An interview with Dan Appledorn and Tim Dale, conducted by James Ives, MPsych. Essen Bioscience is a Sartorius brand.
Please give an overview of how assays of complex cellular models have changed in recent years. What implications have these changes had on the types of cells and the applications that can be studied?
Tim Dale: There's a continued drive towards making in vitro assays ever more translational towards in vivo models and ultimately the clinic. This ties in with the resurgence of phenotypic screening and is a response to the perceived poor translation of the traditional simple cell-based assays, often developed to study just single protein targets.
 © Rattiya Thongdumhyu/Shutterstock.com
© Rattiya Thongdumhyu/Shutterstock.com
To meet these challenges, we’re aiding researchers study complex phenotypic models, where both the microenvironment that the cells reside, and the cell types, have been made to be more representative of the real-world physiology.
For example, co-cultures can dramatically alter cell behavior, such as the growth of supportive astrocytes in neuronal cultures. Additionally, the enthusiasm towards 3D models offers many new opportunities. Oncology researchers for example, may be interested in the existence of the hypoxic core, which is only achievable in a tumor spheroid. The presence of a biomatrix may also provide structural and chemical cues which are more relevant to physiology.
The ability to investigate 3D structures can then potentially lead onto organoid cultures, which ultimately drive towards the development of organ-on-a-chip, allowing researchers to study real world organs in an in vitro environment.
Recombinant, inexhaustible cell types have their utility, but have less translation potential. More and more research in this area is focused on working with induced pluripotent stem cell (iPSC)-derived cells and patient-specific tissues to make a more translational system.
It is also worth noting that as cell types become ever more scarce, and expensive, the need for improved QC and understanding around cell handling only increases. Therefore, the division between cell husbandry and assay becomes ever more blurred.
With an increase in complexity and a move towards spheroid models over 2D assays, how have the research methods adapted and improved? What challenges have these new techniques raised for researchers in the field?
Dan Appledorn: Now that we know that these cultures are more translatable, there's an energy to do more work with them, in order to do so, we've had to develop new auto-focusing and image-processing algorithms to accommodate the new ways in which these cells are cultured, either in 3D matrices, single spheroids, multi-spheroid and models with multiple cell types.
Traditionally it has been difficult, from a biochemistry perspective, to tease out phenotypic changes when two (or more) different cell types are cultured together. For example, if you conducted a luciferase assay in a co-culture, you wouldn’t know which cell type the read-out is coming from. The same applies to sequencing methodologies, RNA-Seq or other transcriptional analyses. Imaging technologies can partially solve that problem by labeling individual cell types enabling more sophisticated measurements in co-culture environments.
We've accomplished this in spheroid cultures, and we are also initiating multi-spheroid analyses. Additionally we’ve been working in more complex iPSC-derived neuronal models, in which we are making both structural (neurite-length) and functional (calcium transients) measurements from neurons in the presence of other cell types (rodent and/or human astrocytes).
What advantages are there for live-cell imaging research of complex cellular models over end-point assays? What does Sartorius offer in the live-cell imaging and analysis space?
Tim Dale: The time course of the complex models is difficult to predict, making it tough to take a measurement at the right time. Live-cell imaging and analysis removes the need to make that estimation. I believe this is the primary advantage of non-invasive kinetic readouts of live-cell imaging of complex models over end-point assays.
Data is captured automatically and can be looked at after the event to determine the ideal timepoint. This is particularly telling when you see multiphasic time courses of biology.
A specific example would be in studies we've conducted investigating Notch signaling within vascular tube formation in an angiogenesis application. At the early timepoints, inhibition of notch signaling has little or no impact on the vascular formation.
