An interview with Kimberley Wicklund, conducted by James Ives, MPsych. Essen Bioscience is a Sartorius brand.
Cells, and cell interactions are extremely complex and highly dynamic. Please give an overview of the challenges facing researchers when analyzing these complex systems.
Your first sentence hit on a key challenge: dynamics. Every cell is a miniature factory taking in raw material (extracellular signals) and processing it to produce something (a biological response). But unlike a factory that can be easily tweaked to run at steady-state and look the same from day to day, cells are dynamic.
CHO HD edited
If you are using cells to investigate responses, how do you know when the response happens? What if you miss it? Knowing what your cells are doing and when they do it is a key challenge. Another challenge is physiological relevance. If you are looking at cell lines or primary cell cultures, great pain is taken to keep them in a consistent environment.
But then they are taken in and out of their happy homes to be poked and prodded on analytical instruments. How do these changes in environment affect cellular health and function? As we progress towards more complex or sensitive models, maintaining environmental conditions consistent is not only important for keeping the cells alive, but also for maintaining relevance to the original biological system and for ensuring reproducibility of your experiment.
Another challenge in maintaining physiological relevance arises when we use reagents to detect biological changes of interest. Most reagents perturb the very biology we are investigating. It’s important to consider how reagents affect cell health and morphology before using them with living cells.
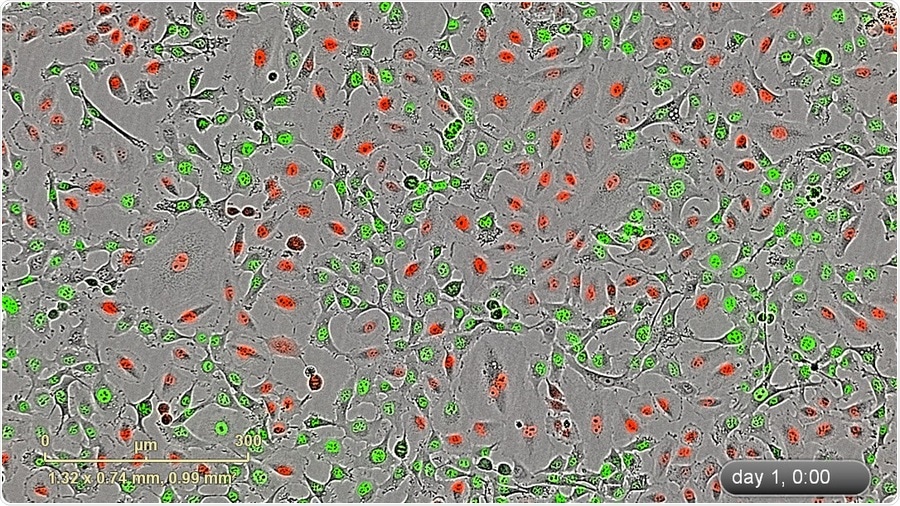
IncuCyte blended HD-phase contrat/fluorescence image (10x) of a mixed stable population of HT-1080 NucLight Green and A549 NucLight Red cells.
Until recently what has been used as the gold standard in analytical methods of cell biology?
For cell biology, key analytical methods have involved instruments such as microscopes, flow cytometers or plate readers used in conjunction with colorimetric or fluorescent reagents to aid in detection of the biology of interest.
Development of fluorescent proteins, of course, has been a key advancement for performing cell analysis. These proteins can be constitutively expressed without significant disruption to normal cell behavior and have truly enabled live-cell imaging.
Please give an overview of the Incucyte S3 and its features.
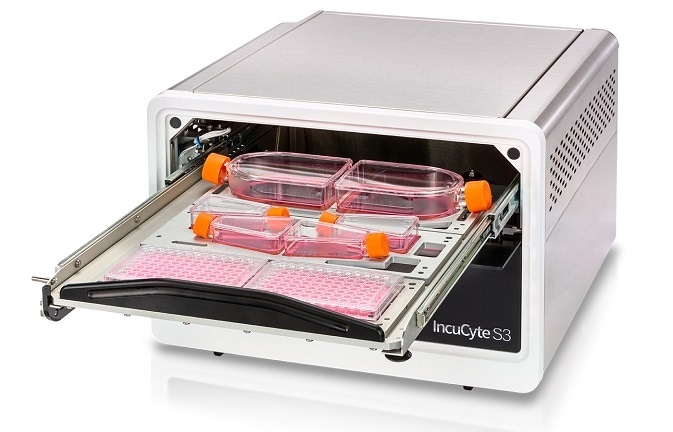
The IncuCyte® S3 is the only real-time live-cell imaging and analysis system that automatically captures and analyzes images of living cells around the clock, for days, weeks, or months, while cells remain undisturbed inside a standard tissue culture incubator.
