Linking Protein Dynamics To Systems-Level Biological Understanding
Role of Proteomics in Systems Biology
Modeling Cellular Pathways Using Proteomics Data
Integration of Proteomics With Multi-Omics Data
Applications in Disease Research
Challenges in Proteomics-Based Systems Biology
Future Directions
Proteomics Enables Dynamic Modeling Of Cellular Systems
References
Further Reading
Proteomics enables systems biology by providing dynamic, protein-level insights into cellular networks, facilitating the modeling of complex biological pathways and disease mechanisms.
 Image credit: bixstock/Shutterstock.com
Image credit: bixstock/Shutterstock.com
Linking Protein Dynamics To Systems-Level Biological Understanding
Systems biology is an integrative approach that examines biological systems as interconnected networks of genes, proteins, and metabolites, aiming to understand their dynamic behavior rather than isolated components. On the other hand, reductionist biology examines organic molecules separately from their context. Systems biology studies how biological systems work as interconnected networks to create new properties. These systems exhibit emergent behavior, in which system-level functions arise from nonlinear interactions, feedback loops, and stochastic processes that cannot be predicted from studying individual components alone.2
In addition, proteomics provides information about large-scale protein interactions by analyzing how proteins relate to the global network of interconnected biological processes. As proteins are the primary executors of cellular processes and reflect real-time biological activity, proteomics enables modeling of dynamic pathways and disease-associated molecular changes across biological conditions.3,5
This article explains how proteomics drives systems biology by modeling dynamic pathways and improving disease understanding and precision medicine.
Save this guide to proteomics-driven systems biology and dynamic disease pathway modeling. Download the full PDF here.
Role of Proteomics in Systems Biology
Systems biology focuses on understanding biological systems as interconnected networks, in which interactions, feedback loops, and emergent properties determine overall function rather than the function of individual components alone. Unlike looking at single genes, studying the whole system is better for understanding complex, changing behaviors. Within this framework, proteomics plays a central role by enabling the large-scale analysis of proteins, which are key functional molecules in cellular processes.1,2
Proteomics facilitates the mapping of protein-protein interactions (PPIs), identification of signaling pathways, and reconstruction of complex regulatory networks. It provides insights into how proteins coordinate cellular responses through interconnected pathways. Importantly, proteome-level measurements reflect post-transcriptional regulation and capture pivotal post-translational modifications, such as phosphorylation, that govern protein function and signaling.
Post-translational modifications (PTMs) are widespread across the proteome and can alter protein structure, localization, stability, and interaction dynamics, thereby significantly expanding functional diversity beyond the genome. Since proteins mediate cellular functions, proteomics provides a dynamic and functional view of biology that is not possible from genomic data alone.1,3,5
Proteomics 101
Video credit: JAMANetwork/Youtube.com
Modeling Cellular Pathways Using Proteomics Data
Proteomics offers a powerful framework for modeling cellular pathways, capturing changes in protein expression, modification, and interactions across biological conditions. Quantitative proteomics measures protein levels to detect changes associated with pathway activation or inhibition. Such information provides a direct insight into cellular responses to physiological or pathological stimuli at the protein level.2,5
