One of the most challenging aspects of imaging mammalian tissues is how to handle their diverse composition. The human body is comprised of hundreds of known cell types, with immeasurably more to be discovered.
There is a significant need for finding imaging techniques with the capacity to recover imaging information on cell-to-cell interactions with good resolution to identify unique phenotypes while demonstrating compatibility with the diversity of cell types..
IBEX protocol
The recently established Iterative Bleaching Extends multiplexity (IBEX) imaging method provides a dynamic approach to the multiplex imaging of various tissue types, including human organs.1,2
IBEX is a comparatively inexpensive protocol that biologists with basic laboratory skills can complete in just 2-5 days. The method has also been designed to recover ultrahigh content information for qualitative and quantitative analysis.
Central to the IBEX protocol is a multiplex antibody-based imaging technique for the spatial characterization of complex phenotypes in tissue by available reagents and equipment. The technique relies on chemical bleaching of fluorophores conjugated to the antibodies without physically removing them. The process involves staining, imaging and bleaching steps repeated in cycles.
While it is possible to manually process IBEX images, the method can also be utilized with automated content imaging and sample processing.
One of the main benefits of the IBEX protocol is how dynamic and flexible the method is. It demonstrates compatibility with a number of different imaging platforms as well as 250 commercially available antibodies and 16 fluorophores.
Protocols that advocate for antibody-fluorophore combinations and tissue preparation steps are also available to stimulate widespread method adoption and minimize the amount of characterization and preparatory work that individual laboratories need to conduct.
Faster immunolabelling and bleaching times are another key advantage of IBEX imaging. The capacity to stain and image numerous biomarkers in a single tissue sample is key to exploring the complete complexity of most tissue samples.
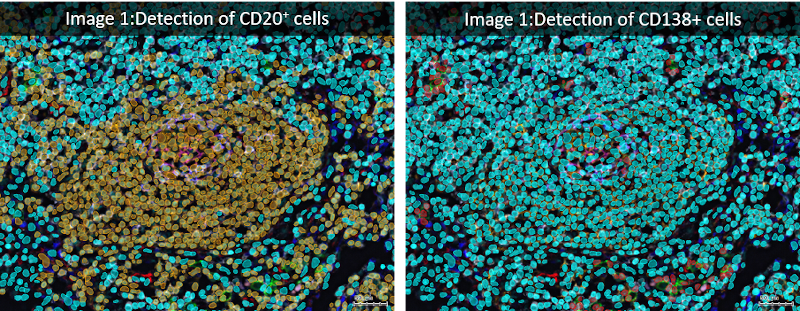
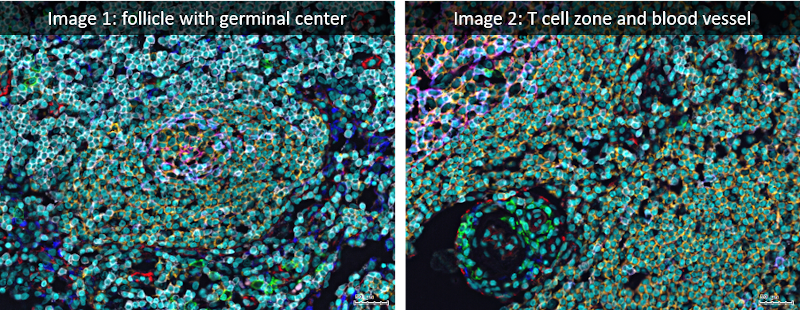
Analyzed lymph node images were acquired using the iterative bleaching extends multiplexity (IBEX) method (PNAS, Nature Protocols), an open source multiplexed antibody-based imaging method (Nature Methods). StrataQuest analysis was performed on a lymph node dataset shared publicly by the authors (Accompanying Zenodo).
PNAS: https://www.pnas.org/doi/10.1073/pnas.2018488117
Nature Protocols: https://www.nature.com/articles/s41596-021-00644-9
Nature Methods: https://www.nature.com/articles/s41592-021-01316-y
Accompanying Zenodo: https://zenodo.org/record/5244551#.YnVAZ-jMIuU
StrataQuest
One way to boost throughput potential, efficiency and detail analysis of the images from IBEX protocols is to take advantage of automated image analysis.
For brightfield or fluorescence image processing, TissueGnostics offers StrataQuest, a world-leading context-based analysis software.3 StrataQuest can be employed in different scenarios, such as the reconstruction of Z stack or entire slide images.
As far as compatibility with IBEX imaging datasets goes, StrataQuest can effortlessly complete all the appropriate processing steps due to the quality of the StrataQuest app design.
StrataQuest was developed with the objective of simplifying and accelerating workflows. To get started with StrataQuest to analyze the output of an IBEX protocol, the workflow may be initiated with image alignment. IBEX procedures can replicate either 3D image stacks or 2D single-z slices.