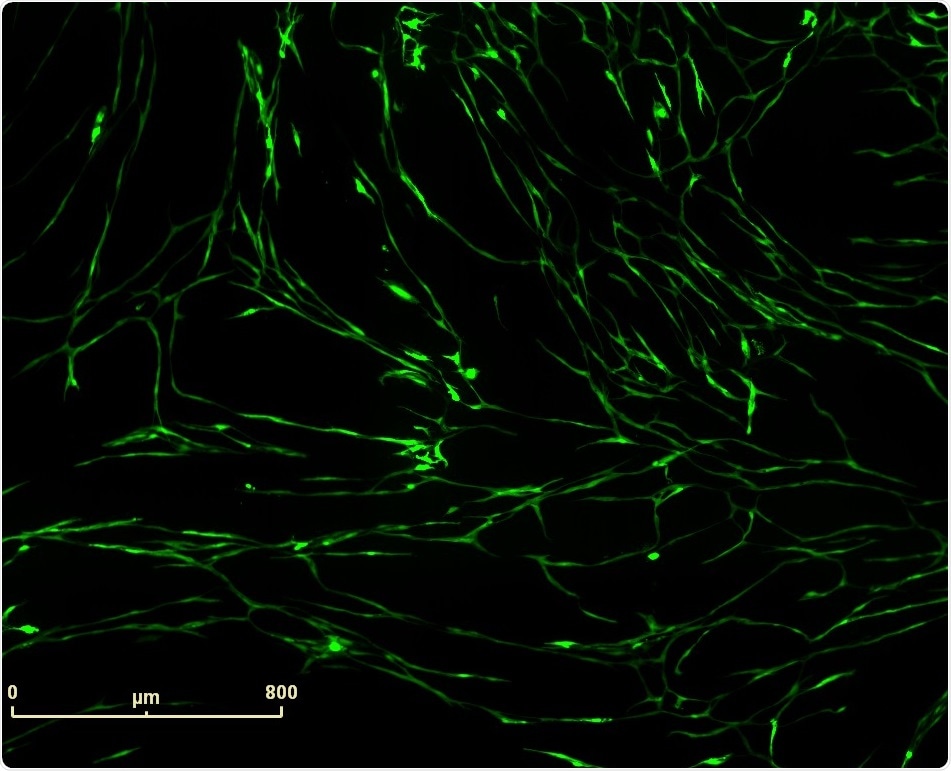 Vascular tube formation induced by Vasoactive Endothelial Growth Factor (VEGF) in PrimeKit HUVEC and human dermal fibroblast co-culture model. IncuCyte™ (10x) fluorescence image of VEGF (4 ng/ml) induced vascular structures.
Vascular tube formation induced by Vasoactive Endothelial Growth Factor (VEGF) in PrimeKit HUVEC and human dermal fibroblast co-culture model. IncuCyte™ (10x) fluorescence image of VEGF (4 ng/ml) induced vascular structures.
However, at very late timepoints, after a week of complex co-culture of endothelial cells with stromal fibroblasts, you start to see a notable change in the differentiation of the endothelial cells, leading to a striking change in morphology, which the imaging and analysis platform can decipher and use to provide mechanistic insight.
Being non-invasive, we are not ruling out endpoint analysis. People can still take the plates that we've read and move them across into other platforms such as flow cytometry or high-content analysis, so live-cell imaging and analysis can be seen as a complimentary approach.
Another aspect is the ability to apply data normalization. With an endpoint read-out, scientists can only compare one condition to another at a given moment in time.
With the ability to look at the full-time course of the biology, researchers can analyze observations after treatment w.r.t before treatment, which can add more statistical power and understanding of what’s going on.
For example, if you see an effect at the end of a study, you can ask yourself “did that truly happen after I added my treatment, or was there something fundamentally different within the well before I added my treatment?” You would never have that insight unless you could look at the data over the entire time course.
Do you think giving researchers access to the entire time course of their samples, opens research up to fraudulent behavior? Could scientists look for statistically significant timepoints and stop their experiments prematurely, disregarding subsequent data?
Tim Dale: I certainly wouldn't think of that as a problem. Instead, having the full time-course adds a huge amount of extra information which can inform the study, and I don't know of any researcher who would select the timepoint to specifically skew the data.
Researchers can be selective, but the full-time course is available to show, and specific questions can be asked about why effects occurred at a certain time point versus others.
In the notch signaling example, the insight that the full time-course gave was consistent with other known effects of notch signaling on the differentiation of endothelial cells, so it was the linking of specific time points to known pathways that really added the power.
Human induced pluripotent stem cells (hiPSCs) are a good example of a research area that requires long-term culture and analysis to model neurological disease. How does Sartorius’ IncuCyte S3 help neurological researchers to improve the quality and consistency of their results?
Dan Appledorn: It's only been 10 years since Professor Yamanaka showed that you could take a human fibroblast and reprogram or de-differentiate that fibroblast into a pluripotent state.
The technology for developing iPSCs is still in its infancy. It has only been in the last five to six years that researchers have figured out how to take those iPSCs and differentiate them into different cell types, with cardiomyocytes, hepatocytes and neurons leading the way.
The characterization of the cells that come out of hiPSC-derived differentiation pathways are understudied because researchers don’t have the tools required to ask the right questions, this is where we think the IncuCyte can provide a lot of advantages.
Using iPSC-derived neurons as an example, differentiating iPSCs into neurons can take 6 or 7 days for induced-neurons or as long as 55 days, where cells going through embryoid bodies, primitive neural stem cells, neural rosettes, neural stem cells and then ultimately neurons.
That is a significant period of time, during which researchers want to be monitoring the health, morphology and fucntional activity of the cells, as well as the neurite outgrowth.