IncuCyte® S3 Live-Cell Analysis System
The IncuCyte has a unique, compact and mobile optical design that allows cells to stay stationary throughout the experiment, which is crucial for non-adherent or sensitive cell types such as immune cells or neurons. Our approach is application-focused and cell health-focused.
Via a recently updated user interface, the IncuCyte is achieving “walk-up and use” status for live-cell imaging and analysis. Plus, the IncuCyte achieves this at scale. A new user can conduct 96- and 384-well experiments in a matter of minutes, and then create movies and graphing metrics that are publication- and presentation-ready with just a few clicks.
I still remember my first day at Essen where I learned how to do a 96-well scratch wound assay using the IncuCyte and our Woundmaker. It took no more than a half hour of hands-on time to seed, create 96 scratches, put the microplate in the IncuCyte, and set up the software to acquire and automatically analyze.
HT-1080 Cell Invasion through Matrigel
Graphs began building immediately, truly real-time analysis. When we finished, I said “Is that it???”. That was it, very simple and accessible, and very very powerful.
In addition to improving productivity for the individual user, the IncuCyte S3 empowers an entire research team. Multiple users can run multiple applications on the IncuCyte in parallel, reducing time spent waiting for an instrument to be available. In addition, data is accessible remotely to any user via unlimited, free networked licenses.
With the new IncuCyte S3, researchers are able to devise new experiments not previously thought possible. They now have access to real-time live-cell analysis that offers not only new insights, but also throughput and a streamlined user experience.

Please give an overview of the importance of ‘real-time’ live-cell imaging and analysis? Why is it so important to keep live-cell samples rather than using procedures that destroy the cells as part of the experimental workflow?
Rather than a single snapshot, continuous analysis ensures you never miss a critical response or interaction. Furthermore, kinetic, image-based measurements provide more physiologically relevant data and deeper, more meaningful insights into active biological processes and cellular function that cannot be achieved with conventional endpoint assays.
To get to an understanding of a time-dependent response with endpoint assays, it requires setup of multiple assays and concatenation of the data to put together a picture of what happened. That is time-consuming and expensive, and you have no ability to further analyze your samples once you have conducted the assay.
Furthermore, if your technology is not image-based, you will not be able to visually validate any trends or assess outliers. With real-time imaging and analysis using non-perturbing reagents, each well or vessel can be analyzed over time to reveal concentration-dependent or cell-specific responses that can be validated with images and movies and enable real-time decision-making.
You also have the benefit that your cells are not destroyed during the experimental process, and can be characterized further using other technologies such as flow cytometry.
Aside from the insight you gain using continuous live-cell imaging and analysis, you also can quickly assess any outliers, as you can evaluate seeding density and morphology of your cells at the start of the experiment and throughout. The quality control aspect of performing live-cell analysis cannot be overlooked.
What applications can be studied when using the Incucyte S3? Are there limitations or restrictions?
The IncuCyte S3 is compatible with a wide range of culture vessels and applications. The IncuCyte S3 System can be used for real-time cell monitoring and surveillance, assaying cell health and viability, migration and invasion, plus a wide range of phenotypic cell-based assays.
Applications for the IncuCyte® Live-Cell Analysis System
The IncuCyte takes a population-based approach to cell analysis. If you are interested in investigating subcellular structural details with a live-cell approach, we recommend first using the IncuCyte to characterize your cells and investigate variables at the population level, and then follow up with a higher resolution technique such as confocal microscopy.
The IncuCyte has a rapidly growing collection of over 1,000 peer reviewed articles, and new applications using the IncuCyte are being published continuously.
Why is it important to be able to leave your cells within the incubator, with imaging equipment inside? What advantages are there to this method?
One day, years ago, one of our founders was walking around the lab and observing scientists walking back and forth to the incubator, taking cells out and bringing them to an instrument to be checked.
He asked “Do you think the cells like that? Wouldn’t it be better if you could see your cells while they are sitting inside the incubator?” That is the beginning of the IncuCyte story and a perfect example of “design thinking” - observe the user in their natural environment and solve a problem with innovation.
The tissue culture incubator is a highly trusted and precise piece of equipment that keeps a large volume of gas at an appropriate mixture, temperature and humidity. It’s designed to keep cells alive.
By putting the IncuCyte inside the incubator, we leverage that trusted environment. Of course, there are instruments that have built-in environmental control or add-on accessories, but precise regulation of substantially smaller volumes of gas mixtures can be cumbersome and difficult.

How does the image processing and analysis software work to help automate experimental analysis? What applications are there for this automated analysis software?
I have spent a substantial portion of my career learning various image processing and analysis software platforms and guiding scientists on their usage. Many software platforms take a “tool box” approach.
With a traditional imaging system, I have to first take steps to calibrate the images and correct artifacts from the imaging device and illumination system, then correct artifacts from the sample itself, then figure out how to draw out my features of interest with a dazzling array of image processing filters, and finally, I work through an assortment of metrics to analyze….a single image.