The alignment step is typically conducted using the intensity of the images or in multi-cycle IBEX experiments by evaluating the image to locate the presence of a repeated marker to fix all images to a fixed position. Establishing a marker means identifying numerous features – either a specific cell organelle or structural feature.
Normalization of the images is performed using a similarity metric on the aligned images, providing a robust registration of a large volume of data in three dimensions of data. The protocol and its application, in combination with StrataQuest, can process more than 160 GB of data with over 20 imaging cycles in a matter of minutes.
Simple interfacing
Once the processing of images is complete, these can be evaluated for the analysis of cell populations and relative positions. Within StrataQuest, it is possible to build workflows that integrate the required image processing for handling the output of IBEX experiments and can be used to automate subsequent data analysis workflows.
The ‘SQ Auto Analysis’ can export any results in the required format. StrataQuest 7.1 has several image analysis and recognition capabilities built-in, including a machine learning-based tissue classifier that can output information such as proximity and distance information and can be utilized for automated contextual tissue cytometry.
StrataQuest’s built-in feature relative to cell, tissue and structure recognition as well as proximity works well with IBEX processed samples.
The range of tissue types that can be given distinct stains in a single image (with up to 29 parameter datasets made possible with automated IBEX) means the automated analysis that StrataQuest performs can help expedite workflows and make sure all possible interactions are captured.
IBEX procedures are especially powerful when visualizing tumor-immune interactions that are vital when trying to better understand the underlying biochemical mechanisms involved in tumor growth and developing new therapies to target specific types of metastatic cells.
Performing a significant number of image cycles with IBEX does not cause cellular degradation, making it possible to utilize great numbers of antibodies when a number of distinct cell types are present in complex human tissue samples.
TissueGnostics
StrataQuest can either be a self-contained analysis program or amalgamated with imaging equipment to conduct automated experiment control. This may involve processes such as auto sampling to determine regions of interest for further imaging utilizing the SQ app to pinpoint ‘meaningful objects.’
This makes choosing StrataQuest an easy choice when selecting software to support either automated or manual IBEX workflows to acquire the maximum amount of information from IBEX images in the fastest timeframes.
References
- Radtke, A. J., Chu, C. J., Yaniv, Z., Yao, L., Marr, J., Beuschel, R. T., Ichise, H., Gola, A., Kabat, J., Lowekamp, B., Speranza, E., Croteau, J., Thakur, N., Jonigk, D., Davis, J., Hernandez, J. M., & Germain, R. N. (2021). IBEX: An open and extensible method for high content multiplex imaging of diverse tissues.
- Radtke, A. J., Kandov, E., Lowekamp, B., Speranza, E., Chu, C. J., Gola, A., Thakur, N., Shih, R., Yao, L., Yaniv, Z. R., Beuschel, R. T., Kabat, J., Croteau, J., Davis, J., Hernandez, J. M., & Germain, R. N. (2020). IBEX: A versatile multiplex optical imaging approach for deep phenotyping and spatial analysis of cells in complex tissues. Proceedings of the National Academy of Sciences of the United States of America, 117(52), 33455–33465. https://doi.org/10.1073/PNAS.2018488117
- TissueGnostics (2022) StrataQuest, https://tissuegnostics.com/products/contextual-image-analysis/strataquest, accessed March 2022

About TissueGnostics
TissueGnostics (TG) is an Austrian company focusing on integrated solutions for high content and/or high throughput scanning and analysis of biomedical, veterinary, natural sciences, and technical microscopy samples.
TG has been founded by scientists from the Vienna University Hospital (AKH) in 2003. It is now a globally active company with subsidiaries in the EU, the USA, and China, and customers in 30 countries.
TissueGnostics portfolio
TG scanning systems are currently based on versatile automated microscopy systems with or without image analysis capabilities. We strive to provide cutting-edge technology solutions, such as multispectral imaging and context-based image analysis as well as established features like Z-Stacking and Extended Focus. This is combined with a strong emphasis on automation, ease of use of all solutions, and the production of publication-ready data.
The TG systems offer integrated workflows, i.e. scan and analysis, for digital slides or images of tissue sections, Tissue Microarrays (TMA), cell culture monolayers, smears, and other samples on slides and oversized slides, in Microtiter plates, Petri dishes and specialized sample containers. TG also provides dedicated workflows for FISH, CISH, and other dot structures.
TG analysis software apart from being integrated into full systems is fully standalone capable and supports a wide variety of scanner image formats as well as digital images taken with any microscope.
TG also provides routine hematology scanning and analysis systems for peripheral blood, bone marrow, and body fluids.
TG cooperations
TG continuously cooperates with research groups and other companies in the industry to provide novel tools and applications to its customers.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.Net which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.