The IncuCyte can monitor all these attributes, allowing researchers to optimize critical components of the cellular environment, e.g. growth factors, medias and surface coatings. Ultimately, the goal is to develop the most translatable models in order to positively impact our knowledge and treatment of human disease.
Sartorius has the application specific NeuroTrack Software. How might this help neurologists to analyze their data?
Dan Appledorn: NeuroTrack Software allows you to make neurite outgrowth measurements in either phase contrast or in fluorescence.
Initially our NeuroTrack Software was designed to make measurements of neurite outgrowth in monocultured neurons. Over time, we have learned that researchers are interested in more complex cultures and in making neurite outgrowth measurements in co-cultures.
Therefore, we also provide the software, image acquisition tools, reagents and protocols that enable researchers to make measurements of neurite outgrowth in mixed cultures.
Unsurprisingly, we’ve also learned that making simple measurements of neurite outgrowth does not necessarily predict neuronal function. As a result, we're launching a new application that measures spontaneous neuronal activity by capturing movies of calcium transients within neurons. Interestingly, we've observed instances where the neurite outgrowth can be quite extensive, yet no activity can be measured.
With this new application, the IncuCyte technology now enables parallel measurements of both neurite outgrowth and spontaneous neuronal activity within the same instrument and in the researchers own incubator.
How can real-time kinetic data provide researchers with a better understanding of their assays?
Tim Dale: Understanding the temporal profile of biological modulation can provide an understanding into the mechanism of action. Often the insight is even more powerful when you don't get the expected outcome.
The ability to look back through time to understand what happened can be very insightful. By being able to see the full time course researchers can be sure that there wasn’t an initial effect that was then masked later on, which is a powerful example of how kinetic data can be really useful.
One example is in cancer cell migration, when PI-3 kinase inhibitors are used, observations after 24 hours suggests the PI-3 kinase inhibitor has little or no effect, whereas if you look after 8 hours it has had a marked effect.
PI-3 seems to just cause a delay in migration of those cells and that cell type, which would be missed with an end-point assay.
That understanding of the alteration through the kinetic profile and kinetic signature of particular biological events, would be lost without this kinetic data.
Further, by having kinetic data as a whole, researchers can start to utilize parameters that were previously completely unthought of, such as the rate of change in biological process.
Also, with real-time data, i.e. analyzed images, rather than just the images themselves, decisions can be made about the biology as it happens, which may change the experimental design according to the biology rather than the design remaining static, based on previous assumptions.
How is throughput affected using the IncuCyte S3 live-cell system?
Dan Appledorn: The maximum capacity of an IncuCyte S3 system is six 384 well plates. When imaging all those plates twice a day for three days, that results in more than 13,800 data points or images acquired that are automatically processed and automatically collated.
These can be displayed in a graphical way in real-time so that researchers can make scientific decisions very quickly.
Most of the time, the IncuCyte is being used as a secondary screening tool. We consider it a moderate throughput instrument, but the data acquired is certainly very rich and enables quick scientific decisions.
Is there a high level of technical expertise required?
Dan Appledorn: I think we have one of the easiest-to-use assay systems available on the market. Our goal while developing our instrumentation, software, protocols and applications, has been to make the applications and all the tools associated with it available to any level of researcher.
IncuCyte® S3 Live-Cell Analysis System
Image acquisition, image processing, and data display tools are designed with the end-user in mind. Our software engineers have developed simple, easy to understand tools that are validated by our team of biologists.
Are there high costs to run these assays?
Dan Appledorn: It really depends on the application. Label-free applications have no additional cost w.r.t consumables, cells or media because the similar cells and media would be used as in any traditional assay.
Additional costs are only incurred when specialized live-cell reagents need to be added. For example, in neurite outgrowth applications in a co-culture where labeling or apoptosis reagents are used.
However, those reagent costs aren't any different from those typically found for other biochemistry analyses. This type of analysis also provides opportunities to multiplex different measurements, generating additional data on top of label-free data, thus increasing value to the researcher.
I would argue that the enabling aspect of the IncuCyte, and the rich, kinetic data it generates leads to higher quality, and faster scientific decision making, ultimately saving a significant amount of time and money in the long term.
Is cleaning and sterilization of the IncuCyte S3 also automated? What mechanisms are in place to ensure that there is no contamination between assays?
Tim Dale: Currently, the sterilization of IncuCyte is not automated and relies upon a similar process employed for a standard incubator. Periodically, the IncuCyte can be sterilized using a hydrogen peroxide fumigation, either in situ inside the incubator or it can be removed and sent back for sterilization here at Sartorius.
With recent advancements in situ hydrogen peroxide sterilization is becoming an ever-simpler process and doing that with the IncuCyte still residing inside the incubator is becoming routine.