To repeat this procedure, I build a script or hopefully there is some batch process menu. It is cumbersome, to say the least, and I am constantly trying to remember exactly what steps I took and where my images are.
The IncuCyte offers several advantages. First, the instrument has a calibration process that corrects artifacts automatically. Second, the software interface presents only the tools you need for the task at hand.
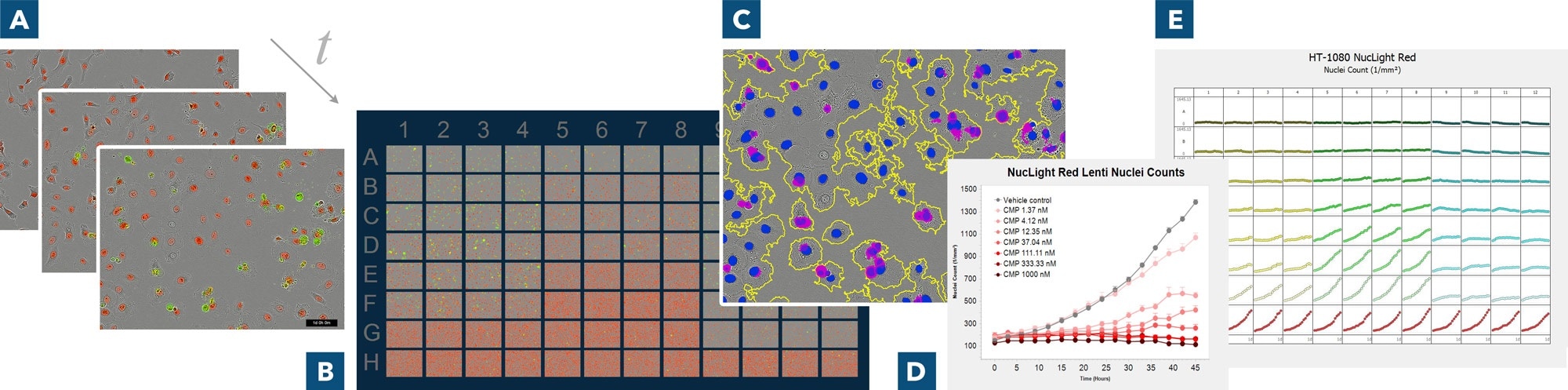
Derive deeper insights with real-time live-cell analysis. (A) Automatically acquire images over time, (B) use IncuCyte® VesselView to view images of all locations in the vessel at once and quickly assess experimental results, plus zoom in on images of interest (C) automatically identify regions of interest via masks (D) generate presentation-ready timelapse graphs, and (E) view all 96- or 384-well kinetic trends at once with IncuCyte® PlateGraph and export data to calculate EC50 or IC50 response values.
If you are doing a cell proliferation assay that is counting red blobs of nuclear protein, we present you the tools to count red blobs. If the analysis is more complex, like analyzing neurite outgrowth, we give you the tools do that as well. Third, the analysis is applied to your entire experiment, all time points and all wells automatically.
Users are able to set up a processing definition one time using a representative experiment, and then apply that week after week, month after month with a click of a button. Last, but most certainly not least, all of the images and analyses are neatly stored together in a searchable database.
What can be gained by observing cell morphology and cell movements during studies?
Observing changes in cell morphology and movement provide insight into many normal and pathological processes including immune response and tumor metastasis. The ability to study these changes kinetically and in physiologically relevant conditions is pertinent to understanding the effects of drug treatments and subsequent cellular response, allowing you to precisely pinpoint when these changes occur.
Visual assessment of cell morphology and movement, not only verifies the quantitative data, but ultimately allows for additional phenotypic information in order to profile cell-specific and time-dependent biological activity.
What does the future hold for IncuCyte S3 technology? How will this further help researchers to conduct experiments in the future?
We do have some exciting developments ahead. While I can’t give too much detail, we are actively assessing more complex models such as spheroids and also iPSC-derived cells, such as neurons. We are always pushing towards more physiological relevance and deeper insight into function. Make sure to follow us on Facebook and Linkedin to keep up to date with our activities!
Where can readers find more information?
Please visit our website at www.incucyte.com.
About Kimberley Wicklund
Kimberly Wicklund spent a decade working with research microscopy hardware and software prior to joining Essen in 2015. She began her career in life science instrumentation as an applications scientist and technical support manager before landing in product management.
She has launched a number of imaging systems, devices, and software platforms to the market and has served as vendor faculty at many microscopy courses at Wood’s Hole, Cold Spring Harbor, and Mount Desert Island Biological Laboratories.
She obtained her PhD in chemical engineering at the University of California-Berkeley and studied breast cancer cell metabolism in hypoxic conditions, so she has long appreciated the importance of physiologically relevant environments. She is now the Director of Product Management at Essen Bioscience, a Sartorius company and lives in Ann Arbor, Michigan.