In addition to the identification and characterization of individual cell morphology, a more complex phenotypic profile will include the cell’s activation state and role within the tissue microenvironment.
Image credits: Komsan Loonprom | Shutterstock
Contemporary advancements in quantitative imaging software and hardware have enabled researchers to examine immune cells in situ. However, as only four biomarkers can be examined at any one time using conventional microscopy systems, this does not provide a comprehensive representation of the cell’s activity.
Likewise, although flow cytometry is a well-established methodology which permits multiple biomarkers to be examined at any one time, it also disassociates the cells from key information regarding the tissue microenvironment.
The establishment of a system that can provide flow cytometry-like data on cell biomarkers, while preserving information on the tissue microenvironment, as in conventional microscopy, would thus be exceptionally advantageous in terms of satisfying the rigors of contemporary clinical practice.
A recent study by researchers at Yale University School of Medicine (Blenman K & Bosenberg M, 2019) presents a methodology that can overcome the limitations above. They demonstrate that the application of novel analytic technology to a standard microscopy configuration enables the complex characterization of immune cells in situ.
Adapting a Familiar Setup
Dr Kim Blenman examined tissue samples from the spleen in a mouse model of melanoma developed by Dr Marcus Bosenberg. Dr Blenman utilized a standard microscopy setup for performing multiplexed fluorochrome-based histology on formalin-fixed paraffin-embedded tissue sections, with biomarkers conjugated to Cy3 or Cy5.
Dr Blenman focused on the CD4 transmembrane protein – a biomarker of a number of T-cell subtypes – and six biomarkers of cell activity, including proliferation and transcription-associated with these cells.
She utilized multiple staining rounds, inactivating the dyes between every round, to examine the divergent biomarkers of cell activity, such as Foxp3, Granzyme B, IL-6, Ki-67, RORγ(t) and T-bet.
Dr Blenman utilized an all-in-one system from TissueGnostics. High-magnification images were obtained using the TissueFAXS Quantitative Imaging System for downstream analysis. This was integrated with sophisticated image processing software (TissueGnostics StrataQuest) which can reconstruct whole images in silico and perform cell phenotyping and tissue cytometry.
The software first utilizes an algorithm to stitch the tiled images together, thus recreating the whole image. It subsequently generates a composite image that comprises every biomarker from the staining rounds. Cell isolation is then undertaken utilizing two algorithms: one for identifying the cell nucleus and a second for identifying the biomarker for the cell phenotype.
Quantification was subsequently accomplished by establishing which cells were positive for each biomarker. This was undertaken using a backgating algorithm for determining cutoffs at which cells or cell clusters should or should not be included.
Utilizing this methodology on the spleen tissue samples, Dr Blenman could demonstrate that it is capable of obtaining multiple biomarkers at one time in situ.
This information enabled the phenotypical characterization of cells within the tissue sample and, in turn, they could assign those cells to categories based on the combination of biomarkers expressed by individual and clusters of cells. In doing so, they demonstrated that levels of CD4 expression within cell clusters were correlative with explicit patterns of biomarker expression and cell proliferation.
Visualizing the Future
The immune system is exceptionally complex and the restriction to profiling only four biomarkers per cell type represents a significant limitation. One benefit of the system utilized by the authors is that it utilizes a microscopy configuration that already exists in the majority of laboratories, making it extremely accessible. Moreover, while the authors elected to examine six biomarkers, there are, in theory, no limits to the number of biomarkers that can be examined at one time.
Dr Blenman states that this methodology has the advantage of being able to quantify cell biomarkers objectively. Writing in Cytometry Part A, she asserts it could be extremely efficacious in cancer treatment, where there is an increase in targeting therapies to an individual’s disease and its underlying mechanism. For instance, the technique might be utilized for detecting response to immunotherapy via the analysis of biopsies before, during, and after treatment.
The methodology, facilitated by flow cytometry-like capabilities such as gating, backgating and histogram/dot scatterplot outputs, allows the operator to retain spatial information about the cells during phenotypical characterization.
TissueGnostics Innovation
The process elaborated by Dr Blenman in the Cytometry A paper was facilitated by the TissueFAXS Quantitative Imaging System and TissueGnostics StrataQuest analysis software. When integrated, these technologies generate an efficient, highly automated process, thus freeing the operator from hands-on time. The authors emphasize the potential of TissueFAXs for incorporating MATLAB scripts into the StrataQuest analysis software. There are also more than 50 compatible ready-to-load apps which can enable specialized analyses. Each app provides a specific analysis from start to finish, delivering data that is ready to export to the operator’s chosen program.
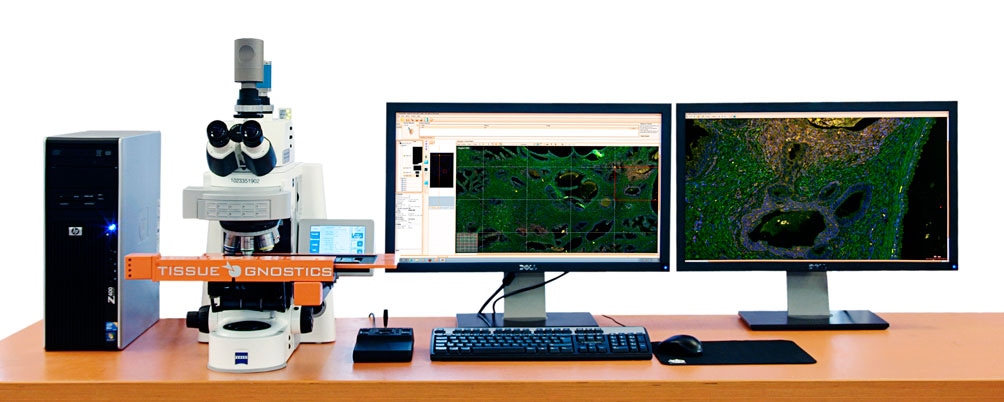

Image credit: TissueGnostics
The TissueFAXS system is a versatile upright system which is capable of scanning and analyzing cytospins, slides, smears and tissue microarrays. Although it is already fitted with both brightfield and fluorescence, with supplemental components, it is also customizable for contrast microscopy methods.
The system can be configured with one of three software options, of which StrataQuest is the most comprehensive and advanced. StrataQuest can automatically identify structures within a digital slide, such as blood vessels, and can detect and quantify millions of cells in a cell sample. The software is also available as a standalone product for utilization with existing scanning systems and is compatible with imported images and slides in a broad array of formats.
References
About TissueGnostics
TissueGnostics (TG) is an Austrian company focusing on integrated solutions for high content and/or high throughput scanning and analysis of biomedical, veterinary, natural sciences and technical microscopy samples.
TG was founded by scientists from the Vienna University Hospital (AKH) in 2003. It is now a globally active company with subsidiaries in the EU, the USA and China and customers in 28 countries.
TG systems offer integrated workflows, i.e. scan and analysis, for digital slides or images of tissue sections, Tissue Microarrays (TMA), cell culture monolayers, smears and of other samples on slides and oversized slides, in Microtiter plates, Petri dishes and specialized sample containers.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.Net which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.