We have never observed any greater occurrences of contamination inside the IncuCyte than inside a standard incubator. There also has been no evidence that any higher level of sterilization and fumigation is required.
When we think more specifically about a contamination between assays, or plates or wells. It's worth noting that the plates, flasks and any other plasticware inside the IncuCyte, do not move, it is the microscope imaging system, that moves to the various locations.
Therefore, the risk of contact-based contamination is minimal, and the risk of aerosol generation is mitigated by the stationary nature of the plasticware.
On the rare occasions we have seen infections in specific wells, we've been able to attribute this to operator error during re-feeding or treatment points, and never seen any cross-contamination from one plate to another.
What new challenges have arisen with the use of real-time live-cell imaging and analysis of complex cellular models?
Dan Appledorn: We are challenged every day. There are challenges both on the technology and the application side. With our advancements, new areas and concepts are triggered in researcher’s’ minds and therefore the requests they make have also expanded.
The possible live-cell imaging applications are endless. Our challenge is to take on the right problems and provide the support needed to develop applications which will answer the questions most important for researchers.
One big challenge has been providing reagents. A big focus for us and a requirement for live-cell studies are that the reagents are non-perturbing to the biology that researchers want to interrogate.
Unfortunately, the majority of available of reagents outside of our IncuCyte reagents are perturbing to biology. In some cases, they're called live-cell reagents, but those are only intended to label the cells when they're living, without fixing them. The act of putting the reagent on the cells may actually kill them.
Our customers quickly find out that there aren't many reagents out there that are non-perturbing to the biology. Therefore, onus is on us at Sartorius to provide the reagents for our customers. That is a challenge we are certainly happy to have.
In a similar vein, the complexity of the model systems that are being used by researchers poses a challenge in terms of image acquisition, auto-focusing algorithms, image-processing algorithms and reagents. The tools we provide are constantly being improved in order to facilitate measurements in increasingly complex systems.
Lastly, designing an instrument that functions inside of an incubator where the humidity is over 90% and the temperature is at 37 degrees Celsius (98.6oF) is not easily accomplished. It is a testament to our engineering team here at Sartorius.
What does the future hold for real time live-cell imaging and analysis of live-cells? What developments are there for the IncuCyte S3 technology and the accompanying analysis software?
Tim Dale: In general terms, Sartorius' ethos has been of offering application-centric solutions and I think that any future developments will follow this approach.
We need to ensure that we maintain the principles of simple workflows and fully integrated solutions, such that the researcher can focus on the biology and not get distracted by the technology.
Although we've talked throughout about how the IncuCyte S3 enables researchers to approach a broad range of assays, there's still a large number that we are striving to develop.
We have a continued focus around 3D in vitro biology and want to extend further, not only in tumor cell lines for oncology research, but we are keen to support organoid research and ultimately be able to think about creating organs in wells.
Where can readers find more information?
About Tim Dale
 Tim co-founded the European laboratories of Essen BioScience in 2009 and is currently Director of Biology, managing R&D operations. A highly experienced industrial pharmacologist and leader with >20 years of service, Tim has a track record in drug discovery.
Tim co-founded the European laboratories of Essen BioScience in 2009 and is currently Director of Biology, managing R&D operations. A highly experienced industrial pharmacologist and leader with >20 years of service, Tim has a track record in drug discovery.
Appling his knowledge and expertise to drug discovery programs in the neurology, psychiatry and gastrointestinal therapeutic areas, he has contributed to the progression of a number of candidate molecules into clinical studies.
Tim has co-authored over 20 publications and patents relating to pharmacology, ion channels and cell-based assays. Most recently, Tim is applying his many years of experience to extend the portfolio of cell-based applications utilizing live-cell analysis in the fields of oncology, immunology and neuroscience.
About Dan Appledorn
 Dan is the Director of US Biology R&D at Essen BioScience. Dan joined Essen in the fall of 2010 and is leading a team of scientists to develop new applications, biological reagents and instruments for drug discovery with a focus on building assays for quantitative live cell analysis using the IncuCyte system.
Dan is the Director of US Biology R&D at Essen BioScience. Dan joined Essen in the fall of 2010 and is leading a team of scientists to develop new applications, biological reagents and instruments for drug discovery with a focus on building assays for quantitative live cell analysis using the IncuCyte system.
Prior to joining Essen, Dan was a postdoctoral research fellow at Michigan State University where his primary focus was investigating the interaction between adenovirus vectors and the innate and adaptive immune systems in mouse models with the goal of developing gene therapy and/or vaccine vectors for the treatment of human disease.
Over the course of his 4 year postdoc, Dan’s pioneering work in this field resulted in several patents and 20+ publications describing the utility of Ad vectors in both gene therapy and vaccine platforms.