How did you produce a 3D picture of the surface of the Ebola virus?
We used a very powerful electron microscope that can take high magnification images of individual Ebola proteins.
Did this 3D picture reveal weak spots on the surface of the Ebola virus?
Yes. We have found two primary weak spots and are searching for more.
In what way can this information be used to target antibodies or drugs?
We learned that two of the three antibodies in the Zmapp cocktail attack the same site, indicating that there may be some redundancy in the cocktail and it may be possible to reformulate Zmapp in a simpler two antibody format.
We also learned that one of the antibodies is unlikely to prevent viral entry into cells. Thus, it acts by binding infected cells and signalling the immune system to eliminate the cells before new viruses are released.
This multi-prong attack on viral entry and reproduction provides a powerful one-two punch against the virus.
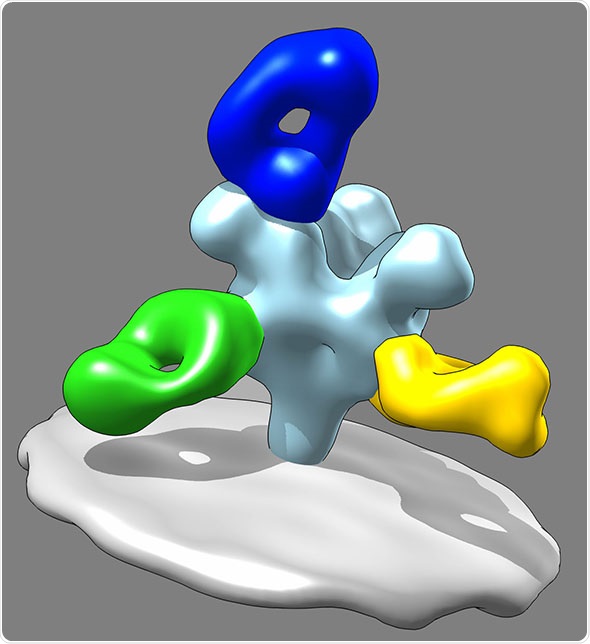
The structure of the ZMapp, an anti-Ebola antibody cocktail, binding to a part of the Ebola virus (in grey). Credit: Daniel Murin, Ward lab.
How many anti-Ebola antibodies have been discovered so far and how will these be compared?
There are hundreds of other antibodies against Ebola that we are in the process of imaging using the electron microscope.
We are looking for new sites of vulnerability as well as subtle differences in the way the known sites are attacked.
In particular we are looking for antibodies that the virus is unlikely to escape from when it mutates.
What are the main hurdles in formulating an immunotherapeutic cocktail to target Ebola?
The biggest hurdle is the clinical testing due to the expense and stringent safety conditions under which trials must be conducted.
Thus, we are using techniques such as electron microscopy to characterize the hundreds of available antibodies and make recommendations on the best combinations to advance to clinical testing.
How many genetic changes has the Ebola virus undergone during the current outbreak and are these likely to affect the weak spots of the surface of the virus?
Ebola has undergone over one hundred mutations and is still evolving. Luckily, the majority of these mutations are located in the so-called mucin domains that are not targeted by Zmapp antibodies.
What are the next steps for the National Institutes of Health-funded Viral Hemorrhagic Fever Immunotherapeutic Consortium?
Sorting through hundreds of antibodies that have been raised in the lab as well as from human survivors of Ebola in order to create better antibody cocktails and understand the mechanistic details of antibody-mediated Ebola therapy.
Will it be necessary to develop a back-up cocktail in case the Ebola virus mutates in a way that makes it resistant to treatment?
That is one of our primary concerns. Thus, by mapping all available antibodies we can predict which ones will be effective against particular mutants.
Where can readers find more information?
About Andrew Ward
Andrew B. Ward obtained a B.S. in Biology at Duke University and Ph.D. in Biophysics at The Scripps Research Institute in the laboratory of Ronald Milligan where he studied membrane proteins by electron microscopy.
During his postdoctoral work in the laboratory of Geoffrey Chang at The Scripps Research Institute he continued to study various membrane proteins by X-ray crystallography.
Dr. Ward became an Assistant Professor at The Scripps Research Institute in 2010 and his lab studies membrane proteins including viral envelope proteins that reside on the surface of viruses are the targets of antibodies.
His lab has developed a pipeline to address a variety of biological and biomedical questions, including how viruses are recognized and neutralized by the human immune system, and how we can use the structural information to design novel or better vaccines.
More recently his lab has also begun to study the molecular mechanisms of antibody-mediated protection against viruses.