PTMs further refine pathway modeling by regulating protein activity, localization, and stability. Modifications such as phosphorylation play essential roles in controlling signaling cascades and enabling rapid, reversible cellular responses, making them critical for understanding signal transduction networks. For example, phosphorylation can modulate enzyme activity, promote or disrupt protein–protein interactions, and trigger conformational changes that propagate signaling events across networks.1,3
In addition, advances in proteomics technologies allow the study of dynamic and spatial aspects of cellular processes. High-throughput mass spectrometry enables the monitoring of protein changes over time and across different cellular compartments, supporting a more context-specific understanding of pathway behavior. Modern workflows, including bottom-up and top-down proteomics, enable both peptide-level quantification and intact-protein characterization, thereby improving the resolution of proteoforms and complex assemblies.2,5
These datasets are integrated into network-based models using computational and graph-theoretical methods, in which proteins are represented as interconnected nodes. Such models facilitate the reconstruction of signaling networks and enable simulations that predict system behavior under varying biological conditions. Quantitative modeling approaches, including differential equation-based and constraint-based models, are increasingly used to simulate dynamic system responses and predict perturbation effects.1,2,4
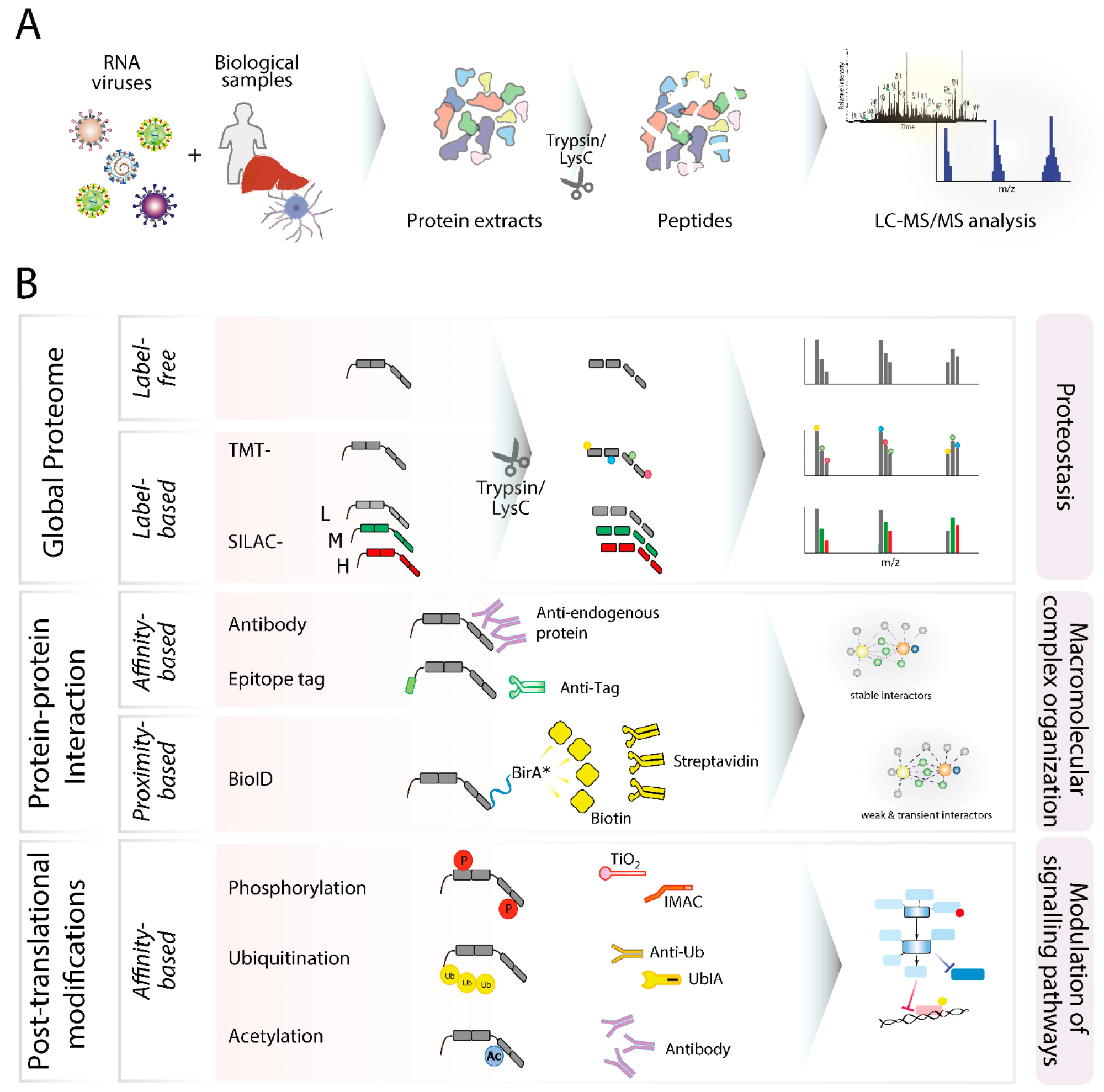 Overview of mass spectrometry–based proteomics workflows for studying RNA virus biology. (A) General proteomics pipeline illustrating sample collection, protein extraction, enzymatic digestion (e.g., trypsin/LysC), and LC-MS/MS analysis. (B) Comparative strategies for proteome analysis, including label-free and label-based quantification, protein–protein interaction mapping (affinity- and proximity-based methods), and post-translational modification profiling, highlighting their roles in proteostasis, macromolecular complex organization, and signaling pathway modulation.5
Overview of mass spectrometry–based proteomics workflows for studying RNA virus biology. (A) General proteomics pipeline illustrating sample collection, protein extraction, enzymatic digestion (e.g., trypsin/LysC), and LC-MS/MS analysis. (B) Comparative strategies for proteome analysis, including label-free and label-based quantification, protein–protein interaction mapping (affinity- and proximity-based methods), and post-translational modification profiling, highlighting their roles in proteostasis, macromolecular complex organization, and signaling pathway modulation.5
Integration of Proteomics With Multi-Omics Data
Proteomic integration with other omics levels is crucial to systems biology, as it offers a more complete view of biological systems. The different omics methods provide complementary views of the biology of interest; genomics provides the genetic blueprint, transcriptomics documents gene activity, proteomics records protein activity, and metabolomics operates downstream of all previous levels to demonstrate biochemical activity. Studying these layers together allows researchers to better understand how molecular interactions translate into cellular function and disease phenotypes.4
Multi-omics integration strategies involve combining datasets through approaches such as data layering, correlation analysis, and network-based modeling. Machine learning is now commonly used to extract patterns and predict outcomes from complex data. These approaches are often supported by systems-level databases and bioinformatics platforms that facilitate data sharing, annotation, and interpretation across biological domains. Integration strategies may be simultaneous (analyzing all omics layers together) or step-wise (integrating results sequentially), depending on data availability and study design.4,5
