the production of publication-ready data.
The TG systems offer integrated workflows, i.e. scan and analysis, for digital slides or images of tissue sections, Tissue Microarrays (TMA), cell culture monolayers, smears, and other samples on slides and oversized slides, in Microtiter plates, Petri dishes and specialized sample containers. TG also provides dedicated workflows for FISH, CISH, and other dot structures.
TG analysis software apart from being integrated into full systems is fully standalone capable and supports a wide variety of scanner image formats as well as digital images taken with any microscope.
TG also provides routine hematology scanning and analysis systems for peripheral blood, bone marrow, and body fluids.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.Net which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.