Despite advances, integrating multi-omics data remains challenging, as variability across testing platforms (batch effects) introduces bias and complicates comparisons. Differences in data scale, resolution, and measurement techniques further complicate integration, while the high dimensionality and heterogeneity of datasets make harmonization and interpretation difficult. The lack of standardized pipelines and incomplete overlap of samples across omics layers further limit the robustness of integrative analyses.4,5
Applications in Disease Research
Proteomics has emerged as a prototype for investigating disease mechanisms, enabling large-scale observations of protein expression and function across diverse biological states. By comparing proteomic profiles between healthy and diseased conditions, researchers can identify biological pathways in cancer or Alzheimer’s. These insights offer a systems-level perspective on how molecular alterations drive disease onset and development. Such analyses can reveal dysregulated signaling networks and identify key regulatory nodes involved in disease pathogenesis.2,5
Proteomics is also very important in biomarker discovery. The identification of disease-specific protein signatures helps in early diagnosis and more accurate prognosis. Protein biomarkers detected in tissues or biofluids can reflect underlying pathological changes, making them valuable tools for clinical assessment and monitoring.5
By identifying key regulatory proteins and interaction hubs within disease networks, proteomics helps identify potential therapeutic targets. This information can be beneficial for developing new drugs or repurposing existing drugs. By combining proteomic findings with clinical data, precision medicine initiatives will be enhanced through patient stratification, enabling the development of targeted, individualized treatment regimens. Proteomic profiling can also be used to monitor treatment response and identify mechanisms of drug resistance, improving clinical decision-making. These approaches support the transition toward personalized medicine by linking proteomic profiles to patient-specific therapeutic strategies.3,5
 Image credit: angellodeco/Shutterstock.com
Image credit: angellodeco/Shutterstock.com
Challenges in Proteomics-Based Systems Biology
Proteomics-based systems biology faces several challenges related to the complexity and scale of the data generated. High-throughput technologies produce large, high-dimensional datasets that require advanced tools for analysis, yet understanding this multi-layered biological information remains difficult. The integration of diverse omics layers further increases analytical complexity and demands robust data processing strategies.1,4
Reproducibility and standardization in experimentation are major issues in proteomics and systems biology, and variability in experimental methods (e.g., protocols for sample preparation, handling, and the use of different instruments) may lead to inconsistencies, underscoring the need for standardized workflows and robust data validation procedures.2,4
The use of computational models in systems biology may be overly simplistic and not able to represent the nonlinearities, feedback cycles, and stochasticity found in many cellular processes. Additionally, incomplete proteome coverage, particularly for low-abundance proteins, and technical limitations in detecting transient protein interactions remain significant barriers.3,5
Finally, challenges in data integration and scalability persist, as combining heterogeneous datasets across platforms and scaling analyses to larger populations remain difficult due to limited interoperability and data harmonization issues.4
Future Directions
Due to advancements in proteomics technology, the ability to conduct systems biology research will significantly increase through enhanced sensitivity, resolution, and analytical depth. The advent of quantitative and single-cell proteomics will also allow researchers to detect many proteins at lower levels of abundance than previously possible. This will enable a more robust assessment of cell-to-cell variability and a more accurate depiction of heterogeneous biological systems. These advancements now enable adequate capture of the dynamic processes that occur in a cell.2,5
At the same time, artificial intelligence (AI) and machine learning are becoming key tools for analyzing complex proteomics and multi-omics datasets. These techniques enhance pattern analysis, develop predictive models, and enable the integration of diverse input data to enhance biological interpretation.4,5
Systems biology is also advancing as computational frameworks improve. Multi-scale approaches focus on improving the modeling of dynamic biological systems to better simulate real-time cellular activity and its involvement in molecular networks.3,4
Lastly, clinical translation of proteomics has recently gained momentum and is being integrated into diagnostics and therapeutic monitoring. Researchers believe that proteomics-based information will augment precision medicine, leading to more tailored therapeutic interventions and enhanced disease management. Continued improvements in mass spectrometry, bioinformatics, and data integration frameworks will be essential for bridging the gap between research and clinical application.3,4
Proteomics Enables Dynamic Modeling Of Cellular Systems
Proteomics is a key component in understanding biology and how proteins act, interact, and are regulated in complex cellular networks. Proteomics will enable modeling of dynamic pathway features and provide deeper insight into the underlying mechanisms of disease over time, as it captures real-time, dynamic molecular changes. By integrating with other omics layers, proteomics enhances predictive modeling and increases the potential to translate molecular data into biological or clinical insights.
Advances in analytical technologies and computational methodologies will further enhance the impact of proteomic data in biomedical research and precision medicine.
References
- Moore, J. B., & Weeks, M. E. (2011). Proteomics and systems biology: current and future applications in the nutritional sciences. Advances in nutrition. 2(4). 355-364. DOI:10.3945/an.111.000554, https://academic.oup.com/advances/article/2/4/355/4591630
- Simanjuntak, Y., Schamoni-Kast, K., Grün, A., Uetrecht, C., & Scaturro, P. (2021). Top-Down and Bottom-Up Proteomics Methods to Study RNA Virus Biology. Viruses. 13(4). DOI:10.3390/v13040668, https://www.mdpi.com/1999-4915/13/4/668
- Audagnotto, M., & Dal Peraro, M. (2017). Protein post-translational modifications: In silico prediction tools and molecular modeling. Computational and structural biotechnology journal. 15. 307-319. DOI:10.1016/j.csbj.2017.03.004, https://www.sciencedirect.com/science/article/pii/S2001037016301052
- Wörheide, M. A., Krumsiek, J., Kastenmüller, G., & Arnold, M. (2021). Multi-omics integration in biomedical research–A metabolomics-centric review. Analytica chimica acta. 1141. 144-162. DOI:10.1016/j.aca.2020.10.038, https://www.sciencedirect.com/science/article/pii/S0003267020314745
- Xie, Y., Chen, X., Xu, M., & Zheng, X. (2025). Application of the Human Proteome in Disease, Diagnosis, and Translation into Precision Medicine: Current Status and Future Prospects. Biomedicines. 13(3). 681. DOI:10.3390/biomedicines13030681, https://www.mdpi.com/2227-9059/13/3/681
Further Reading
Last Updated: Apr 6, 2026